Impact Factor
Theranostics 2022; 12(1):290-306. doi:10.7150/thno.63463 This issue Cite
Research Paper
Integrated analysis of circulating immune cellular and soluble mediators reveals specific COVID19 signatures at hospital admission with utility for prediction of clinical outcomes
1. Fundación Instituto de Investigación Sanitaria Aragón (IIS-Aragón), Biomedical Research Centre of Aragon (CIBA), 50009 Zaragoza, Spain.
2. Servicio de Enfermedades Infecciosas, Hospital Clinico Universitario Lozano Blesa, 50009 Zaragoza, Spain.
3. Unit of Biomarkers and Susceptibility, Oncology Data Analytics Program (ODAP), Catalan Institute of Oncology (ICO), Oncobell Program, Bellvitge Biomedical Research Institute (IDIBELL) and CIBERESP, L'Hospitalet de Llobregat, Barcelona, Spain.
4. Department of Immunology, University Clinic Hospital Lozano Blesa, 50009, Zaragoza, Spain and Department of Pathology, University Clinic Hospital Lozano Blesa, University of Zaragoza, IIS Aragón, 50009 Zaragoza, Spain.
5. Instituto de Carboquímica ICB-CSIC, 50018 Zaragoza, Spain.
6. Biobanco de Aragón, Instituto Aragonés de Ciencias de la Salud (IACS), 50009, Zaragoza, Spain.
7. Servicio de Enfermedades Infecciosas, Hospital Clinico Universitario Lozano Blesa, 50009 Zaragoza, Spain.
8. Nanoscience Institute of Aragon (INA), Consejo Superior de Investigaciones Científicas (CSIC), University of Zaragoza, 50018 Zaragoza, Spain.
9. Department Microbiology, Pediatry, Radiology and Public Health, University of Zaragoza.
10. Unidad de nanotoxicología e inmunotoxicología experimental (UNATI). Fundación Instituto de Investigación Sanitaria Aragón (IIS-Aragón), Biomedical Research Centre of Aragón (CIBA), 50009 Zaragoza, Spain.
11. Centro de Investigación Biomédica en Red (CIBER) de Enfermedades Infecciosas, 50018 Madrid, Spain.
12. Aragon I+D Foundation (ARAID), Zaragoza, Spain.
* These authors contributed equally
# These authors share corresponding and senior authorship
Abstract
Coronavirus disease 2019 (COVID19), caused by SARS-CoV-2, is a complex disease, with a variety of clinical manifestations ranging from asymptomatic infection or mild cold-like symptoms to more severe cases requiring hospitalization and critical care. The most severe presentations seem to be related with a delayed, deregulated immune response leading to exacerbated inflammation and organ damage with close similarities to sepsis.
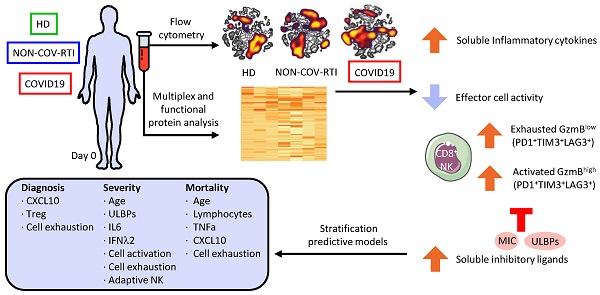
Methods: In order to improve the understanding on the relation between host immune response and disease course, we have studied the differences in the cellular (monocytes, CD8+ T and NK cells) and soluble (cytokines, chemokines and immunoregulatory ligands) immune response in blood between Healthy Donors (HD), COVID19 and a group of patients with non-COVID19 respiratory tract infections (NON-COV-RTI). In addition, the immune response profile has been analyzed in COVID19 patients according to disease severity.
Results: In comparison to HDs and patients with NON-COV-RTI, COVID19 patients show a heterogeneous immune response with the presence of both activated and exhausted CD8+ T and NK cells characterised by the expression of the immune checkpoint LAG3 and the presence of the adaptive NK cell subset. An increased frequency of adaptive NK cells and a reduction of NK cells expressing the activating receptors NKp30 and NKp46 correlated with disease severity. Although both activated and exhausted NK cells expressing LAG3 were increased in moderate/severe cases, unsupervised cell clustering analyses revealed a more complex scenario with single NK cells expressing more than one immune checkpoint (PD1, TIM3 and/or LAG3). A general increased level of inflammatory cytokines and chemokines was found in COVID19 patients, some of which like IL18, IL1RA, IL36B and IL31, IL2, IFNα and TNFα, CXCL10, CCL2 and CCL8 were able to differentiate between COVID19 and NON-COV-RTI and correlated with bad prognosis (IL2, TNFα, IL1RA, CCL2, CXCL10 and CXCL9). Notably, we found that soluble NKG2D ligands from the MIC and ULBPs families were increased in COVID19 compared to NON-COV-RTI and correlated with disease severity.
Conclusions: Our results provide a detailed comprehensive analysis of the presence of activated and exhausted CD8+T, NK and monocyte cell subsets as well as extracellular inflammatory factors beyond cytokines/chemokines, specifically associated to COVID19. Importantly, multivariate analysis including clinical, demographical and immunological experimental variables have allowed us to reveal specific immune signatures to i) differentiate COVID19 from other infections and ii) predict disease severity and the risk of death.
Keywords: COVID19, GzmA, GzmB, NK cells, CXCL9, CXCL10, MIC, ULBP
 Global reach, higher impact
Global reach, higher impact