Impact Factor
Theranostics 2022; 12(6):2948-2962. doi:10.7150/thno.70373 This issue Cite
Research Paper
Assay design for unambiguous identification and quantification of circulating pathogen-derived peptide biomarkers
1. Center for Cellular and Molecular Diagnostics, Department of Biochemistry and Molecular Biology, School of Medicine, Tulane University, New Orleans, Louisiana, USA.
2. Sichuan Provincial Key Laboratory for Human Disease Gene Study, Department of Medical Genetics, Department of Laboratory medicine, Sichuan Academy of Medical Sciences & Sichuan Provincial People's Hospital, Chengdu, China.
3. Translational Research Unit, National Institute for Infectious Diseases Lazzaro Spallanzani-IRCCS, Rome, Italy.
4. Departments of Medicine, Division of Allergy and Infectious Diseases, and Global Health, University of Washington, Seattle, USA.
5. Southwest National Primate Research Center, Texas Biomedical Research Institute, San Antonio, Texas, USA.
6. Department of Pediatrics and Child Health, University of Nairobi, Nairobi, Kenya.
*these authors are co-first authors and contribute equally to this work.
Abstract
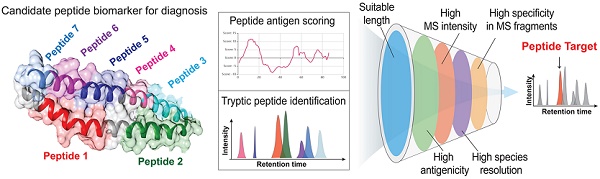
Rationale: Circulating pathogen-derived proteins can serve as useful biomarkers for infections but may be detected with poor sensitivity and specificity by standard immunoassays due to masking effects and cross-reactivity. Mass spectrometry (MS)-read immunoassays for biomarker-derived peptides can resolve these issues, but lack standard workflows to select species-specific peptides with strong MS signal that are suitable for antibody generation.
Methods:Using a Mycobacterium tuberculosis (Mtb) protein as an example, candidate peptides were selected by length, species-specificity, MS intensity, and antigenicity score. MS data from spiked healthy serum was employed to define MS feature thresholds, including a novel measure of internal MS data correlation, to produce a peak detection algorithm.
Results: This algorithm performed better in rejecting false positive signal than each of its criteria, including those currently employed for this purpose. Analysis of an Mtb peptide biomarker (CFP-10pep) by this approach identified tuberculosis cases not detected by microbiologic assays, including extrapulmonary tuberculosis and tuberculosis cases in children infected with HIV-1. Circulating CFP-10pep levels measured in a non-human primate model of tuberculosis distinguished disease from asymptomatic infection and tended to correspond with Mtb granuloma size, suggesting that it could also serve as a surrogate marker for Mtb burden and possibly treatment response.
Conclusions: These biomarker selection and analysis approach appears to have strong potential utility for infectious disease diagnosis, including cryptic infections, and possibly to monitor changes in Mtb burden that may reflect disease progression or a response to treatment, which are critical needs for more effective disease control.
Keywords: MRM, peptide biomarker, immunoprecipitation, mass spectrometry, tuberculosis
 Global reach, higher impact
Global reach, higher impact