13.3
Impact Factor
Theranostics 2022; 12(8):3818-3833. doi:10.7150/thno.71833 This issue Cite
Research Paper
Single-cell landscape reveals active cell subtypes and their interaction in the tumor microenvironment of gastric cancer
1. Lanzhou University Second Hospital, Lanzhou 730030, China.
2. BIOPIC, Beijing Advanced Innovation Center for Genomics, School of Life Sciences, Peking University, Beijing 100871, China.
3. Clabee Genomics, Urban Garden Building, Bookstore Road, Luohu District, Shenzhen 518000, Guangdong, China.
4. Department of Genetics, Harvard Medical School, Boston, Massachusetts 02115, USA.
5. Department of Molecular and Cellular Biology, Harvard University, Cambridge, Massachusetts 02138, United States.
6. National Cancer Center/National Clinical Research Center for Cancer/Cancer Hospital, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing 100021, China.
7. Center on Translational Neuroscience, College of Life and Environmental Sciences, Minzu University of China, Beijing 100081, China.
8. Wyss Institute for Biologically Inspired Engineering, Harvard University, Boston, Massachusetts 02115, USA.
#These authors contributed equally to this work.
Abstract
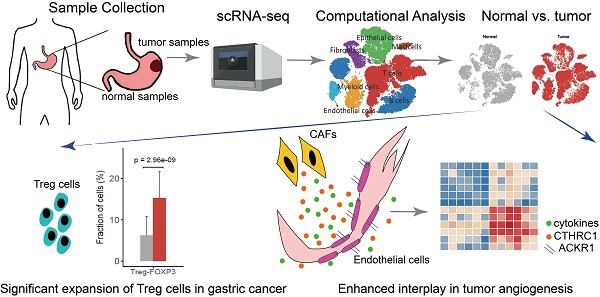
Background: Gastric cancer remains the third most common cause of cancer-related death worldwide. The development of novel therapeutic strategies for gastric cancer requires a deep understanding of the tumor cells and microenvironment of gastric cancer.
Methods: We performed the single-cell RNA sequencing (scRNA-seq) on nine untreated non-metastatic gastric cancer patients. The transcriptomic atlas and ligand-receptor-based intercellular communication networks of the single cells were characterized.
Results: Here, we profiled the transcriptomes of 47,304 cells from nine patients with gastric cancer. Tregs cells were significantly enriched in the gastric tumor tissues with increased expression of immune suppression related genes, which suggest a more immunosuppressive microenvironment. We also observed the absence of separate exhausted CD8+ T cell cluster, and the low expression level of exhaustion markers PDCD1, CTLA4, HAVCR2, LAG-3, and TIGIT in those specific cells. These may serve as molecular-level evidence for the limited benefit of immunotherapy among gastric cancer patients. In addition, we found ACKR1 specifically expressed in tumor endothelial cells, associated with poor prognosis in the cohort data and potentially provided a novel target of gastric cancer treatment. Furthermore, the tight interaction between endothelial cells and fibroblast implied the important roles of fibroblast in tumor angiogenesis and the maintenance of tumor vasculature.
Conclusions: In conclusion, this single-cell atlas provide understanding the cellular heterogeneity from molecular level in gastric cancer and will serve as a valuable resource for developing innovative early and companion diagnostics, as well as discovering novel targeted therapies for gastric cancer.
Keywords: Gastric cancer, tumor microenvironment, single-cell RNA sequencing.
 Global reach, higher impact
Global reach, higher impact