Impact Factor
Theranostics 2018; 8(20):5772-5783. doi:10.7150/thno.28949 This issue Cite
Research Paper
An Aptamer-Based Probe for Molecular Subtyping of Breast Cancer
1. State Key Laboratory of Bioelectronics, National Demonstration Center for Experimental Biomedical Engineering Education (Southeast University), School of Biological Science and Medical Engineering, Southeast University, Nanjing 210096, P. R. China
2. School of Chemistry and Chemical Engineering, Southeast University, Nanjing 211189, P. R. China
3. Department of Pathology, Third Xiangya Hospital, Central South University, Changsha 410013, P. R. China
4. The Medical School, Ningbo University, Ningbo 315211, P. R. China
5. Economical Forest Cultivation and Utilization of 2011 Collaborative Innovation Center in Hunan Province, Hunan Key Laboratory of Biomedical Nanomaterials and Devices, Hunan University of Technology, Zhuzhou 412007, P. R. China
6. Department of Medical Imaging, Jinling Hospital, School of Medicine, Nanjing University, Nanjing 210002, P.R. China
7. State Key Laboratory of Analytical Chemistry for Life Science, School of Chemistry and Chemical Engineering, Nanjing University, Nanjing 210093, P.R. China
*These three authors contributed equally to this work.
Abstract
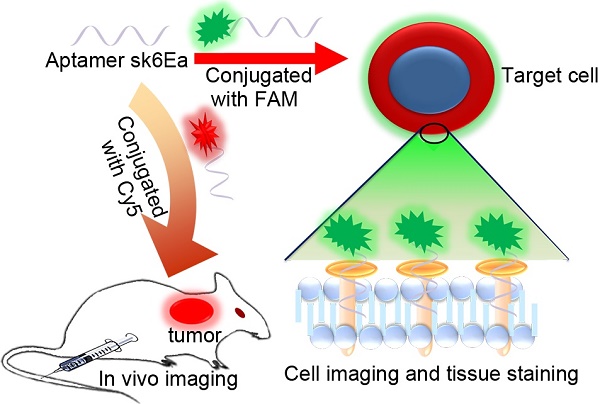
Molecular subtyping of breast cancer is of considerable interest owing to its potential for personalized therapy and prognosis. However, current methodologies cannot be used for precise subtyping, thereby posing a challenge in clinical practice. The aim of the present study is to develop a cell-specific single-stranded DNA (ssDNA) aptamer-based fluorescence probe for molecular subtyping of breast cancer.
Methods: Cell-SELEX method was utilized to select DNA aptamers. Flow cytometry and confocal microscopy were used to study the specificity, binding affinity, temperature effect on the binding ability and target type analysis of the aptamers. In vitro and in vivo fluorescence imaging were used to distinguish the molecular subtypes of breast cancer cells, tissue sections and tumor-bearing mice.
Results: Six SK-BR-3 breast cancer cell-specific ssDNA aptamers were evolved after successive in vitro selection over 21 rounds by Cell-SELEX. The Kd values of the selected aptamers were all in the low-nanomolar range, among which aptamer sk6 showed the lowest Kd of 0.61 ± 0.14 nM. Then, a truncated aptamer-based probe, sk6Ea, with only 53 nt and high specificity and binding affinity to the target cells was obtained. This aptamer-based probe was able to 1) differentiate SK-BR-3, MDA-MB-231, and MCF-7 breast cancer cells, as well as distinguish breast cancer cells from MCF-10A normal human mammary epithelial cells; 2) distinguish HER2-enriched breast cancer tissues from Luminal A, Luminal B, triple-negative breast cancer tissues, and adjacent normal breast tissues (ANBTs) in vitro; and 3) distinguish xenografts of SK-BR-3 tumor-bearing mice from those of MDA-MB-231 and MCF-7 tumor-bearing mice within 30 min in vivo.
Conclusion: The results suggest that the aptamer-based probe is a powerful tool for fast and highly sensitive subtyping of breast cancer both in vitro and in vivo and is also very promising for the identification, diagnosis, and targeted therapy of breast cancer molecular subtypes.
Keywords: aptamer, cell-SELEX, molecular subtype, breast cancer, imaging
 Global reach, higher impact
Global reach, higher impact