Impact Factor
Theranostics 2021; 11(19):9519-9537. doi:10.7150/thno.65398 This issue Cite
Research Paper
Glycoproteome remodeling in MLL-rearranged B-cell precursor acute lymphoblastic leukemia
1. Institute for Glycomics, Griffith University, Gold Coast Campus, QLD, Australia.
2. Department of Systems Biology, Beckman Research Institute City of Hope, Monrovia, CA, USA.
3. Division of Hematology/Oncology and Bone Marrow Transplant, Children's Hospital Los Angeles, Los Angeles, CA, USA.
4. Glyco-Oncology, Ovarian Cancer Research, Department of Biomedicine, University Hospital Basel and University of Basel, Basel, Switzerland.
5. Department of Chemistry and Biomolecular Sciences, Macquarie University, Sydney, NSW, Australia.
6. ARC Centre of Excellence for Nanoscale BioPhotonics, Griffith University, QLD and Macquarie University, NSW, Australia.
#Present address: Emmanuelle Charpentier's Laboratory, Max Planck Unit for the Science of Pathogens, Charitéplatz 1 | D-10117 Berlin | Germany.
§Present address: Ionopticks, 68-70 Hanover Street, Fitzroy VIC 3065, Australia.
*Present address: Division of Immunobiology, Cincinnati Children's Hospital Medical Center, Cincinnati, OH 45229, USA.
Abstract
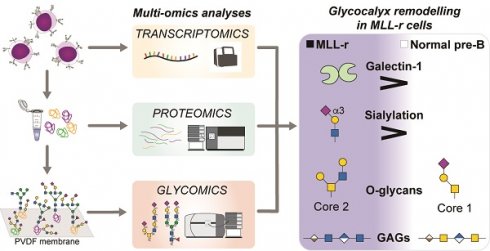
B-cell precursor acute lymphoblastic leukemia (BCP-ALL) with mixed-lineage leukemia gene rearrangement (MLL-r) is a poor-prognosis subtype for which additional therapeutic targets are urgently needed. Currently no multi-omics data set for primary MLL r patient cells exists that integrates transcriptomics, proteomics and glycomics to gain an inclusive picture of theranostic targets.
Methods: We have integrated transcriptomics, proteomics and glycomics to i) obtain the first inclusive picture of primary patient BCP-ALL cells and identify molecular signatures that distinguish leukemic from normal precursor B-cells and ii) better understand the benefits and limitations of the applied technologies to deliver deep molecular sequence data across major cellular biopolymers.
Results: MLL-r cells feature an extensive remodeling of their glycocalyx, with increased levels of Core 2-type O-glycans and complex N-glycans as well as significant changes in sialylation and fucosylation. Notably, glycosaminoglycan remodeling from chondroitin sulfate to heparan sulfate was observed. A survival screen, to determine if glycan remodeling enzymes are redundant, identified MGAT1 and NGLY1, essential components of the N-glycosylation/degradation pathway, as highly relevant within this in vitro screening. OGT and OGA, unique enzymes that regulate intracellular O-GlcNAcylation, were also indispensable. Transcriptomics and proteomics further identified Fes and GALNT7-mediated glycosylation as possible therapeutic targets. While there is overall good correlation between transcriptomics and proteomics data, we demonstrate that a systematic combined multi-omics approach delivers important diagnostic information that is missed when applying a single omics technology.
Conclusions: Apart from confirming well-known MLL-r BCP-ALL glycoprotein markers, our integrated multi-omics workflow discovered previously unidentified diagnostic/therapeutic protein targets.
Keywords: Leukemia, Glycomics, Proteomics, Transcriptomics, multi-omics
 Global reach, higher impact
Global reach, higher impact