13.3
Impact Factor
Theranostics 2021; 11(18):8674-8691. doi:10.7150/thno.60233 This issue Cite
Research Paper
Deficiency of the X-inactivation escaping gene KDM5C in clear cell renal cell carcinoma promotes tumorigenicity by reprogramming glycogen metabolism and inhibiting ferroptosis
1. Department of Medical Genetics, School of Basic Medical Sciences, Wuhan University, Wuhan 430071, China.
2. Department of Gynecology and Obstetrics, Zhongnan Hospital of Wuhan University, Wuhan 430072, China.
3. Department of Immunology, School of Basic Medical Sciences, Wuhan University, Wuhan 430071, China.
4. Hubei Provincial Key Laboratory of Allergy and Immunology, Wuhan 430071, China.
#These authors contributed equally to this paper.
Abstract
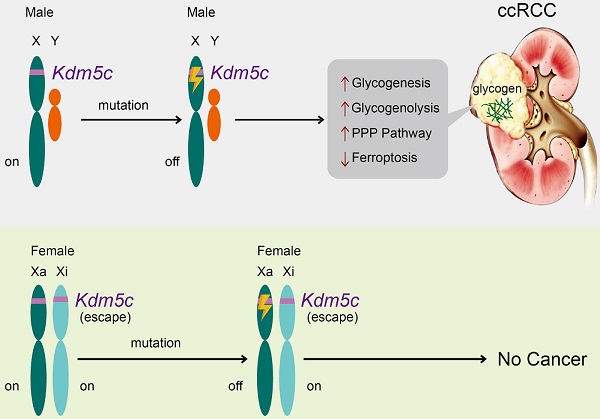
Background: Clear cell renal cell carcinoma (ccRCC) is characterized by glycogen-laden, unexplained male predominance, and frequent mutations in the Von Hippel-Lindau (VHL) gene and histone modifier genes. Besides, poor survival rates of ccRCC patients seem to be associated with up-regulation of the pentose phosphate pathway (PPP). However, the mechanism underlying these features remains unclear.
Methods: Whole exome sequencing was used to identify the gene mutation that implicated in the rewired glucose metabolism. RNA-seq analyses were performed to evaluate the function of KDM5C in ccRCC. Furthermore, heavy isotope tracer analysis and metabolites quantification assays were used to study how KDM5C affects intracellular metabolic flux. To provide more in vivo evidence, we generated the Kdm5c-/- mice by CRISPR-Cas9 mediated gene knockout and performed the xenografts with KDM5C overexpressing or depleted cell lines.
Results: A histone demethylase gene KDM5C, which can escape from X-inactivation and is predominantly mutated in male ccRCC patients, was identified to harbor the frameshift mutation in the ccRCC cell line with the highest glycogen level, while the restoration of KDM5C significantly reduced the glycogen level. Transcriptome and metabolomic analysis linked KDM5C to metabolism-related biological processes. KDM5C specifically regulated the expression of several hypoxia-inducible factor (HIF)-related genes and Glucose-6-phosphate dehydrogenase (G6PD) that were involved in glycogenesis/glycogenolysis and PPP, respectively, mainly through the histone demethylase activity of KDM5C. Depletion of KDM5C increased the production of glycogen, which was then directed to glycogenolysis to generate glucose-6-phosphate (G6P) and subsequently PPP to produce nicotinamide adenine dinucleotide phosphate hydride (NADPH) and glutathione (GSH), thus conferring cells resistance to reactive oxygen species (ROS) and ferroptosis. KDM5C re-expression suppressed the glucose flux through PPP and re-sensitized cancer cells to ferroptosis. Notably, Kdm5c-knockout mice kidney tissues exhibited elevated glycogen level, reduced lipid peroxidation and displayed a transformation of renal cysts into hyperplastic lesions, implying a cancer-protective benefit of ferroptosis. Furthermore, KDM5C deficiency predicted the poor prognosis, and clinically relevant KDM5C mutants failed to suppress glycogen accumulation and promoted ferroptosis as wild type.
Conclusion: This work revealed that a histone modifier gene inactive mutation reprogramed glycogen metabolism and helped to explain the long-standing puzzle of male predominance in human cancer. In addition, our findings may suggest the therapeutic value of targeting glycogen metabolism in ccRCC.
Keywords: KDM5C, glycogen, male predominance, X-inactivation escaping gene, pentose phosphate pathway
 Global reach, higher impact
Global reach, higher impact