Impact Factor
Theranostics 2022; 12(12):5522-5536. doi:10.7150/thno.74428 This issue Cite
Research Paper
DNA aptamers inhibit SARS-CoV-2 spike-protein binding to hACE2 by an RBD- independent or dependent approach
Department of Chemistry and Center for Photochemical Sciences, Bowling Green State University, Bowling Green, Ohio 43403, United States.
Abstract
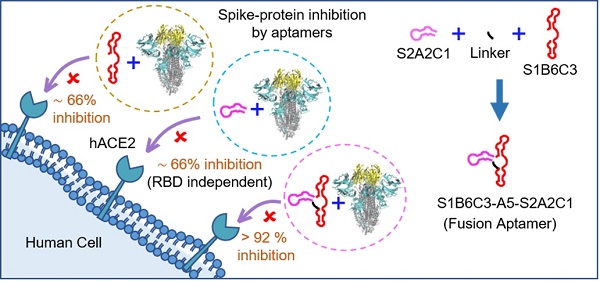
Objective: Nobody knows when the COVID-19 pandemic will end or when and where the next coronavirus will outbreak. Therefore, it is still necessary to develop SARS-CoV-2 inhibitors for different variants or even the new coronavirus. Since SARS-CoV-2 uses its surface spike-protein to recognize hACE2, mediating its entry into cells, ligands that can specifically recognize the spike-protein have the potential to prevent infection.
Methods: We have recently discovered DNA aptamers against the S2-domain of the WT spike-protein by exploiting the selection process called SELEX. After optimization, among all candidates, the aptamer S2A2C1 has the shortest sequence and the best binding affinity toward the S2-protein. More importantly, the S2A2C1 aptamer does not bind to the RBD of the spike-protein, but it efficiently blocks the spike-protein/hACE2 interaction, suggesting an RBD-independent inhibition approach. To further improve its performance, we conjugated the S2A2C1 aptamer with a reported anti-RBD aptamer, S1B6C3, using various linkers and constructed hetero-bivalent fusion aptamers. Binding affinities of mono and fusion aptamers against the spike-proteins were measured. The inhibition efficacies of mono and fusion aptamers to prevent the hACE2/spike-protein interaction were determined using ELISA.
Results: Anti-spike-protein aptamers, including S2A2C1 and S1B6C3-A5-S2A2C1, maintained high binding affinity toward the WT, Delta, and Omicron spike-proteins and high inhibition efficacies to prevent them from binding to hACE2, rendering them well-suited as diagnostic and therapeutic molecular tools to target SARS-CoV-2 and its variants.
Conclusions: Overall, we discovered the anti-S2 aptamer, S2A2C1, which inhibits the hACE2/spike-protein interaction via an RBD-independent approach. The anti-S2 and anti-RBD aptamers were conjugated to obtain the fusion aptamer, S1B6C3-A5-S2A2C1, which recognizes the spike-protein by an RBD-dependent approach. Our strategies, which discovered aptamer inhibitors targeting the highly conserved S2-protein, as well as the design of fusion aptamers, can be used to target new coronaviruses as they emerge.
Keywords: SARS-CoV-2, DNA aptamer, S2-protein, fusion aptamer, spike-protein
 Global reach, higher impact
Global reach, higher impact