13.3
Impact Factor
Theranostics 2021; 11(10):5010-5027. doi:10.7150/thno.55645 This issue Cite
Research Paper
Deciphering the autophagy regulatory network via single-cell transcriptome analysis reveals a requirement for autophagy homeostasis in spermatogenesis
1. State Key Laboratory of Organ Failure Research, Department of Developmental Biology, School of Basic Medical Sciences, Southern Medical University, Guangzhou, Guangdong, 510515, P. R. China.
2. Department of Biochemistry and Molecular Biology, Shenzhen University Health Science Center, Shenzhen, Guangdong, 518060, P. R. China.
3. Department of Bioinformatics, School of Basic Medical Sciences, Southern Medical University, Guangzhou, Guangdong, 510515, P. R. China.
4. Department of Health Management, Nanfang Hospital, Southern Medical University, Guangzhou, Guangdong, 510515, P. R. China.
5. Guangdong Key Laboratory of Construction and Detection in Tissue Engineering, Southern Medical University, Guangzhou, Guangdong, 510515, P. R. China.
6. Zhuhai Precision Medical Center, Zhuhai People's Hospital (Zhuhai Hospital affiliated with Jinan University), Zhuhai, Guangdong, 519000, P. R. China.
7. Reproductive Medicine Center of The Third Affiliated Hospital of Guangzhou Medical University, Guangzhou 510150, P. R. China.
8. Department of Thoracic Surgery, Nanfang Hospital, Southern Medical University, Guangzhou, Guangdong, 510515, P. R. China.
9. Cancer Center, Faculty of Health Sciences, University of Macau, Macau SAR, China.
10. Bioland Laboratory (Gangzhou Regenerative Medicine and Health Guangdong Laboratory), Guangzhou, Guangdong, 510005, P. R. China.
11. Department of Gynecology, Zhujiang Hospital, Southern Medical University, Guangzhou, Guangdong, 510280, P. R. China.
12. China National Center for Bioinformation Key Laboratory of Genomic and Precision Medicine, Beijing Institute of Genomics, Chinese Academy of Sciences, Beijing 100101, P. R. China.
* These authors contributed equally to this work.
Abstract
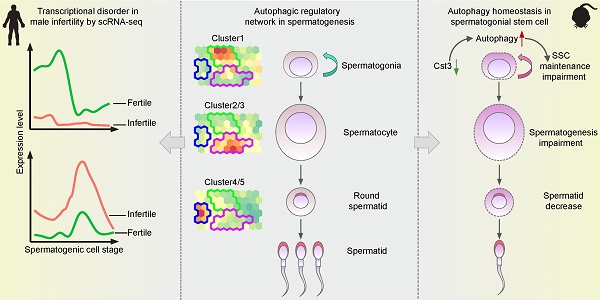
Background: Autophagy has been implicated as a crucial component in spermatogenesis, and autophagy dysfunction can lead to reproductive disorders in animal models, including yeast, C. elegans and mice. However, the sophisticated transcriptional networks of autophagic genes throughout human spermatogenesis and their biological significance remain largely uncharacterized.
Methods: We profiled the transcriptional signatures of autophagy-related genes during human spermatogenesis by assessing specimens from nine fertile controls (including two normal persons and seven obstructive azoospermia (OA) patients) and one nonobstructive azoospermia (NOA) patient using single-cell RNA sequencing (scRNA-seq) analysis. Dysregulation of autophagy was confirmed in two additional NOA patients by immunofluorescence staining. Gene knockdown was used to identify the role of Cst3 in autophagy during spermatogenesis.
Results: Our data uncovered a unique, global stage-specific enrichment of autophagy-related genes. Human-mouse comparison analysis revealed that the stage-specific expression pattern of autophagy-related genes was highly conserved in mammals. More importantly, dysregulation of some clusters of autophagy-related genes was observed in NOA patients, suggesting the association of autophagy with male infertility. Cst3, a human-mouse conserved and autophagy-related gene that is actively expressed in spermatogonia and early spermatocytes, was found to regulate spermatogonial stem cell (SSC) maintenance and subsequent male germ cell development. Knockdown of Cst3 increased autophagic activity in mouse SSCs and subsequently suppressed the transcription of SSC core factors such as Oct4, Id1, and Nanos3, which could be efficiently rescued by manipulating autophagic activity.
Conclusions: Our study provides comprehensive insights into the global transcriptional signatures of autophagy-related genes and confirms the importance of autophagy homeostasis in SSC maintenance and normal spermatogenesis, opening new avenues for further dissecting the significance of the autophagy regulatory network in spermatogenesis as well as male infertility.
Keywords: spermatogenesis, autophagy, single-cell RNA sequencing, spermatogonial stem cells, meiosis, male infertility.
 Global reach, higher impact
Global reach, higher impact