13.3
Impact Factor
Theranostics 2021; 11(19):9605-9622. doi:10.7150/thno.63763 This issue Cite
Research Paper
Visualization of endogenous p27 and Ki67 reveals the importance of a c-Myc-driven metabolic switch in promoting survival of quiescent cancer cells
1. School of Biomedical Sciences and Pharmacy, The University of Newcastle, NSW, 2308, Australia.
2. Translational Research Institute, Henan Provincial People's Hospital and People's Hospital of Zhengzhou University, Henan Provincial and Zhengzhou City Key laboratory of Long Non-coding RNA and Cancer Metabolism, Henan, 450053, China.
3. Centre for Excellence in Molecular Plant Sciences, Chinese Academy of Sciences, Shanghai, 200032, China.
4. Department of Pulmonary and Critical Care Medicine, Henan Provincial People's Hospital, Zhengzhou University People's Hospital, Henan 450003, China.
5. Biomedical Imaging Facility, University of New South Wales, NSW, 2052, Australia.
6. Department of Orthopaedics, John Hunter Hospital, Hunter New England Health, NSW, 2305, Australia.
7. Central Clinical School and Charles Perkins Centre, The University of Sydney, Sydney 2006, Australia.
8. Children's Cancer Institute Australia for Medical Research, University of New South Wales, NSW 2750, Australia.
9. Department of Nephrology, Henan Provincial People's Hospital, Zhengzhou University People's Hospital, Henan Provincial Clinical Research Canter for Kidney Disease, Henan 450003, China.
#These authors contributed equally to this work.
Abstract
Rationale: Recurrent and metastatic cancers often undergo a period of dormancy, which is closely associated with cellular quiescence, a state whereby cells exit the cell cycle and are reversibly arrested in G0 phase. Curative cancer treatment thus requires therapies that either sustain the dormant state of quiescent cancer cells, or preferentially, eliminate them. However, the mechanisms responsible for the survival of quiescent cancer cells remain obscure.
Methods: Dual genome-editing was carried out using a CRISPR/Cas9-based system to label endogenous p27 and Ki67 with the green and red fluorescent proteins EGFP and mCherry, respectively, in melanoma cells. Analysis of transcriptomes of isolated EGFP-p27highmCherry-Ki67low quiescent cells was conducted at bulk and single cell levels using RNA-sequencing. The extracellular acidification rate and oxygen consumption rate were measured to define metabolic phenotypes. SiRNA and inducible shRNA knockdown, chromatin immunoprecipitation and luciferase reporter assays were employed to elucidate mechanisms of the metabolic switch in quiescent cells.
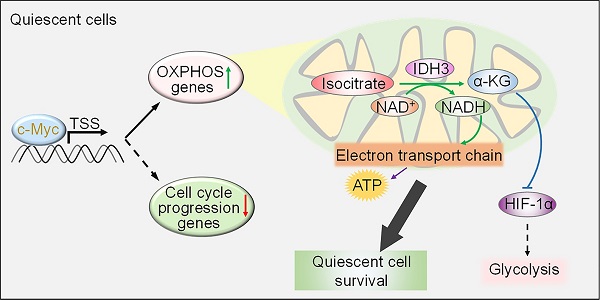
Results: Dual labelling of endogenous p27 and Ki67 with differentiable fluorescent probes allowed for visualization, isolation, and analysis of viable p27highKi67low quiescent cells. Paradoxically, the proto-oncoprotein c-Myc, which commonly drives malignant cell cycle progression, was expressed at relatively high levels in p27highKi67low quiescent cells and supported their survival through promoting mitochondrial oxidative phosphorylation (OXPHOS). In this context, c-Myc selectively transactivated genes encoding OXPHOS enzymes, including subunits of isocitric dehydrogenase 3 (IDH3), whereas its binding to cell cycle progression gene promoters was decreased in quiescent cells. Silencing of c-Myc or the catalytic subunit of IDH3, IDH3α, preferentially killed quiescent cells, recapitulating the effect of treatment with OXPHOS inhibitors.
Conclusion: These results establish a rigorous experimental system for investigating cellular quiescence, uncover the high selectivity of c-Myc in activating OXPHOS genes in quiescent cells, and propose OXPHOS targeting as a potential therapeutic avenue to counter cancer cells in quiescence.
Keywords: c-Myc, IDH3, quiescence, quiescent cells, oxidative phosphorylation
 Global reach, higher impact
Global reach, higher impact