13.3
Impact Factor
Theranostics 2021; 11(18):8945-8963. doi:10.7150/thno.61390 This issue Cite
Research Paper
The methods and advances of adaptive immune receptors repertoire sequencing
1. Hunan Key Laboratory of Biomedical Nanomaterials and Devices, Hunan University of Technology, Zhuzhou 412007, China.
2. State Key Laboratory of Bioelectronics, Southeast University, Nanjing 210096, China.
3. Department of Dermatology, Second Xiangya Hospital, Central South University, Hu-nan Key Laboratory of Medical Epigenomics, Changsha, Hunan, China.
4. Department of Microbiology and Immunology, Keio University School of Medicine, Tokyo, Japan.
#Authors contributed equally to this work.
Abstract
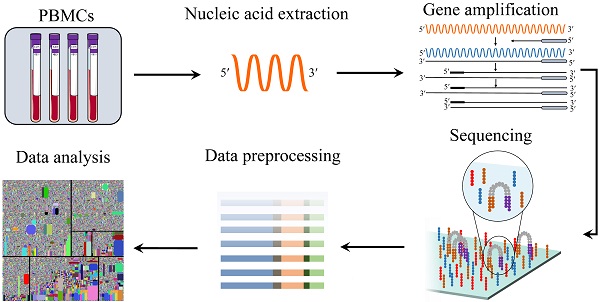
The adaptive immune response is a powerful tool, capable of recognizing, binding to, and neutralizing a vast number of internal and external threats via T or B lymphatic receptors with widespread sets of antigen specificities. The emergence of high-throughput sequencing technology and bioinformatics provides opportunities for research in the fields of life sciences and medicine. The analysis and annotation for immune repertoire data can reveal biologically meaningful information, including immune prediction, target antigens, and effective evaluation. Continuous improvements of the immunological repertoire sequencing methods and analysis tools will help to minimize the experimental and calculation errors and realize the immunological information to meet the clinical requirements. That said, the clinical application of adaptive immune repertoire sequencing requires appropriate experimental methods and standard analytical tools. At the population cell level, we can acquire the overview of cell groups, but the information about a single cell is not obtained accurately. The information that is ignored may be crucial for understanding the heterogeneity of each cell, gene expression and drug response. The combination of high-throughput sequencing and single-cell technology allows us to obtain single-cell information with low-cost and high-throughput. In this review, we summarized the current methods and progress in this area.
Keywords: adaptive immune repertoire sequencing, T or B cell receptors, high-throughput sequencing, bioinformatics
 Global reach, higher impact
Global reach, higher impact