13.3
Impact Factor
Theranostics 2022; 12(10):4779-4790. doi:10.7150/thno.72339 This issue Cite
Review
Engineering viral genomics and nano-liposomes in microfluidic platforms for patient-specific analysis of SARS-CoV-2 variants
1. Department of Bioengineering, School of Engineering, University of California, Los Angeles, California, USA.
2. Skin Research Center, Shahid Beheshti University of Medical Science, Tehran, Iran.
3. Medical Engineering, California Institute of Technology, California, Pasadena, USA.
4. Department of Bioengineering, Henry Samueli School of Engineering & Applied Science, University of California, CA, USA.
5. Institute for Quantitative Health Science & Engineering and Department of Biomedical Engineering, College of Engineering, Michigan State University, MI, USA.
6. Department of Medicine, Greater Los Angeles VA Healthcare System, Los Angeles, California, USA.
7. Division of Cardiology, Department of Medicine, School of Medicine, University of California, Los Angeles, California, USA.
8. Section of Hematology, Oncology, and Blood & Marrow Transplantation, Department of Medicine, College of Medicine, University of Iowa, Iowa, USA.
9. Department of Microbiology and Immunology, College of Medicine, University of Iowa, USA.
Abstract
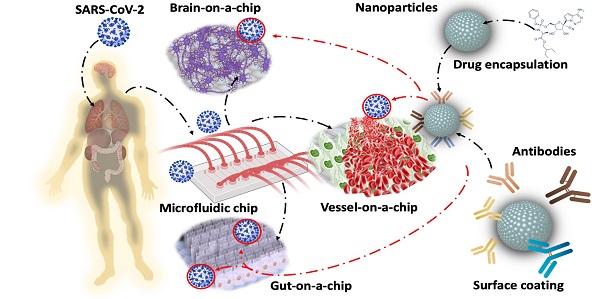
New variants of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) are continuing to spread globally, contributing to the persistence of the COVID-19 pandemic. Increasing resources have been focused on developing vaccines and therapeutics that target the Spike glycoprotein of SARS-CoV-2. Recent advances in microfluidics have the potential to recapitulate viral infection in the organ-specific platforms, known as organ-on-a-chip (OoC), in which binding of SARS-CoV-2 Spike protein to the angiotensin-converting enzyme 2 (ACE2) of the host cells occurs. As the COVID-19 pandemic lingers, there remains an unmet need to screen emerging mutations, to predict viral transmissibility and pathogenicity, and to assess the strength of neutralizing antibodies following vaccination or reinfection. Conventional detection of SARS-CoV-2 variants relies on two-dimensional (2-D) cell culture methods, whereas simulating the micro-environment requires three-dimensional (3-D) systems. To this end, analyzing SARS-CoV-2-mediated pathogenicity via microfluidic platforms minimizes the experimental cost, duration, and optimization needed for animal studies, and obviates the ethical concerns associated with the use of primates. In this context, this review highlights the state-of-the-art strategy to engineer the nano-liposomes that can be conjugated with SARS-CoV-2 Spike mutations or genomic sequences in the microfluidic platforms; thereby, allowing for screening the rising SARS-CoV-2 variants and predicting COVID-19-associated coagulation. Furthermore, introducing viral genomics to the patient-specific blood accelerates the discovery of therapeutic targets in the face of evolving viral variants, including B1.1.7 (Alpha), B.1.351 (Beta), B.1.617.2 (Delta), c.37 (Lambda), and B.1.1.529 (Omicron). Thus, engineering nano-liposomes to encapsulate SARS-CoV-2 viral genomic sequences enables rapid detection of SARS-CoV-2 variants in the long COVID-19 era.
Keywords: Organ on-a-chip, microfluidics, viral genomics, mutations, variants of concerns, COVID-19, nano-liposomes
 Global reach, higher impact
Global reach, higher impact