Impact Factor
Theranostics 2020; 10(23):10360-10377. doi:10.7150/thno.49922 This issue Cite
Review
Managing therapeutic resistance in breast cancer: from the lncRNAs perspective
1. State Key Laboratory of Biotherapy and Cancer Center, West China Hospital, and West China School of Basic Medical Sciences & Forensic Medicine, Sichuan University, and Collaborative Innovation Center for Biotherapy, Chengdu, 610041, P.R. China.
2. Department of Urology, Institute of Urology, West China Hospital, Sichuan University, Chengdu, 610041, P.R. China.
3. Department of Biochemistry and Molecular Biology, Monash University, Clayton, Victoria, Australia.
4. West China School of Basic Medical Sciences & Forensic Medicine, Sichuan University, Chengdu, 610041, P.R. China.
#These authors contributed equally to this work.
Received 2020-6-25; Accepted 2020-8-4; Published 2020-8-18
Abstract
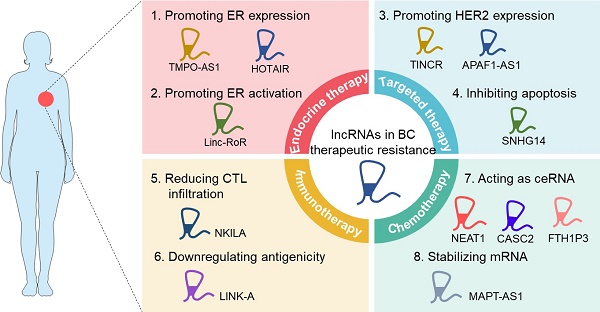
Breast cancer (BC) is the most common female malignancy and the second leading cause of cancer-related death worldwide. In spite of significant advances in clinical management, the mortality of BC continues to increase due to the frequent occurrence of treatment resistance. Intensive studies have been conducted to elucidate the molecular mechanisms underlying BC therapeutic resistance, including increased drug efflux, altered drug targets, activated bypass signaling pathways, maintenance of cancer stemness, and deregulated immune response. Emerging evidence suggests that long noncoding RNAs (lncRNAs) are intimately involved in BC therapy resistance through multiple modes of action. Therefore, an in-depth understanding of the implication of lncRNAs in resistance to clinical therapies may improve the clinical outcome of BC patients. Here, we highlight the role and underlying mechanisms of lncRNAs in regulating BC treatment resistance with an emphasis on lncRNAs-mediated resistance in different clinical scenarios, and discuss the potential of lncRNAs as novel biomarkers or therapeutic targets to improve BC therapy response.
Keywords: Long noncoding RNA, Breast cancer, Drug resistance, Biomarkers, Therapeutic targets
Introduction
Breast cancer (BC) is one of the three most frequent cancers worldwide and also the most common cancer in females, with approximately one in ten women at risk of suffering it during their lifetime [1]. Moreover, BC is the leading cause of cancer-related death in less developed countries and ranks second in more developed countries [1]. Currently, the mainstay treatments of BC involve endocrine therapy, anti-human epidermal receptor 2 (HER2) target therapy and cytotoxic chemotherapy depending on individual BC subtypes. Based on emerging preclinical and clinical studies, combinational treatment of targeted drugs (e.g. mTOR inhibitor everolimus [2], cyclin-dependent kinase 4/6 (CDK 4/6) inhibitor palbociclib [3]) with endocrine therapy has prolonged progression-free survival (PFS) in certain BC patients. These therapeutic strategies along with early screening tools (e.g. mammography [4], tomosynthesis [5] and magnetic resonance imaging (MRI) [6]) have partly decreased BC mortality; however, the clinical therapeutic outcome is far from satisfied. The major barrier for complete BC cure is the development of therapeutic tolerance. BC cells are equipped with numerous mechanisms to cope with different treatment strategies, such as increased drug efflux, altered drug targets and activated bypass signaling pathways, maintenance of cancer stemness, as well as deregulated immune response. Therefore, there is still an urgent need for extensive research on BC therapy resistance to develop novel biomarkers and therapeutic targets that could predict therapeutic response or improve clinical outcomes of BC patients.
Basic research on BC therapeutic resistance has focused more on protein-coding genes as their products are thought to play a central role in regulating biological activities. However, only about 2% of the human transcriptome encodes proteins [7], while the remainder of the transcriptome has no obvious protein-coding potential (referred to as noncoding RNAs, ncRNAs). The past few years have witnessed exciting advances in the functional and mechanistic characterization of ncRNAs, especially long noncoding RNAs (lncRNAs) which contain the largest percentage of the noncoding transcriptome [8]. Since the identification of the first lncRNA H19 in fetal liver tissue in 1900 [9], thousands of lncRNAs have been found and investigated. Notably, an increasing number of lncRNAs have been found to be associated with pathological scenarios such as neurodegenerative disorders, cardiovascular diseases and cancers. For example, several lncRNAs identified in HOX gene foci have been shown to play essential roles in cancer initiation and development [10].
Although the specific functions and exact underlying mechanisms of most BC-related lncRNAs remain elusive, there have been increasing studies on dysregulated lncRNAs in BC therapeutic resistance (Table 1). For example, upregulated expression of lncRNA HOTAIR and nuclear paraspeckle assembly transcript 1 (NEAT1) are responsible for BC therapeutic resistance to endocrine therapy and chemotherapy, respectively [11-13]. In particular, Cantile et al. have recently published a comprehensive review on the role of HOTAIR in BC therapy, which introduces numerous recent and influential studies [14]. Furthermore, growing evidence has indicated the great potential of lncRNAs to serve as biomarkers to predict treatment response, such as HOXB-AS5 [15], Lnc712 [16] and the long intergenic non-coding RNA for kinase activation (LINK-A) [17]. LncRNAs can also be applied as therapeutic targets to help tackle treatment resistance in BC. For example, therapeutic delivery of locked nucleic acids (LNAs) targeting LINK-A has been proven to confer BC sensitivity to immune checkpoint inhibitors in a preclinical study [17].
Mechanisms of BC therapeutic resistance mediated by lncRNAs
| LncRNA | Therapeutic strategy | Pathway/target | Action modes | Effection | Refs |
|---|---|---|---|---|---|
| HOTAIR | Tamoxifen | ER | Promoting ligand-independent ER activities, increasing cancer stemness | Inducing | [11] |
| NEAT1 | Paclitaxel , cisplatin, 5-FU | miR-129/ZEB2, miR-211/HMGA2 | CeRNA, regulating apoptosis and cell cycle progression, facilitating cell growth | Inducing | [12, 13] |
| HOXB-AS5 | PI3K/AKT/mTOR inhibitors | PI3K/AKT/mTOR | Promoting cell growth, migration and invasion | Inducing | [15] |
| Lnc712 | CDK inhibitors | HSP90 | Regulating CDK2 activation and triggering cell proliferation | Inducing | [16] |
| LINK-A | Immune checkpoint blockers | PIP3/GPCR, PLC | Reducing antigenicity to avoid detection by antitumor lymphocytes | Inducing | [17] |
| H19 | Tamoxifen, fulverstrant | ER | Regulating ERα expression at the transcript and protein levels | Inducing | [19] |
| MIR2052HG | Aromatase inhibitors | EGR1, ER | Promoting ESR1 transcription and limiting ubiquitin-mediated ERα degradation | Inducing | [20] |
| TINCR | Trastuzumab | miR-125b/ERBB2 | CeRNA, regulating the expression level of HER2 | Inducing | [21] |
| Linc-RoR | Tamoxifen | DUSP7, MAPK/ERK | Promoting estrogen-independent cell growth | Inducing | [25] |
| DCST1-AS1 | Doxorubicin, paclitaxel | ANXA1 | Unknown | Inducing | [37] |
| NKILA | Immunotherapy | NF-κB | Facilitating T cell vulnerability to AICD and decreasing CTL infiltration | Inducing | [52] |
| TMPO-AS1 | Endocrine therapy | ER | Stabilizing ESR1 mRNA through interaction with ESR1 mRNA | Inducing | [61] |
| LINP1 | Tamoxifen | ER | Attenuating the estrogen response | Inducing | [62] |
| DSCAM-AS1 | Tamoxifen | hnRNPL | Unknown | Inducing | [63] |
| GAS5 | Tamoxifen | miR-222/PTEN | CeRNA | Reversing | [64] |
| UCA1 | Tamoxifen | miR-18a/HIF1α | CeRNA, regulating cell cycle | Inducing | [65] |
| CYTOR | Tamoxifen | miR‑125a‑5p/SRF, Hippo, MAPK | CeRNA, promoting cell survival | Inducing | [66] |
| DSCAM‐AS1 | Tamoxifen | miR‐137/EPS8 | CeRNA, promoting cell proliferation and suppressing apoptosis | Inducing | [67] |
| HOTAIRM1 | Tamoxifen | EZH2 | Preventing H3K27me3 of HOXA1 | Inducing | [69] |
| AFAP1-AS1 | Trastuzumab | AUF1/ERBB2 | Enhancing HER2 translation, exosome-mediated dissemination | Inducing | [75] |
| AGAP2-AS1 | Trastuzumab | hnRNPA2B1 | Exosome-mediated dissemination | Inducing | [76] |
| SNHG14 | Trastuzumab | Bcl-2/Bax, PABPC1 | Inhibiting apoptosis, exosome-mediated dissemination | Inducing | [77, 78] |
| AGAP2-AS1 | Trastuzumab | CBP, MyD88, NF-κB | Activating NF-κB signaling pathway, promoting cell growth | Inducing | [80] |
| LINK-A | MK2206 | AKT | Facilitating the enzymatic activation of AKT | Inducing | [89] |
| AK023948 | AKT inhibitors | DHX9/p85 | Sustaining the stability of p85 | Inducing | [90] |
| Linc-ROR | mTOR inhibitor (rapamycin) | miR-194-3p/ MECP2 | CeRNA | Inducing | [91] |
| lncRNA-JADE | PARP inhibitors | BRCA1, Jade1 | Increasing transcription of DNA damage repair-related genes | Inducing | [110] |
| GUARDIN | PARP inhibitors | BRCA1, TRF2 | Maintaining genome integrity | Inducing | [115] |
| PHACTR2-AS1 | PARP inhibitors | Ribosome DNA genes | Triggering H3K9 methylation-mediated silencing of ribosome DNA genes | Inducing | [117] |
| FTH1P3 | Paclitaxel | miR-206/ABCB1 | CeRNA | Inducing | [124] |
| Linc00518 | Paclitaxel | miR-199a/MRP1 | CeRNA | Inducing | [125] |
| NONHSAT101069 | Epirubicin | miR-129-5p/Twist1 | CeRNA | Inducing | [126] |
| CASC2 | Paclitaxel | miR-18a-5p/CDK19 | CeRNA | Inducing | [127] |
| MAPT-AS1 | Paclitaxel | MAPT | Increasing the stability of MAPT mRNA | Inducing | [128] |
| NONHSAT141924 | Paclitaxel | p-CREB/Bcl-2 apoptosis pathway | Unknown | Inducing | [129] |
| LINC00968 | Paclitaxel, adriamycin | WNT2 | Inhibiting the Wnt2/β-catenin signaling pathway | Reversing | [132] |
| GAS5 | Adriamycin | miR-221-3p/DDK2 | CeRNA | Reversing | [133] |
| AC073284.4 | Paclitaxel | miR‐18b‐5p/DOCK4 | CeRNA | Reversing | [134] |
In this review, we will discuss biological functions and underlying mechanisms of dysregulated lncRNAs in BC therapy resistance with an emphasis on lncRNAs-mediated therapeutic resistance in BC clinical scenarios. Additionally, we also highlight the advantages and challenges lying ahead for the application of lncRNAs as biomarkers or targets for restraining treatment resistance in BC.
LncRNAs-mediated BC therapeutic resistance
The mechanisms of resistance to different BC therapeutic strategies share many similarities, including increased drug efflux, altered drug targets, activated bypass signaling pathways, as well as maintenance of cancer stemness. Besides, deregulated immune response has also been identified as an essential contributor to BC immunotherapy resistance. Emerging evidence has demonstrated that lncRNAs participate in therapeutic resistance of BC through multiple modes of action. As the role of lncRNAs in increased drug efflux has been comprehensively reviewed elsewhere [18], here we focus on the molecular mechanisms of lncRNAs-mediated BC therapeutic resistance including the alteration of drug targets, downstream bypass pathways, cancer stemness, and immune response.
LncRNAs in drug targets and bypass signaling alteration
Drug efficacy is largely determined by the protein level and mutation state of drug targets, as well as the activation of bypass signaling pathways. One example is the altered expression of estrogen receptor α (ERα), which is a target of endocrine therapy for BC. It has been reported that Notch and HGF signaling-mediated upregulation of lncRNA H19 could promote ERα expression at both mRNA and protein levels. Thus, H19 counteracts endocrine therapy-mediated downregulation of ERα protein and is responsible for therapeutic resistance in BC cells [19]. Similarly, lncRNA MIR2052HG increases the transcription of the ERα encoding gene ESR1 and reduces ERα degradation through LMTK3, thus resulting in resistance to endocrine therapy [20]. LncRNA TINCR upregulated in BC cells sponges miR-125b to increase the expression level of HER2, resulting in the resistance of BC cells to anti-HER2 targeted therapy [21].
In addition to drug target alteration, lncRNAs are implicated in activation of bypass signaling pathways to mediate BC therapeutic tolerance. A typical example is the ligand-independent ERα activation that impedes the efficacy of endocrine agents. ERα can be phosphorylated by MAPK and AKT independent of the ligand binding, leading to endocrine resistance [22-24]. According to a recent study, linc-RoR promotes ligand-independent cell growth through activating MAPK/ERK pathway and results in BC endocrine resistance [25]. Taken together, these findings indicate that lncRNAs play essential roles in modulating therapeutic resistance by bypassing the original drug targets. Further investigations are required to unmask the lncRNAs-associated redundant pro-resistant signaling pathways involved in BC therapeutic strategies. Targeting fundamental lncRNAs in these redundant pro-resistant signaling pathways may exhibit promising effects in clinical situations.
LncRNAs in cancer stemness maintenance
CSCs are considered as a self-renewing subpopulation of neoplastic cells among heterogeneous tumors and were first documented in 1997 in the hematopoietic system [26]. Later in 2003, BC became the first solid tumor in which CSCs were discovered. The existence of breast cancer stem cells (BCSCs) poses a tremendous challenge for BC treatment due to their inherent therapeutic resistance to conventional therapies. Therefore, more in-depth research is required to explore the regulatory networks of BCSCs formation and maintenance.
A recent study reveals that lncRNA Peblr20 can enhance pluripotent reprogramming thus maintaining pluripotency of induced pluripotent stem cells (iPSCs) [27], implicating that lncRNAs may also orchestrate the preservation of cancer stemness. Previous studies have illustrated a direct nexus between epithelial-mesenchymal transition (EMT) and cancer stemness, especially in BC [28-29]. Emerging studies have revealed that lncRNAs are vital novel players in the regulation of EMT-associated BCSC stemness [30-31]. For instance, several lncRNAs have been demonstrated to maintain or enhance both EMT traits and CSC-like characteristics of BC cells, including LINC01638 [32], lncRNA RP1 [33], LINC-ZNF469-3 [34] and LINC-ROR [35]. A recent study provides a more detailed possible mechanism through which lncRNA-Hh upregulated by Twist directly targets the hedgehog signaling (Hh) enhancer GAS1, to activate Hh and increase the expression of Gli1, SOX2 and OCT4 for BCSC maintenance [36]. The Twist-lncRNA-Hh-Hh-SOX2/OCT4 axis partly explains why epithelial BC cells with EMT phenotype also gain CSC-like properties. Recently, Tang et al. established the direct nexus between lncRNA-regulated EMT and resistance to BC therapy. According to their study, DCST1-AS1 enhances EMT and promotes TNBC chemoresistance to doxorubicin and paclitaxel by directly interacting with ANXA1 [37]. However, the underlying mechanism of DCST1-AS1-ANXA1 axis-mediated doxorubicin resistance remains to be further elucidated.
Furthermore, it has been well documented that some of the pluripotency factors (e.g. OCT3/4, SOX2, KLF4, LIN28) and CSC markers (e.g. ALDH1A3) [38-45] are capable of promoting stemness in BCSCs. Increasing evidence has implicated the pivotal role of lncRNAs in BCSCs maintenance through their interplay with these stemness-associated factors. Recently, Xu and colleagues have demonstrated that lncRNA CCAT2 enhances the expression of OCT4, Nanog and KLF4, as well as increases the ALDH+ CSC subpopulation in TNBC [46]. In addition, a number of lncRNAs have been found to act as competing endogenous RNAs (ceRNAs), which compete against the limited microRNAs (miRNAs) pool, to regulate the expression of pluripotency factors and CSC markers. For example, highly expressed lncRNA H19 in BCSCs acts as a ceRNA to sponge miRNA Let-7, resulting in the increased expression of a Let-7 target LIN28 thus promoting the preservation of BCSCs [47]. Intriguingly, LIN28 reversely inhibits Let-7 production and maturation, further de-repressing H19 expression in BCSCs [47]. The positive feedback loop formed by H19, Let-7 and LIN28 maintains the stemness of BCSCs, indicating that the disruption of this axis may provide opportunities for reversal of treatment tolerance. Likewise, mesenchymal stem cells (MSCs)-induced lncRNA LINC01133 positively regulates KLF4 to promote phenotypic and growth characteristics of BCSCs [48]. Moreover, lncRNA-Hh promotes the activation of the hedgehog signaling molecule Hh to upregulate SOX2 and OCT4 for BCSC maintenance [36]. The lncRNA HOTTIP regulates the miR-148a-3p/WNT1 pathway to maintain the CSC-like properties of BCSCs and facilitate BC growth [49]. Notably, NRAD1 was identified as the first lncRNA which can be activated by a CSC marker [50]. Mechanistically, ALDH1A3 and its product retinoic acid positively regulate the expression of NRAD1, thus enhancing the interaction between NRAD1 and genes involved in differentiation and catabolism, eventually promoting cell survival and increasing the number of BCSCs [50]. Taken together, lncRNAs are widely involved in BCSCs preservation and may lead to intrinsic therapeutic tolerance, however, the underlying mechanisms and clinical value remain to be thoroughly explored.
LncRNAs in immune response deregulation
Cancer immunotherapy is an emerging treatment option taking advantage of the cytotoxic potential of the immune system. In spite of the encouraging progress in BC immunotherapy, cancer cells have been reported to develop numerous mechanisms to evade immune elimination, including reduced tumor antigenicity, increased activation-induced cell death (AICD) of T lymphocytes and re-activation of oncogenic signaling [51]. For example, lncRNA LINK-A plays a central role in antigenicity loss and immune checkpoints evasion in BC through directly interacting with phosphatidylinositol-(3,4,5)-trisphosphate and inhibitory G-protein-coupled receptor (GPCR). Such interactions lead to reduced cyclic AMP (cAMP) concentrations and subsequent protein kinase A (PKA)-mediated TRIM71 phosphorylation. Consequently, phosphorylated-TRIM71 enhances proteasome-mediated degradation of peptide-loading complex (PLC) components, thus resulting in decreased antigen presentation to the surface of BC cells [17]. Another independent study has revealed that lncRNA NKILA enhances T cell vulnerability to AICD by interacting with NF-κB. Thus, the apoptosis and subsequent reduced infiltration of cytotoxic T lymphocytes (CTLs) might contribute to immunotherapy resistance [52]. Overall, lncRNAs are intimately related to deregulated immune response, thus conferring resistance to immunotherapy in cancer cells. Targeting lncRNAs may present a promising strategy to reverse therapeutic resistance and achieve better clinical outcome for BC patients.
LncRNAs-mediated BC resistance in different clinical scenarios
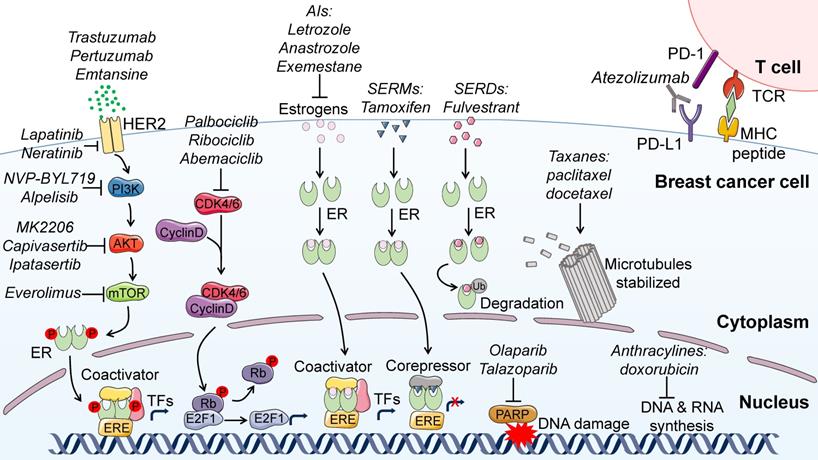
Based on the expression of specific biomarkers including ER, progesterone receptor (PR) and HER2, BC has been classified into at least four clinically relevant subtypes, including luminal A, luminal B, HER2-enriched, and basal like subtype. The two luminal subtypes are commonly characterized with positive ER or PR (or both) expression and negative HER2 expression, among which luminal A subtype is characterized with less proliferative potential. HER2 subtype BC is characterized by the overexpression of HER2, which can be subdivided into non-luminal (ER and PR negative) and luminal (ER or PR positive, or both) subtype. The basal-like subtype BC shows positive basal marker expression and usually negative expression of ER, PR and HER2, thus most of basal-like subtype BC is triple-negative breast cancer (TNBC) [1]. Among the four main subtypes, the luminal A subtype shows favorable prognosis, while the basal-like subtype exhibits the most aggressive clinical behavior. Based on these different clinical subtypes of BC, endocrine therapy, anti-HER2 targeted therapy and chemotherapy constitute the backbone of BC treatment (Figure 1). These clinical mainstay strategies, along with immunotherapy and targeted therapies beyond HER2, have, to a large extent, improved the PFS of BC patients. However, resistance inevitably occurs due to multifaceted factors and thus impedes therapeutic efficacy. Here, we highlight the mechanisms responsible for treatment resistance to each BC therapeutic strategy in the perspective of lncRNAs.
Clinical therapeutic strategies for BC. Currently, endocrine therapy, anti-HER2 targeted therapy and chemotherapy constitute the backbone of BC treatment in clinic. Drugs for endocrine therapies include selective estrogen receptor modulators (SERMs), selective estrogen receptor degraders (SERDs) and aromatase inhibitors. SERMs bind ER to antagonize the activity of estrogen, SERDs bind ER to promote its proteasome-mediated degradation, while aromatase inhibitors block the biosynthesis of estrogens from adrenal steroids. Drugs available for anti-HER2-targeted therapy include monoclonal antibodies (trastuzumab, pertuzumab and emtansine) and small molecules (lapatinib and neratinib). Chemotherapy standards for BC treatment are anthracylines and taxanes. In addition, novel therapeutic strategies including targeted therapies beyond HER2 and immunotherapy have been administered in clinical situations (drugs for BC treatment are shown in italic).
LncRNAs in the development of endocrine resistance
The underlying mechanism of most BC risk factors (e.g. menstrual factors such as early menarche, late menopause and short menstrual cycles [53], reproductive factors such as late pregnancies [54]) is the overexposure of mammary epithelium to ovarian hormones, especially estrogens and progesterone [55]. This suggests that aberrant female hormones may be the primary stimulus for uncontrolled breast cell proliferation. In agreement, around 75% of BC patients are diagnosed as HER2-negative luminal subtypes (with positive expression of hormone receptors), suggesting that the ER signaling pathway driven by estrogen is a major oncogenic pathway of most BC [56]. For the treatment of these luminal BC patients, endocrine therapies including selective estrogen receptor modulators (SERMs), selective estrogen receptor degraders (SERDs) and aromatase inhibitors are highly effective through the disruption of receptor binding or estrogen deprivation (Figure 1). SERMs such as the first-line endocrine agent tamoxifen can bind ER to antagonize the activity of estrogen, leading to transcriptional repression of ER target genes [57, 58]. SERDs are clinically effective by not only antagonizing ERs but also degrading them, as exampled by the only FDA-approved drug fulvestrant [59]. Aromatase inhibitors such as letrozole, anastrozole, and exemestane block the biosynthesis of estrogens from adrenal steroids [60]. Unfortunately, most of the patients treated with these endocrine therapies finally develop treatment resistance accompanied by poor prognosis. Therefore, a deeper understanding of the potential mechanisms is urgently needed for exploiting new effective therapeutic strategies.
BC cells develop multiple mechanisms to promote intrinsic and acquired resistance to endocrine therapy. Ligand-independent transactivation of ER is one of the major mechanisms responsible for de novo endocrine therapeutic resistance, in which certain lncRNAs play essential roles. It has been long described that ERK-induced ER phosphorylation elicits the estrogen-independent activation of ER signaling [22, 23]. Recently, it has been reported that linc-RoR is able to activate the MAPK/ERK signaling pathway through the regulation of the ERK-specific phosphatase DUSP7 in ER+ BC, which bypasses the ER signaling pathway thus facilitating the development of intrinsic resistance to endocrine therapy [25] (Figure 2). The above study demonstrates an alternative lncRNA-dependent pathway to activate ER signaling, further illustrating the fundamental role of lncRNAs in BC endocrine resistance. Disrupting linc-RoR-mediated ER activation may help reverse endocrine resistance.
LncRNAs-mediated endocrine therapy resistance. LncRNA TMPO-AS1 directly interacts and stabilizes the mRNA of the ERα encoding gene ESR1, leading to the hyper-proliferation of ER+ BC and endocrine resistance. In addition, linc-RoR promotes the degradation of the ERK-specific phosphatase DUSP7 thus enhancing ERK phosphorylation. The upregulation of MAPK/ERK pathway activates ER signaling independent of estrogen, resulting in intrinsic resistance to endocrine therapy. Furthermore, HOTAIR is transcriptionally suppressed by ER. Upon blocking ER signaling through endocrine therapy, HOTAIR is upregulated and promotes the expression of ER at the protein level, leaing to enhanced transcriptional activity of ER and accelerated endocrine resistance.
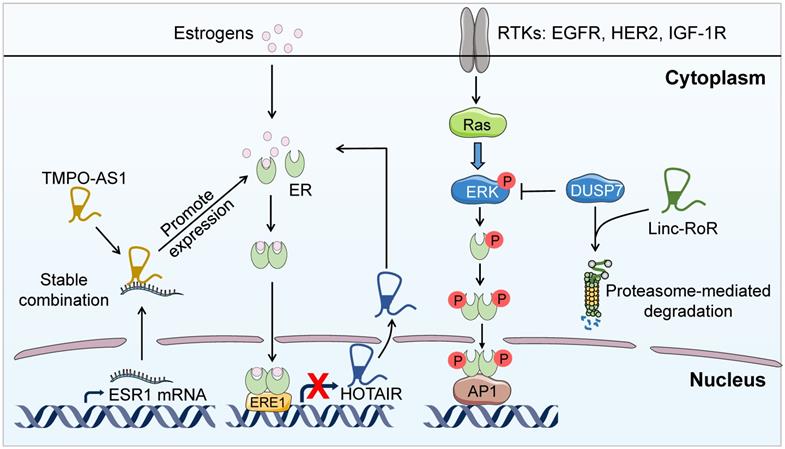
In addition, BC cells can survive endocrine therapeutic pressures through deregulation of the ER signaling components. Accumulating evidence indicates the involvement of lncRNAs in this mechanism. For example, lncRNA thymopoietin antisense transcript (TMPO-AS1) could interact with and stabilize the mRNA of the ERα encoding gene ESR1, leading to the hyper-proliferation of ER+ BC and possible endocrine resistance [61] (Figure 2). Besides, some lncRNAs are direct targets of ER and can possibly accelerate endocrine resistance resulting from ER signaling blockade. For instance, HOTAIR [11] and LINP1 [62] are transcriptionally suppressed by ER and therefore upregulated upon blocking ER signaling following endocrine therapy. Paradoxically, upregulated HOTAIR in turn promotes the expression of ER at the protein level and facilitates its transcriptional activity [11] (Figure 2). However, LINP1 overexpression downregulates the protein level of ER and diminishes the estrogen response to mediate anti-estrogen resistance [62]. The study of HOTAIR in endocrine resistance elucidates a positive feedback loop between ER and HOTAIR. Disrupting this loop may exhibit promising efficacy for the reversal of endocrine resistance. Moreover, DSCAM-AS1 has been found to be transcriptionally regulated by ER and confer tamoxifen resistance in BC by interacting with hnRNPL, but the detailed mechanism remaining to be determined [63].
Remarkably, a large number of lncRNAs regulate endocrine resistance in BC through the lncRNA-miRNA-mRNA axis, involving GAS5 [64], urothelial carcinomaassociated 1 (UCA1) [65], CYTOR [66], DSCAM‐AS1 [67] and lncRNA-ROR [68]. This indicates that lncRNAs as ceRNAs have profound implication in controlling endocrine response of BC. In addition, a newly published study has demonstrated that lncRNA HOTAIRM1 promotes acquired tamoxifen resistance in BC by interacting with EZH2, thus preventing the PRC2 complex-mediated H3K27me3 of the putative HOXA1 promoter [69]. This study indicates that HOTAIRM1 is a promising therapeutic target for BC patients with tamoxifen resistance.
LncRNAs in targeted therapy resistance
BC patients with HER2 overexpression or amplification have benefited significantly from HER2-tergeted therapeutics since the first anti-HER2 monoclonal antibody trastuzumab was developed in 1990 [70]. Currently, the first-line treatment for HER2-positive metastatic BC is trastuzumab and pertuzumab plus docetaxel [71], while the antibody-drug conjugate trastuzumab emtansine (T-DM1) is used as second-line therapy [72] (Figure 1). Broader HER2-targeted therapies, especially small molecules targeting HER2 such as lapatinib [73] and neratinib [74], have also been approved for the treatment of HER2+ BC patients. Anti-HER2 targeted therapies have appreciably prolonged overall survival of BC patients. In addition, targeted therapies beyond HER2, including inhibitors of PI3K/AKT/mTOR pathway, cyclin-dependent kinases (CDKs) and poly (ADP-ribose) polymerase (PARP), have been applied in clinical practice to prolong the survival for patients diagnosed with BC (Figure 1). These targeted therapies have exhibited superior efficacy in clinical trials and have greatly benefited BC patients. However, the effectiveness of targeted therapies has been largely restricted by high rates of resistance, and lncRNAs are emerging as pivotal regulators to mediate therapeutic tolerance.
Therapies targeting HER2
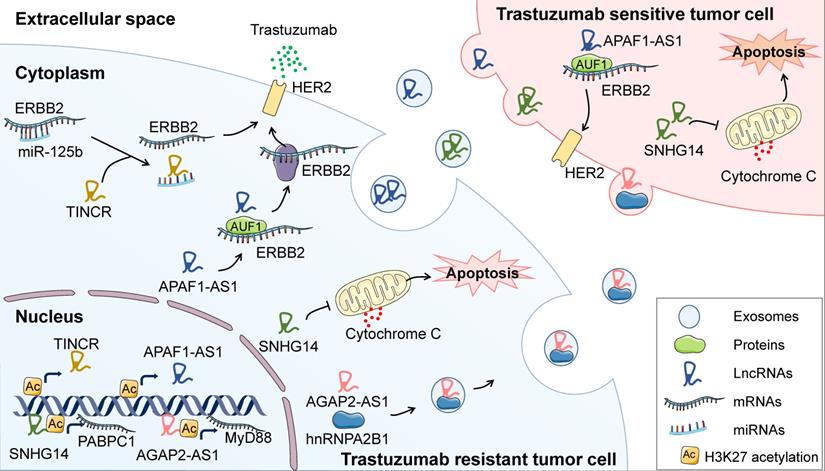
LncRNAs have attracted increasing attention as pivotal regulators of trastuzumab resistance in BC. Some studies have revealed that lncRNAs which confer acquired trastuzumab resistance in BC cells can be incorporated into exosomes thus disseminating the resistance to surrounding cells (Figure 3). For example, in trastuzumab resistant BC cells, lncRNA AFAP1-AS1 is upregulated through the H3K27 acetylation at its promoter region and guides AUF1, which improves the translation of target mRNA, to bind to HER2 mRNA thus enhancing HER2 translation and trastuzumab resistance. Strikingly, AFAP1-AS1 in trastuzumab resistant cells can be packaged into exosomes and promote resistance in recipient cells [75]. In addition, lncRNA AGAP2-AS1 [76] and SNHG14 [77, 78] can also facilitate trastuzumab tolerance of BC cells through exosome-mediated dissemination. These findings reveal that exosomes play fundamental roles in lncRNAs-mediated BC resistance to targeted therapies. Further, exosomal lncRNAs have also been widely documented to modulate cancer therapeutic resistance in other types of tumors [79]. Therefore, in addition to target lncRNA, blocking the packaging or secretion of exosomes may become a promising strategy for attenuating therapeutic resistance.
Regulatory function of lncRNAs in BC resistance to anti-HER2 targeted therapy. Under trastuzumab exposure, TINCR is upregulated by H3K27 acetylation and induces trastuzumab resistance through sponging miR-125b and releasing HER2 in BC cells. Moreover, AFAP1-AS1 can be upregulated through H3K27 acetylation at its promoter region and guides AUF1 to bind to HER2 mRNA, leading to enhanced translation of HER2. AGAP2-AS1 increases H3K27 acetylation at the promoter region of MyD88, resulting in the activation of NF-κB signaling pathway and therapeutic resistance to trastuzumab. Moreover, another lncRNA, known as SNHG14, could inhibit trastuzumab-induced apoptosis through upregulating Bcl2. Intriguingly, several lncRNAs have been reported to confer trastuzumab resistance in surrounding cells through being engulfed in exosomes and incorporated by neighbor cells.
According to a recent study, another lncRNA, TINCR, can be activated by CREB-binding protein (CBP)-mediated H3K27 acetylation, leading to trastuzumab resistance in BC [21]. Mechanistically, TINCR acts as a sponge for miR-125b, thus releasing HER2 to compromise the anti-tumor effect of trastuzumab [21]. Perhaps the most common functional mechanism of lncRNAs is remodeling chromatin structure to regulate gene expression, through which lncRNAs may enhance the resistance to trastuzumab treatment. For example, lncRNA AGAP2-AS1 induced by the transcription factor SP1 binds to CBP and increases H3K27 acetylation at the promoter region of MyD88, resulting in the activation of the NF-κB signaling pathway and resistance to trastuzumab [80]. Other lncRNAs, as exampled by H19 [81], UCA1 [82] and GAS5 [83] have been documented to be closely related to trastuzumab resistance; however, the mechanisms are as yet to be determined. In spite of increased understanding of lncRNAs-regulated trastuzumab resistance, the involvement of lncRNAs in resistance to pertuzumab and other small molecules targeting HER2 needs further exploration.
Therapies targeting PI3K/AKT/mTOR pathway
The PI3K/AKT/mTOR pathway is commonly hyperactivated in BC due to frequent somatic PIK3CA mutations and HER2-triggered oncogenic signaling. Some PI3K inhibitors have been launched for clinical use in BC treatment (Figure 1). For example, the novel and specific PI3Kα inhibitor NVP-BYL719 is now under active preclinical and clinical studies for the treatment of BC patients with PIK3CA mutations and/or HER2 amplification [84]. In particular, alpelisib, which is indicated for use in combination with fulvestrant in ER+, HER2- BC patients, is the first PI3K inhibitor that has been approved by the FDA [85]. The mTOR inhibitor everolimus increases PFS by more than twofold in ER+, HER2- advanced BC patients after failure of treatment with nonsteroidal aromatase inhibitors [2]. In addition, a phase-II clinical trial has demonstrated that the pan-AKT inhibitor MK2206 could increase pathologic complete response rates when combined with standard neoadjuvant therapy in ER-/PR- and HER2+ BC [86]. The AKT inhibitors capivasertib and ipatasertib in combination with first-line paclitaxel therapy significantly prolong the PFS of TNBC patients, especially those with PIK3CA/AKT1/PTEN-alterations [87, 88]. However, the PI3K/AKT/mTOR axis may be reactivated by compensatory signaling pathways dependent on lncRNAs, resulting in resistance to targeted therapies.
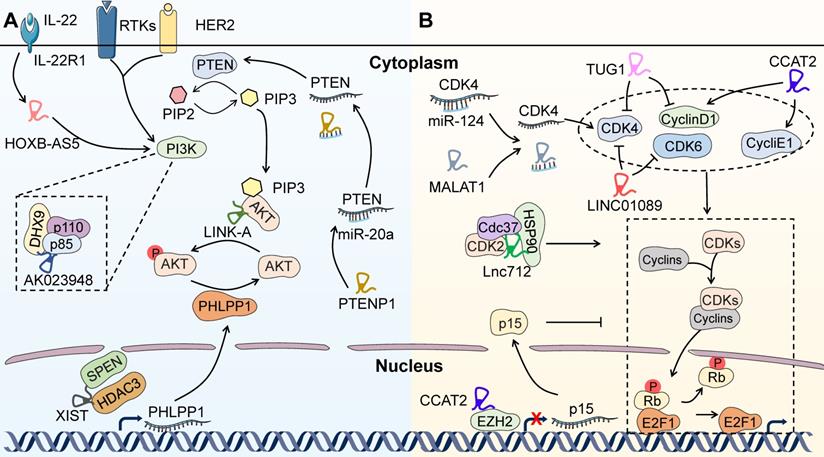
A recent study has shown that LINK-A directly binds to AKT and PIP3 to enhance the interaction between them, thus facilitating the enzymatic activation of AKT. LINK-A-induced hyperactivation of AKT is responsible for BC resistance to MK2206, which raises a hurdle to AKT targeted therapy in BC patients [89] (Figure 4A). Therefore, LINK-A may be a promising biomarker and therapeutic target for predicting AKT inhibitor efficacy and reversing treatment tolerance. Besides, a CRISPR/Cas9-based synergistic activation mediator (SAM) system has been developed and identified lncRNA AK023948 as a positive AKT regulator in BC through interacting with ATP-dependent RNA helicase A (RHA/DHX9) and p85 to sustain the stability of p85 [90] (Figure 4A). Therefore, AK023948-regulated AKT activation may facilitate resistance to AKT-targeted therapies in BC patients. Recently, Zhou et al. have documented that Linc-ROR decreases BC cell sensitivity to mTOR inhibitor rapamycin by sponging miR-194-3p and releasing methyl CpG-binding protein 2 (MECP2) [91]. In spite of the puzzling mechanism, this study proves a direct link between lncRNA and BC resistance to therapies targeting PI3K/AKT/mTOR pathways, again suggesting that lncRNAs are promising therapeutic targets to overcome BC resistance. In addition, it has been reported that IL-22 and the lncRNA HOXB-AS5 could synergistically activate the PI3K/AKT/mTOR pathway [15] (Figure 4A). Remarkably, IL-22 and HOXB-AS5 are detectable in the serum of BC patients, both of which are upregulated and closely related to the clinical stage of BC. Therefore, HOXB-AS5 represents an ideal biomarker to predict therapeutic response to PI3K/AKT/mTOR targeted therapy, which still requires further validation before entering the clinic [15]. Many other lncRNAs have also been reported to be capable of regulating PI3K/AKT/mTOR pathway, such as PTENP1 [92, 93], MALAT1 [94] and Xist [95], indicating the possible involvement of these lncRNAs in regulating resistance to the targeted therapies in BC (Figure 4A). However, the underlying mechanism remains largely unknown and requires further elucidation.
LncRNAs-regulated resistance to targeted therapies beyond HER2. LncRNAs are emerging as pivotal regulators to mediate therapeutic tolerance to targeted therapies beyond HER2, including inhibitors of PI3K/AKT/mTOR pathway, CDKs and PARP. (A) LncRNAs-mediated resistance to therapies targeting PI3K/AKT/mTOR pathway. LINK-A directly binds to AKT and PIP3 to facilitate the enzymatic activation of AKT, leading to BC resistance to MK2206. AK023948 interacts with DHX9 and p85 to positively regulate AKT. In addition, IL-22-induced HOXB-AS5 expression could activate PI3K. Conversely, PTENP1 sponges miR-20a to release PTEN and negatively regulates PI3K/AKT/mTOR pathway. Furthermore, XIST can sequester HDAC3 to enhance the transcription of PHLPP1 and dephosphorylation of AKT. (B) LncRNAs-mediated resistance to therapies targeting CDKs. Lnc712 activates CDK2 through directly interacting with HSP90 and forming a complex of Lnc712/HSP90/Cdc37. In addition, MALAT1, TUG1, CCAT2 and LINC01089 are associated with the altered function of CDKs and Cyclins, which may be required for the drug resistance to CDK inhibitors in BC cells.
Therapies targeting CDKs
Preclinical and clinical studies have demonstrated the synergistic effect of CDKs inhibitors with anti-estrogen agents, probably due to the special dependence of ER+ BC cells on cyclin D1 and estrogen-mediated activation of CDKs [96]. CDK4/6 inhibitors such as palbociclib [97], ribociclib [98] and abemaciclib [99] combined with endocrine agents have exhibited significant PFS benefit and are approved by the FDA as the standard treatment for ER+, HER2- advanced BC patients (Figure 1). Furthermore, a potent dual inhibitor of CDK12/CDK13 has been developed and demonstrated to provoke TNBC cell death by suppressing the pivotal DNA damage response genes and triggering lethal accumulation of DNA damage [100]. This study raises the exciting possibility of developing targeted therapies for TNBC, but still needs further clinical investigation. As with all the other therapeutic strategies, the emergence of resistance to CDK inhibitors is a major clinical obstacle.
The most common mechanism underlying resistance to CDK4/6 inhibitors is cell cycle alterations [101], with some clues implying the participation of lncRNAs (Figure 4B). It has been shown that the responsiveness of CDK4/6 inhibitor palbociclib could be restricted by elevated CDK2 expression or activity, suggesting that redundant CDK functions may predict treatment failure for CDK inhibitors [102, 103]. A newly identified lncRNA, Lnc712, could activate CDK2 by directly interacting with heat-shock protein 90 (HSP90) and forming a complex of Lnc712/HSP90/cell division cycle 37 (Cdc37) in BC [16]. These findings indicate that Lnc712 may enhance resistance to palbociclib and become a promising biomarker for the prediction of drug response in BC. Similarly, lncRNAs associated with other CDKs, including MALAT1 [104], TUG1 [105], CCAT2 [106] and LINC01089 [107], may also mediate resistance to CDK inhibitors. Overall, the direct nexus between lncRNAs and CDK inhibitors is not yet well established. Further investigation focused on the regulation of CDK inhibitor responsiveness by lncRNAs may help to understand the mechanisms underlying treatment resistance.
Therapies targeting PARP
PARP is an enzyme able to initiate single-strand DNA break repair by synthesizing a polymeric adenosine diphosphate ribose (PAR) chain and recruiting critical DNA-repairing enzymes. PARP inhibition leads to DNA double-strand breakage, which is normally repaired by homologous recombination (HR) dependent on BRCA1 and BRCA2 [108]. Since BRCA mutation predisposes to certain cancers as exampled by BC, it can be anticipated that BRCA-mutated BC will be sensitive to PARP inhibitors. Supporting this, two PARP inhibitors, olaparib (Lynparza) and talazoparib (Talzenna), received FDA approval for the treatment of germline BRCA-mutated, HER2- locally advanced or metastatic BC in 2018.
Nevertheless, the benefit of PARP inhibitors has been shown to be heavily compromised by drug resistance. The main mechanisms of PARP inhibitor resistance identified to date include disrupting cellular drug availability, affecting (de)PARylation enzymes, reactivating HR and restoring replication fork stability [109]. Emerging evidence shows the intimate correlation between lncRNAs and DNA damage repair, strongly supportive of lncRNAs involvement in PARP inhibitor resistance. For example, ataxia-telangiectasia mutated (ATM)-mediated DNA damage response has been found to induce the expression of lncRNA-JADE. lncRNA-JADE directly binds to BRCA1 and transcriptionally activates Jade1, a critical element in human acetylase binding to ORC1 (HBO1) histone acetylation complex. The Jade1 activation mediated by lncRNA-JADE promotes global histone H4 acetylation and increases transcription of DNA damage repair-related genes [110]. Linc00261 has been demonstrated to be epigenetically regulated by FOXA2 and induce phosphorylation and activation of DNA damage machinery [111]. Other lncRNAs, such as mitotically-associated long noncoding RNA (MANCR) [112] and transcribed in the opposite direction of RAD51 (TODRA) [113], are revealed to maintain genomic stability and induce HR. Given the capability of these lncRNAs to enhance DNA damage repair and sustain genomic stability, it can be assumed that lncRNAs may play an essential role in PARP inhibitor resistance.
Moreover, in consideration of the key role of the tumor suppressor p53 in maintaining genome stability [114], it is conceivable that an enigmatic nexus may exist between p53 status and PARP inhibitor response. Recently, some p53-responsive lncRNAs have been reported to play essential roles in BC, indicating their potential involvement in PARP inhibitor resistance. For instance, the lncRNA GUARDIN induced by p53 plays an essential role in the maintenance of genome integrity by sustaining the stability of telomeric repeat-binding factor 2 (TRF2) and BRCA1 [115]. The correlation between GUARDIN and p53 as well as the competence of GUARDIN to maintain BRCA1 expression imply that GUARDIN may confer intrinsic resistance to PARP inhibitor. Another lncRNA in the nonhomologous end joining (NHEJ) pathway (LINP1) is also regulated by p53 and overexpressed in TNBC [116]. LINP1 can facilitate DNA double-strand breaks repair, suggesting its potential to counteract PARP inhibitor-induced DNA damage repair vulnerability [116]. An exciting recent study has revealed that downregulated lncRNA PHACTR2-AS1 in BC is associated with tumor development and poor prognosis. Aberrant activation of EZH2 targets and downregulates PHACTR2-AS1, which triggers H3K9 methylation-mediated silencing of ribosome DNA genes, thus inducing genome instability [117]. This finding further implicates the involvement of lncRNAs in DNA damage signaling. However, the direct correlation between lncRNAs and PARP inhibitor resistance remains to be established.
LncRNAs in BC chemoresistance
Chemotherapy is beneficial to the treatment of almost every subtype of BC. Currently, anthracylines and taxanes are standard chemotherapy for early stage BC [1] (Figure 1). Anthracylines such as doxorubicin exert their cytotoxic functions via pleiotropic mechanisms including macromolecular biosynthesis inhibition, free radical production, and DNA damage induced by histone eviction from open chromatin [118, 119]. Taxanes (e.g. paclitaxel and docetaxel) can bind and stabilize microtubules to prevent depolymerization and block mitosis progression [120, 121]. For HER2+ metastatic BC, as mentioned above, trastuzumab in combination with taxane chemotherapy has been found to improve overall survival (OS) and has been the first-line standard treatment since 2001 [122]. Due to the lack of specific biomarkers in TNBC, targeted therapies have rarely met the need to improve clinical outcomes. Therefore, chemotherapy is always recommended as the standard of care for TNBC patients.
Currently, dysregulated lncRNAs have been widely documented to play dual roles in BC chemoresistance. Among all the studies available to date, most of the lncRNAs enhance chemoresistance in BC by acting as ceRNAs to sponge miRNAs, especially through targeting the ATP-binding cassette (ABC) transporter superfamily. The ABC transporter superfamily as drug efflux pumps have been known to mediate multidrug resistance (MDR) in multiple cancers [123]. For example, lncRNA ferritin heavy chain 1 pseudogene 3 (FTH1P3) and linc00518 respectively sponge miR-206 and miR-199a, which share complementary binding sites with ABCB1 and MRP1 mRNA, thus mediating MDR (e.g. paclitaxel, doxorubicin and vincristine) in BC [124, 125]. Besides, dysregulated NONHSAT101069 is reported to sponge miR-129-5p and release Twist1 to confer resistance to the anthracycline genotoxic drug epirubicin in BC [126], and CASC2 enhances paclitaxel resistance through regulation of the miR-18a-5p-CDK19 axis [127]. On the basis of several independent studies, NEAT1 confers resistance to paclitaxel, cisplatin and 5-fluorouracil in BC cells through miR-129/ZEB2 and miR-211/HMGA2 pathways [12, 13]. Collectively, the mechanism whereby lncRNAs act as ceRNAs to release specific mRNAs is widely seen in BC chemotherapy resistance. Specifically targeting the lncRNA-miRNA-mRNA axis may benefit BC patients resistant to chemotherapy.
In addition to acting as ceRNAs, lncRNAs can exert their function of mediating BC chemoresistance through multiple mechanisms. Of note, a recent study has reported a special mechanism of the antisense lncRNA MAPT-AS1 to mediate chemoresistance in BC [128]. In detail, MAPT-AS1 contributes to paclitaxel resistance in ER- BC cells through the formation of RNA duplex with its natural comparable sense transcripts MAPT. The RNA duplex may alter the spatial structure and increase the stability of MAPT mRNA, which has been demonstrated to be involved in BC chemoresistance [128]. LncRNA NONHSAT141924 enhances BC resistance to paclitaxel through the p-CREB/Bcl-2 apoptosis pathway [129]. Intriguingly, H19 can induce doxorubicin resistance in BC, and be engulfed into exosomes to disseminate the resistance to surrounding sensitive cells [130, 131]. Thus, targeting H19 in BC cells as well as blocking its incorporation into exosomes may exhibit superior efficacy in reversing BC chemoresistance.
Compared with lncRNAs which promote chemoresistance in BC, fewer lncRNAs have been found to suppress chemoresistance in BC and the underlying mechanisms still remain elusive. For instance, LINC00968 targets and silences WNT2 and the downstream β-catenin signaling pathway, thus sensitizing BC cells to chemotherapeutics [132]. LncRNA GAS5 suppresses adriamycin resistance by competing with miR-221-3p for DKK2 release, which downregulates the Wnt/β-catenin signaling pathway [133]. Besides, AC073284.4 attenuates paclitaxel resistance via inhibition of miR-18b-5p/dedicator of cytokinesis protein 4 (DOCK4) axis [134]. Taken together, lncRNAs are extensively involved in BC chemoresistance. Modulating the expression of chemoresistance-related lncRNAs is a feasible strategy for the reversal of resistance.
LncRNAs in BC immunotherapy resistance
Currently, there is a growing enthusiasm for the application of immunotherapy in BC treatment, with the immune checkpoint blockade playing the leading role [135] (Figure 1). Agents causing immune checkpoint blockade are under active clinical investigation, either used alone or in combination with other therapeutics. For example, some chemotherapeutics and targeted agents appear promising in combination with immune checkpoint inhibitors because of their competence in T-cell priming and immunity activation [135]. Of note, in 2019 the immune checkpoint blockade agent atezolizumab (Tecentriq) was approved to be paired with cytotoxic chemotherapeutic nab-paclitaxel (Abraxane) for the first-line treatment of PD-L1+ unresectable locally advanced or metastatic TNBC. This initial approval of an immunotherapeutic marks a milestone in BC treatment, especially in TNBC, since prior to that cytotoxic chemotherapeutics were the only treatment option for these patients.
In spite of the encouraging progress in BC immunotherapy, cancer cells can develop variable mechanisms to escape immunosurveillance and generate intrinsic unresponsiveness to immunotherapeutics. One of the immune-resistant mechanisms involves reducing antigenicity that avoids detection by antitumor lymphocytes. For instance, TNBC exhibits resistance to programmed cell death protein-1(PD-1) blockade through downregulation of PLC, which enhances antigen presentation to the cell surface [136]. More intriguingly, the lncRNA LINK-A plays a key role in this process by promoting the degradation of PLC. In agreement with this, treatment with LINK-A LNAs stabilizes PLC components and sensitizes mammary gland tumors to immune checkpoint inhibitors [17] (Figure 5). This study gives an example of lncRNA-dependent antigenicity reduction and intrinsic immunotherapy resistance, which underscores the significance of lncRNAs as potential therapeutic targets and novel biomarkers to predict immunotherapy responses of BC patients.
LncRNAs-regulated resistance to immunotherapy. In BC cells, LINK-A promotes the crosstalk between PIP3 and inhibitory GPCR pathways, leading to decreased cAMP and PKA-mediated phosphorylation of TRIM71. Decreased phosphorylation of TRIM71 enhances the degradation of PLC, Rb and p53, resulting in downregulation of antigenicity and intrinsic tumor resistance to immunotherapy. In stimulated T cells, calcium influx activates calmodulin, thereby removing HDAC and enhancing STAT1-mediated transcription of NKILA. NKILA directly binds to p65 and prevents its nuclear translocation as well as following transcription of anti-apoptotic genes. Thus, NKILA facilitates T cell vulnerability to AICD, resulting in immune evasion and cancer progression of BC.
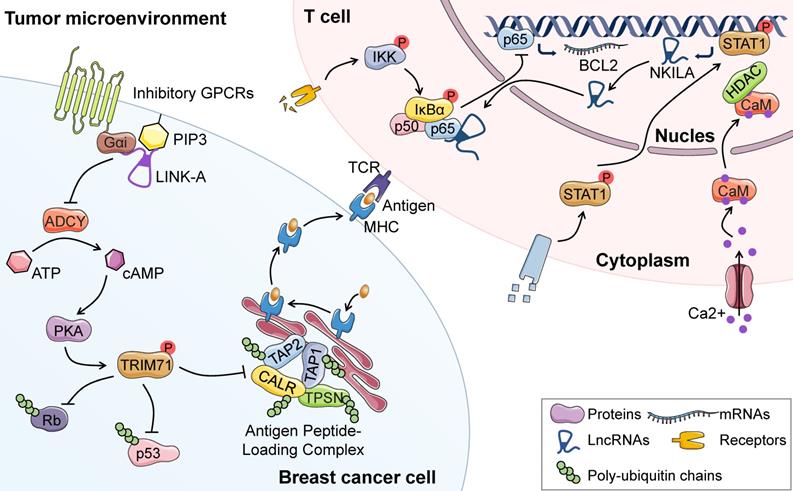
In addition to the downregulation of antigenicity, transformed cells take advantage of AICD of T lymphocytes to escape immunological elimination, in which the lncRNAs play an important role (Figure 5). For example, a recent study has demonstrated that an NF-κB-interacting lncRNA, NKILA, facilitates T cell vulnerability to AICD through inhibition of NF-κB activity, resulting in BC immune evasion and cancer progression. Moreover, knockdown of NKILA in tumor-specific CTLs significantly inhibits the growth of BC patient-derived xenografts due to increased CTL infiltration [52]. Thus, modulating the expression of oncogenic lncRNAs such as NKILA in CTL may represent a novel immunotherapeutic strategy. Moreover, the expression levels of XIST and TSIX are related to PD-L1 in the tissues and body fluids of BC patients with diverse molecular subtypes, indicating the role of the two lncRNAs as predictive biomarkers for immunotherapy in BC patients [137]. Emerging studies have shed light on the mechanism by which XIST attenuates immune suppression. For instance, XIST depletion enhances the secretion of exosomal miRNA-503 to activate M1-M2 conversion of microglia, which is essential for maintaining the suppressive immune microenvironment and inhibiting T cell proliferation [138]. In addition to those lncRNAs mentioned above, multiple lncRNAs have been found to be associated with immune system regulation, including RP4-583P15.10 [139], T-cell leukemia/lymphoma 6 (TCL6) [140], growth hormone secretagogue receptor opposite strand (GHSROS) [141], and linc00152 [142]. These findings demonstrate that lncRNAs may act as promising predictive biomarkers and therapeutic targets for BC immunotherapy.
Clinical implications of lncRNAs in BC therapy
Early stage BC is considered to be potentially curable while the treatment for metastatic or late BC is still challenging, suggesting that early and precise diagnosis is extremely important for BC patients. Currently, two-view mammography is the sole screening method recommended for early detection of BC [6]. However, the high frequency of false positives or negatives limits the reliability of this clinically used screening tool and may impact on survival or impair life quality of patients [143]. Interestingly, a deep learning-based model can outperform radiologists in identifying BC from mammograms, but the clinical benefits are not yet fully established [144]. Many state-of-the-art imaging methods, including abbreviated MRI [6] and digital breast tomosynthesis (DBT) [145], are being developed to expand the sensitivity profile of early stage BC detection. However, the enormous cost, sensitivity and specificity of these screening methods remain to be optimized.
Although ER, PR, and HER2 provide insight into BC molecular classification and clinical management, novel biomarkers for BC subtyping are urgently needed to facilitate individualized treatment due to the high intertumor heterogeneity in BC patients. In addition to the lack of effective biomarkers for early detection and molecular subtyping, resistance against standard therapies has also increased the mortality of BC as discussed above. Therefore, sensitive biomarkers and effective therapeutic targets are urgently required to improve the clinical outcomes of BC patients.
LncRNAs as biomarkers to predict therapeutic response
Mining novel biomarkers for BC management is extremely important for improving clinical outcomes. Some lncRNAs have been found to act as essential regulators in BC, indicating their potential to serve as biomarkers for early diagnosis and molecular subtyping. For instance, Xu et al. has reported that lncRNA RP11-445H22.4 in circulating serum is a sensitive and specific diagnostic biomarker for BC [146]. Liu et al. have developed a novel classification system integrating lncRNA and mRNA expression profiles for TNBC and demonstrated that subtype-specific lncRNAs could act as potential biomarkers for BC classification [147]. Another recent study has demonstrated that the overexpression of HOTAIR is intimately associated with a luminal androgen receptor (LAR) subtype of TNBC, which is characterized by AR expression [148]. These findings indicate the prognostic significance of lncRNAs in BC and raise the possibility of establishing novel detection strategies for BC patients.
In addition, many lncRNAs have been found to be capable of predicting therapeutic response. High expression of HOTAIR and its regulator FOXM1 can help identify endocrine therapy non-responders among ER+ BC patients, illustrating the role of lncRNAs as predictive biomarkers in BC therapy [149]. In addition, the histone deacetylase inhibitor abexinostat has been demonstrated to induce BCSCs differentiation and reduce BCSC population in vivo. XIST functions as a predictive biomarker to distinguish BC patients sensitive to this differentiation therapy [150]. XISTlow BC cells can be specifically eliminated by a FDA-approved chemotherapy medication fludarabine [138]. Further, multigene signatures incorporating lncRNAs have been developed and demonstrated to be capable of predicting taxane responsiveness, tumor recurrence and clinical outcome in patients diagnosed with TNBC [151, 152]. The functionally polymorphic lncRNA MIR2052HG can influence the risk of tumor recurrence in BC patients treated with aromatase inhibitors [153]. These studies reveal that lncRNAs can serve as active biomarkers for therapeutic response prediction, but need further validation in clinical practice. This would be beneficial for personalized/precision medicine.
Interestingly, lncRNAs, including Prostate Cancer gene 3 (PCA3), MALAT1, H19, TINCR and CCAT2, are emerging as circulating biomarkers for early stage cancer detection [154, 155]. A recent study has reported that extracellular RNAs (exRNAs) from a single droplet (5-7 µL) of serum in a liquid biopsy are capable of reflecting human physiological and disease states using SILVER-seq (Small Input Liquid Volume Extracellular RNA Sequencing). More intriguingly, lncRNAs are detectable in this system, and donors with or without BC display significant differences [156]. These findings indicate that lncRNAs can act as sensitive biomarkers in BC non-invasive liquid biopsy, but again this needs further clinical validation. Unfortunately, the BC patient cohorts used in the above studies are relatively small, and larger studies are required to validate the clinical value of these lncRNAs as biomarkers.
LncRNAs as promising BC therapeutic targets
Notwithstanding the several therapeutic strategies available for BC patients, overcoming treatment tolerance remains a major challenge for improving clinical outcome. Theoretically, the regulatory roles of lncRNAs in BC therapeutic resistance confer them potential to be targeted for reversing treatment tolerance. Furthermore, in practice, the development of lncRNA-targeting technologies makes it possible to translate lncRNA-based therapies to the clinic. To date, the most general approaches for suppressing lncRNA involve antisense oligonucleotides (ASOs) and small interfering RNAs (siRNAs) [157]. ASOs refer to synthetic single-stranded oligonucleotides which are complementary to target lncRNAs and can form a DNA/RNA heteroduplex which can be cleaved by RNase H. siRNAs are double-stranded RNAs, the guide strand of which can be loaded into argonaute 2 (AGO2) and then form a RNA-induced silencing complex (RISC) to target lncRNA for degradation. ASOs predominantly target lncRNAs residing in the nucleus while cytoplasmic lncRNAs are more sensitive to siRNAs [158].
Both of these two methods have been applied to silence lncRNAs and have shown considerable efficacy against BC in preclinical studies. Genetic knockout or knockdown of MALAT1 [159], LINC02273 [160] and LINC00673 [161] using ASOs has exhibited superior efficacy in attenuating BC growth and metastasis in vivo. siRNAs targeting BC-related lncRNAs such as HOTAIR have also been demonstrated to inhibit BC growth and invasion [162]. Remarkably, silence of lnc-BM with siRNAs encapsulated within nanoparticles has shown considerable efficacy against brain metastasis in BC [163], indicating that ASOs or siRNAs formulated into suitable drug delivery vehicles (e.g. liposomes, nanoparticles and viruses) may accelerate the clinical application of lncRNAs-based treatment strategies.
LNAs are well accepted as a novel class of RNA used in therapeutics. The ribose moiety of the RNA nucleotide is locked by an extra oxymethylene bridge linking the C (2') and C (4') to facilitate base stacking and enhance hybridization properties [164, 165]. The unique characteristics of LNAs, including high binding affinity, high stability and improved mismatch discrimination, make them emerging approaches for BC treatment [166]. Therapeutic delivery of LNAs targeting LINK-A [17] and BCAR4 [167, 168] has been proven to sensitize BC to immune checkpoint inhibitors and attenuate BC growth and metastasis in vivo, respectively. Preclinical studies on lncRNA-based BC therapy have made substantial progress and offered an opportunity to reverse BC therapeutic resistance. However, these studies are still preliminary and great challenges lie ahead for translating these treatment methods into the clinic, requiring efforts in oligonucleotide chemistry and development of appropriate delivery systems.
Conclusions
In the past few decades, studies on lncRNAs in the field of oncology have drawn considerable attention of researchers, largely due to the rapid and substantial progress in high-throughput sequencing technologies. Additionally, the aberrant expression and multifaceted roles of lncRNAs in cancer have also attracted increasing scientific interests. In this review, we summarize the dysregulated lncRNAs associated with resistance to current BC therapeutic strategies and highlight the underlying mechanisms to facilitate the understanding of BC treatment tolerance.
The lncRNAs implicated in BC therapy resistance hold potential to serve as predictive biomarkers or therapeutic targets to benefit clinical management of BC patients. Firstly, lncRNAs have been widely documented as biomarkers for BC diagnosis, prognosis and therapeutic response prediction. The sensitivity and specificity have been demonstrated in several cohorts of BC patients. It therefore appears that lncRNAs are promising biomarker candidates to be translated into clinical applications. Unfortunately, the size of many clinical studies conducted to date is not large enough to verify their clinical value. Thus, larger BC patient cohorts are needed to validate the potential of these lncRNAs as clinical biomarkers.
Furthermore, the prerequisite of translating lncRNAs to the clinic as therapeutic targets is to elucidate their elaborated mechanisms of action in vitro and in vivo. However, the major impediment that lies ahead is their often obscure mode of action. Additionally, the low sequence conservation of lncRNAs poses a challenge in validating their biological functions. It has been suggested that the secondary structure of lncRNAs are evolutionarily conserved and could be considered as the major functional unit to regulate biological activities [169]. However, very few studies focus on the influence of lncRNA secondary structure on their biological functions. Additionally, the same lncRNA may have multiple targets and could even exert opposite functions in different types of tumors, resulting in dramatic side effects. Lack of animal models remains another limitation in mechanistic studies on lncRNAs. Further exploration on the character of lncRNAs and more technical breakthroughs are required to address these problems. Taken together, investigation of lncRNAs in BC therapeutic resistance may further enhance our understanding of the mechanisms responsible for resistance to BC treatment. Hopefully, some of the specific lncRNAs will enter the clinic as promising prognostic biomarkers or effective therapeutic targets.
Abbreviations
ABC: ATP-binding cassette; AGO2: argonaute 2; AICD: activation-induced cell death; ASOs: antisense oligonucleotides; ATM: ataxia-telangiectasia mutated; BC: breast cancer; BCSCs: breast cancer stem cells; cAMP: cyclic AMP; CBP: CREB-binding protein; Cdc37: cell division cycle 37; CDKs: cyclin-dependent kinases; ceRNAs: competing endogenous RNAs; CTLs: cytotoxic T lymphocytes; DBT: digital breast tomosynthesis; DOCK4: dedicator of cytokinesis protein 4; EMT: epithelial-mesenchymal transition; ER: estrogen receptor; exRNAs: extracellular RNAs; FTH1P3: ferritin heavy chain 1 pseudogene 3; GPCR: G-protein-coupled receptor; HBO1: human acetylase binding to ORC1; HER2: human epidermal receptor 2; Hh: hedgehog signaling; HR: homologous recombination; HSP90: heat-shock protein 90; iPSCs: induced pluripotent stem cells; LAR: luminal androgen receptor; LINK-A: long intergenic non-coding RNA for kinase activation; LNAs: locked nucleic acids; lncRNAs: long noncoding RNAs; MANCR: mitotically-associated long noncoding RNA; MDR: multidrug resistance; MECP2: methyl CpG-binding protein 2; miRNAs: microRNAs; MRI: magnetic resonance imaging; MSCs: mesenchymal stem cells; NEAT1: nuclear paraspeckle assembly transcript 1; NHEJ: nonhomologous end joining; OS: overall survival; PAR: polymeric adenosine diphosphate ribose; PARP: poly (ADP-ribose) polymerase; PCA3: Prostate Cancer gene 3; PD-1: programmed cell death protein-1; PFS: progression-free survival; PKA: protein kinase A; PLC: peptide-loading complex; PR: progesterone receptor; RHA/DHX9: RNA helicase A; RISC: RNA-induced silencing complex; SAM: synergistic activation mediator; SERDs: selective estrogen receptor degraders; SERMs: selective estrogen receptor modulators; SILVER-seq: Small Input Liquid Volume Extracellular RNA Sequencing; siRNAs: small interfering RNAs; TCL6: T-cell leukemia/lymphoma 6; T-DM1: trastuzumab emtansine; TMPO-AS1: thymopoietin antisense transcript; TNBC: triple-negative breast cancer; TRF2: telomeric repeat-binding factor 2; AICD: activation- induced cell death; CBP: CREB-binding protein; CDKs: cyclin-dependent kinases; ceRNAs: competing endogenous RNAs; CTLs: cytotoxic T lymphocytes; DOCK4: dedicator of cytokinesis protein 4; ER: estrogen receptor; FTH1P3: ferritin heavy chain 1 pseudogene 3; GPCR: G-protein-coupled receptor; HER2: human epidermal receptor 2; LINK-A: long intergenic non-coding RNA for kinase activation; lncRNAs: long noncoding RNAs; MECP2: methyl CpG-binding protein 2; NEAT1: nuclear paraspeckle assembly transcript 1; PARP: poly (ADP-ribose) polymerase; PLC: peptide-loading complex; TMPO-AS1: thymopoietin antisense transcript; TRF2: telomeric repeat-binding factor 2; UCA1: urothelial carcinoma associated 1.
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (81821002, 81790251, 81672381, 81972665 and 81770580), Guangdong Basic and Applied Basic Research Foundation (2019B030302012), the National 973 Basic Research Program of China (2013CB911300).
Author contributions
Na Xie and Yuan-Yuan Zhang conceived the structure of manuscript and revised the manuscript. Edouard C. Nice made critical revisions and polished the manuscript. Liyuan Peng and Jingwen Jiang collected the related paper and finished the initial manuscript. Bo Tang made the Figures and Table. All authors read and approved the final manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Harbeck N, Gnant M. Breast cancer. Lancet. 2017;389:1134-50
2. Piccart M, Hortobagyi GN, Campone M, Pritchard KI, Lebrun F, Ito Y. et al. Everolimus plus exemestane for hormone-receptor-positive, human epidermal growth factor receptor-2-negative advanced breast cancer: overall survival results from BOLERO-2dagger. Ann Oncol. 2014;25:2357-62
3. Finn RS, Crown JP, Lang I, Boer K, Bondarenko IM, Kulyk SO. et al. The cyclin-dependent kinase 4/6 inhibitor palbociclib in combination with letrozole versus letrozole alone as first-line treatment of oestrogen receptor-positive, HER2-negative, advanced breast cancer (PALOMA-1/TRIO-18): a randomised phase 2 study. Lancet Oncol. 2015;16:25-35
4. Siu AL, Force USPST. Screening for Breast Cancer: U.S. Preventive Services Task Force Recommendation Statement. Ann Intern Med. 2016;164:279-96
5. Friedewald SM, Rafferty EA, Rose SL, Durand MA, Plecha DM, Greenberg JS. et al. Breast cancer screening using tomosynthesis in combination with digital mammography. JAMA. 2014;311:2499-507
6. Kuhl CK. Abbreviated Magnetic Resonance Imaging (MRI) for Breast Cancer Screening: Rationale, Concept, and Transfer to Clinical Practice. Annu Rev Med. 2019;70:501-19
7. Consortium EP. An integrated encyclopedia of DNA elements in the human genome. Nature. 2012;489:57-74
8. Beermann J, Piccoli MT, Viereck J, Thum T. Non-coding RNAs in Development and Disease: Background, Mechanisms, and Therapeutic Approaches. Physiol Rev. 2016;96:1297-325
9. Brannan CI, Dees EC, Ingram RS, Tilghman SM. The product of the H19 gene may function as an RNA. Mol Cell Biol. 1990;10:28-36
10. Botti G, De Chiara A, Di Bonito M, Cerrone M, Malzone MG, Collina F. et al. Noncoding RNAs within the HOX gene network in tumor pathogenesis and progression. J Cell Physiol. 2018;234:395-413
11. Xue X, Yang YA, Zhang A, Fong KW, Kim J, Song B. et al. LncRNA HOTAIR enhances ER signaling and confers tamoxifen resistance in breast cancer. Oncogene. 2016;35:2746-55
12. Shin VY, Chen J, Cheuk IW, Siu MT, Ho CW, Wang X. et al. Long non-coding RNA NEAT1 confers oncogenic role in triple-negative breast cancer through modulating chemoresistance and cancer stemness. Cell Death Dis. 2019;10:270
13. Li X, Wang S, Li Z, Long X, Guo Z, Zhang G. et al. The lncRNA NEAT1 facilitates cell growth and invasion via the miR-211/HMGA2 axis in breast cancer. Int J Biol Macromol. 2017;105:346-53
14. Cantile M, Di Bonito M, Cerrone M, Collina F, De Laurentiis M, Botti G. Long Non-Coding RNA HOTAIR in Breast Cancer Therapy. Cancers (Basel). 2020 12
15. Rui J, Chunming Z, Binbin G, Na S, Shengxi W, Wei S. IL-22 promotes the progression of breast cancer through regulating HOXB-AS5. Oncotarget. 2017;8:103601-12
16. Cui Y, Lu C, Zhang Z, Mao A, Feng L, Fu L. et al. A Long Non-coding RNA Lnc712 Regulates Breast Cancer Cell Proliferation. Int J Biol Sci. 2020;16:162-71
17. Hu Q, Ye Y, Chan LC, Li Y, Liang K, Lin A. et al. Oncogenic lncRNA downregulates cancer cell antigen presentation and intrinsic tumor suppression. Nat Immunol. 2019;20:835-51
18. Liu K, Gao L, Ma X, Huang JJ, Chen J, Zeng L. et al. Long non-coding RNAs regulate drug resistance in cancer. Mol Cancer. 2020;19:54
19. Basak P, Chatterjee S, Bhat V, Su A, Jin H, Lee-Wing V. et al. Long Non-Coding RNA H19 Acts as an Estrogen Receptor Modulator that is Required for Endocrine Therapy Resistance in ER+ Breast Cancer Cells. Cell Physiol Biochem. 2018;51:1518-32
20. Cairns J, Ingle JN, Kalari KR, Shepherd LE, Kubo M, Goetz MP. et al. The lncRNA MIR2052HG regulates ERalpha levels and aromatase inhibitor resistance through LMTK3 by recruiting EGR1. Breast Cancer Res. 2019;21:47
21. Dong H, Hu J, Zou K, Ye M, Chen Y, Wu C. et al. Activation of LncRNA TINCR by H3K27 acetylation promotes Trastuzumab resistance and epithelial-mesenchymal transition by targeting MicroRNA-125b in breast Cancer. Mol Cancer. 2019;18:3
22. Kato S, Endoh H, Masuhiro Y, Kitamoto T, Uchiyama S, Sasaki H. et al. Activation of the estrogen receptor through phosphorylation by mitogen-activated protein kinase. Science. 1995;270:1491-4
23. Bunone G, Briand PA, Miksicek RJ, Picard D. Activation of the unliganded estrogen receptor by EGF involves the MAP kinase pathway and direct phosphorylation. EMBO J. 1996;15:2174-83
24. Vilgelm A, Lian Z, Wang H, Beauparlant SL, Klein-Szanto A, Ellenson LH. et al. Akt-mediated phosphorylation and activation of estrogen receptor alpha is required for endometrial neoplastic transformation in Pten+/- mice. Cancer Res. 2006;66:3375-80
25. Peng WX, Huang JG, Yang L, Gong AH, Mo YY. Linc-RoR promotes MAPK/ERK signaling and confers estrogen-independent growth of breast cancer. Mol Cancer. 2017;16:161
26. Bonnet D, Dick JE. Human acute myeloid leukemia is organized as a hierarchy that originates from a primitive hematopoietic cell. Nat Med. 1997;3:730-7
27. Wang C, Jia L, Wang Y, Du Z, Zhou L, Wen X. et al. Genome-wide interaction target profiling reveals a novel Peblr20-eRNA activation pathway to control stem cell pluripotency. Theranostics. 2020;10:353-70
28. Mani SA, Guo W, Liao MJ, Eaton EN, Ayyanan A, Zhou AY. et al. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell. 2008;133:704-15
29. Li J, Zhou BP. Activation of beta-catenin and Akt pathways by Twist are critical for the maintenance of EMT associated cancer stem cell-like characters. BMC Cancer. 2011;11:49
30. Tang T, Guo C, Xia T, Zhang R, Zen K, Pan Y. et al. LncCCAT1 Promotes Breast Cancer Stem Cell Function through Activating WNT/beta-catenin Signaling. Theranostics. 2019;9:7384-402
31. Li GY, Wang W, Sun JY, Xin B, Zhang X, Wang T. et al. Long non-coding RNAs AC026904.1 and UCA1: a "one-two punch" for TGF-beta-induced SNAI2 activation and epithelial-mesenchymal transition in breast cancer. Theranostics. 2018;8:2846-61
32. Luo L, Tang H, Ling L, Li N, Jia X, Zhang Z. et al. LINC01638 lncRNA activates MTDH-Twist1 signaling by preventing SPOP-mediated c-Myc degradation in triple-negative breast cancer. Oncogene. 2018;37:6166-79
33. Jia X, Shi L, Wang X, Luo L, Ling L, Yin J. et al. KLF5 regulated lncRNA RP1 promotes the growth and metastasis of breast cancer via repressing p27kip1 translation. Cell Death Dis. 2019;10:373
34. Wang PS, Chou CH, Lin CH, Yao YC, Cheng HC, Li HY. et al. A novel long non-coding RNA linc-ZNF469-3 promotes lung metastasis through miR-574-5p-ZEB1 axis in triple negative breast cancer. Oncogene. 2018;37:4662-78
35. Hou P, Zhao Y, Li Z, Yao R, Ma M, Gao Y. et al. LincRNA-ROR induces epithelial-to-mesenchymal transition and contributes to breast cancer tumorigenesis and metastasis. Cell Death Dis. 2014;5:e1287
36. Zhou M, Hou Y, Yang G, Zhang H, Tu G, Du YE. et al. LncRNA-Hh Strengthen Cancer Stem Cells Generation in Twist-Positive Breast Cancer via Activation of Hedgehog Signaling Pathway. Stem Cells. 2016;34:55-66
37. Tang L, Chen Y, Chen H, Jiang P, Yan L, Mo D. et al. DCST1-AS1 Promotes TGF-beta-Induced Epithelial-Mesenchymal Transition and Enhances Chemoresistance in Triple-Negative Breast Cancer Cells via ANXA1. Front Oncol. 2020;10:280
38. Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126:663-76
39. Okita K, Ichisaka T, Yamanaka S. Generation of germline-competent induced pluripotent stem cells. Nature. 2007;448:313-7
40. Takahashi K, Tanabe K, Ohnuki M, Narita M, Ichisaka T, Tomoda K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell. 2007;131:861-72
41. Yu J, Vodyanik MA, Smuga-Otto K, Antosiewicz-Bourget J, Frane JL, Tian S. et al. Induced pluripotent stem cell lines derived from human somatic cells. Science. 2007;318:1917-20
42. Yu F, Li J, Chen H, Fu J, Ray S, Huang S. et al. Kruppel-like factor 4 (KLF4) is required for maintenance of breast cancer stem cells and for cell migration and invasion. Oncogene. 2011;30:2161-72
43. Leis O, Eguiara A, Lopez-Arribillaga E, Alberdi MJ, Hernandez-Garcia S, Elorriaga K. et al. Sox2 expression in breast tumours and activation in breast cancer stem cells. Oncogene. 2012;31:1354-65
44. Thomas ML, de Antueno R, Coyle KM, Sultan M, Cruickshank BM, Giacomantonio MA. et al. Citral reduces breast tumor growth by inhibiting the cancer stem cell marker ALDH1A3. Mol Oncol. 2016;10:1485-96
45. Zhou J, Ng SB, Chng WJ. LIN28/LIN28B: an emerging oncogenic driver in cancer stem cells. Int J Biochem Cell Biol. 2013;45:973-8
46. Xu Z, Liu C, Zhao Q, Lu J, Ding X, Luo A. et al. Long non-coding RNA CCAT2 promotes oncogenesis in triple-negative breast cancer by regulating stemness of cancer cells. Pharmacol Res. 2020;152:104628
47. Peng F, Li TT, Wang KL, Xiao GQ, Wang JH, Zhao HD. et al. H19/let-7/LIN28 reciprocal negative regulatory circuit promotes breast cancer stem cell maintenance. Cell Death Dis. 2017;8:e2569
48. Tu Z, Schmollerl J, Cuiffo BG, Karnoub AE. Microenvironmental Regulation of Long Noncoding RNA LINC01133 Promotes Cancer Stem Cell-Like Phenotypic Traits in Triple-Negative Breast Cancers. Stem Cells. 2019;37:1281-92
49. Han L, Yan Y, Zhao L, Liu Y, Lv X, Zhang L. et al. LncRNA HOTTIP facilitates the stemness of breast cancer via regulation of miR-148a-3p/WNT1 pathway. J Cell Mol Med. 2020;24:6242-52
50. Vidovic D, Huynh TT, Konda P, Dean C, Cruickshank BM, Sultan M. et al. ALDH1A3-regulated long non-coding RNA NRAD1 is a potential novel target for triple-negative breast tumors and cancer stem cells. Cell Death Differ. 2020;27:363-78
51. Kalbasi A, Ribas A. Tumour-intrinsic resistance to immune checkpoint blockade. Nat Rev Immunol. 2020;20:25-39
52. Huang D, Chen J, Yang L, Ouyang Q, Li J, Lao L. et al. NKILA lncRNA promotes tumor immune evasion by sensitizing T cells to activation-induced cell death. Nat Immunol. 2018;19:1112-25
53. Colditz GA, Rosner BA, Chen WY, Holmes MD, Hankinson SE. Risk factors for breast cancer according to estrogen and progesterone receptor status. J Natl Cancer Inst. 2004;96:218-28
54. MacMahon B, Cole P, Lin TM, Lowe CR, Mirra AP, Ravnihar B. et al. Age at first birth and breast cancer risk. Bull World Health Organ. 1970;43:209-21
55. Brisken C. Progesterone signalling in breast cancer: a neglected hormone coming into the limelight. Nat Rev Cancer. 2013;13:385-96
56. Osborne CK, Schiff R. Mechanisms of endocrine resistance in breast cancer. Annu Rev Med. 2011;62:233-47
57. Frasor J, Stossi F, Danes JM, Komm B, Lyttle CR, Katzenellenbogen BS. Selective estrogen receptor modulators: discrimination of agonistic versus antagonistic activities by gene expression profiling in breast cancer cells. Cancer Res. 2004;64:1522-33
58. Jordan VC. The science of selective estrogen receptor modulators: concept to clinical practice. Clin Cancer Res. 2006;12:5010-3
59. Guan J, Zhou W, Hafner M, Blake RA, Chalouni C, Chen IP. et al. Therapeutic Ligands Antagonize Estrogen Receptor Function by Impairing Its Mobility. Cell. 2019;178:949-63 e18
60. Ghosh D, Lo J, Egbuta C. Recent Progress in the Discovery of Next Generation Inhibitors of Aromatase from the Structure-Function Perspective. J Med Chem. 2016;59:5131-48
61. Mitobe Y, Ikeda K, Suzuki T, Takagi K, Kawabata H, Horie-Inoue K. et al. ESR1-Stabilizing Long Noncoding RNA TMPO-AS1 Promotes Hormone-Refractory Breast Cancer Progression. Mol Cell Biol. 2019 39
62. Ma T, Liang Y, Li Y, Song X, Zhang N, Li X. et al. LncRNA LINP1 confers tamoxifen resistance and negatively regulated by ER signaling in breast cancer. Cell Signal. 2020;68:109536
63. Niknafs YS, Han S, Ma T, Speers C, Zhang C, Wilder-Romans K. et al. The lncRNA landscape of breast cancer reveals a role for DSCAM-AS1 in breast cancer progression. Nat Commun. 2016;7:12791
64. Gu J, Wang Y, Wang X, Zhou D, Shao C, Zhou M. et al. Downregulation of lncRNA GAS5 confers tamoxifen resistance by activating miR-222 in breast cancer. Cancer Lett. 2018;434:1-10
65. Li X, Wu Y, Liu A, Tang X. Long non-coding RNA UCA1 enhances tamoxifen resistance in breast cancer cells through a miR-18a-HIF1alpha feedback regulatory loop. Tumour Biol. 2016;37:14733-43
66. Liu Y, Li M, Yu H, Piao H. lncRNA CYTOR promotes tamoxifen resistance in breast cancer cells via sponging miR125a5p. Int J Mol Med. 2020;45:497-509
67. Ma Y, Bu D, Long J, Chai W, Dong J. LncRNA DSCAM-AS1 acts as a sponge of miR-137 to enhance Tamoxifen resistance in breast cancer. J Cell Physiol. 2019;234:2880-94
68. Zhang HY, Liang F, Zhang JW, Wang F, Wang L, Kang XG. Effects of long noncoding RNA-ROR on tamoxifen resistance of breast cancer cells by regulating microRNA-205. Cancer Chemother Pharmacol. 2017;79:327-37
69. Kim CY, Oh JH, Lee JY, Kim MH. The LncRNA HOTAIRM1 Promotes Tamoxifen Resistance by Mediating HOXA1 Expression in ER+ Breast Cancer Cells. J Cancer. 2020;11:3416-23
70. Oh DY, Bang YJ. HER2-targeted therapies - a role beyond breast cancer. Nat Rev Clin Oncol. 2020;17:33-48
71. Swain SM, Kim SB, Cortes J, Ro J, Semiglazov V, Campone M. et al. Pertuzumab, trastuzumab, and docetaxel for HER2-positive metastatic breast cancer (CLEOPATRA study): overall survival results from a randomised, double-blind, placebo-controlled, phase 3 study. Lancet Oncol. 2013;14:461-71
72. Verma S, Miles D, Gianni L, Krop IE, Welslau M, Baselga J. et al. Trastuzumab emtansine for HER2-positive advanced breast cancer. N Engl J Med. 2012;367:1783-91
73. Geyer CE, Forster J, Lindquist D, Chan S, Romieu CG, Pienkowski T. et al. Lapatinib plus capecitabine for HER2-positive advanced breast cancer. N Engl J Med. 2006;355:2733-43
74. Chan A, Delaloge S, Holmes FA, Moy B, Iwata H, Harvey VJ. et al. Neratinib after trastuzumab-based adjuvant therapy in patients with HER2-positive breast cancer (ExteNET): a multicentre, randomised, double-blind, placebo-controlled, phase 3 trial. Lancet Oncol. 2016;17:367-77
75. Han M, Gu Y, Lu P, Li J, Cao H, Li X. et al. Exosome-mediated lncRNA AFAP1-AS1 promotes trastuzumab resistance through binding with AUF1 and activating ERBB2 translation. Mol Cancer. 2020;19:26
76. Zheng Z, Chen M, Xing P, Yan X, Xie B. Increased Expression of Exosomal AGAP2-AS1 (AGAP2 Antisense RNA 1) In Breast Cancer Cells Inhibits Trastuzumab-Induced Cell Cytotoxicity. Med Sci Monit. 2019;25:2211-20
77. Dong H, Wang W, Chen R, Zhang Y, Zou K, Ye M. et al. Exosome-mediated transfer of lncRNASNHG14 promotes trastuzumab chemoresistance in breast cancer. Int J Oncol. 2018;53:1013-26
78. Dong H, Wang W, Mo S, Liu Q, Chen X, Chen R. et al. Long non-coding RNA SNHG14 induces trastuzumab resistance of breast cancer via regulating PABPC1 expression through H3K27 acetylation. J Cell Mol Med. 2018;22:4935-47
79. Ren J, Ding L, Zhang D, Shi G, Xu Q, Shen S. et al. Carcinoma-associated fibroblasts promote the stemness and chemoresistance of colorectal cancer by transferring exosomal lncRNA H19. Theranostics. 2018;8:3932-48
80. Dong H, Wang W, Mo S, Chen R, Zou K, Han J. et al. SP1-induced lncRNA AGAP2-AS1 expression promotes chemoresistance of breast cancer by epigenetic regulation of MyD88. J Exp Clin Cancer Res. 2018;37:202
81. Sun Z, Zhang C, Wang T, Shi P, Tian X, Guo Y. Correlation between long non-coding RNAs (lncRNAs) H19 expression and trastuzumab resistance in breast cancer. J Cancer Res Ther. 2019;15:933-40
82. Zhu HY, Bai WD, Ye XM, Yang AG, Jia LT. Long non-coding RNA UCA1 desensitizes breast cancer cells to trastuzumab by impeding miR-18a repression of Yes-associated protein 1. Biochem Biophys Res Commun. 2018;496:1308-13
83. Li W, Zhai L, Wang H, Liu C, Zhang J, Chen W. et al. Downregulation of LncRNA GAS5 causes trastuzumab resistance in breast cancer. Oncotarget. 2016;7:27778-86
84. Fritsch C, Huang A, Chatenay-Rivauday C, Schnell C, Reddy A, Liu M. et al. Characterization of the novel and specific PI3Kalpha inhibitor NVP-BYL719 and development of the patient stratification strategy for clinical trials. Mol Cancer Ther. 2014;13:1117-29
85. First PI3K Inhibitor for Breast Cancer. JAMA. 2019; 322: 19.
86. Chien AJ, Tripathy D, Albain KS, Symmans WF, Rugo HS, Melisko ME. et al. MK-2206 and Standard Neoadjuvant Chemotherapy Improves Response in Patients With Human Epidermal Growth Factor Receptor 2-Positive and/or Hormone Receptor-Negative Breast Cancers in the I-SPY 2 Trial. J Clin Oncol. 2020;38:1059-69
87. Schmid P, Abraham J, Chan S, Wheatley D, Brunt AM, Nemsadze G. et al. Capivasertib Plus Paclitaxel Versus Placebo Plus Paclitaxel As First-Line Therapy for Metastatic Triple-Negative Breast Cancer: The PAKT Trial. J Clin Oncol. 2020;38:423-33
88. Kim SB, Dent R, Im SA, Espie M, Blau S, Tan AR. et al. Ipatasertib plus paclitaxel versus placebo plus paclitaxel as first-line therapy for metastatic triple-negative breast cancer (LOTUS): a multicentre, randomised, double-blind, placebo-controlled, phase 2 trial. Lancet Oncol. 2017;18:1360-72
89. Lin A, Hu Q, Li C, Xing Z, Ma G, Wang C. et al. The LINK-A lncRNA interacts with PtdIns(3,4,5)P3 to hyperactivate AKT and confer resistance to AKT inhibitors. Nat Cell Biol. 2017;19:238-51
90. Koirala P, Huang J, Ho TT, Wu F, Ding X, Mo YY. LncRNA AK023948 is a positive regulator of AKT. Nat Commun. 2017;8:14422
91. Zhou Q, Guo J, Huang W, Yu X, Xu C, Long X. Linc-ROR promotes the progression of breast cancer and decreases the sensitivity to rapamycin through miR-194-3p targeting MECP2. Mol Oncol. 2020
92. Gao X, Qin T, Mao J, Zhang J, Fan S, Lu Y. et al. PTENP1/miR-20a/PTEN axis contributes to breast cancer progression by regulating PTEN via PI3K/AKT pathway. J Exp Clin Cancer Res. 2019;38:256
93. Shi X, Tang X, Su L. Overexpression of Long Noncoding RNA PTENP1 Inhibits Cell Proliferation and Migration via Suppression of miR-19b in Breast Cancer Cells. Oncol Res. 2018;26:869-78
94. Meseure D, Vacher S, Lallemand F, Alsibai KD, Hatem R, Chemlali W. et al. Prognostic value of a newly identified MALAT1 alternatively spliced transcript in breast cancer. Br J Cancer. 2016;114:1395-404
95. Huang YS, Chang CC, Lee SS, Jou YS, Shih HM. Xist reduction in breast cancer upregulates AKT phosphorylation via HDAC3-mediated repression of PHLPP1 expression. Oncotarget. 2016;7:43256-66
96. O'Leary B, Finn RS, Turner NC. Treating cancer with selective CDK4/6 inhibitors. Nat Rev Clin Oncol. 2016;13:417-30
97. Finn RS, Martin M, Rugo HS, Jones S, Im SA, Gelmon K. et al. Palbociclib and Letrozole in Advanced Breast Cancer. N Engl J Med. 2016;375:1925-36
98. Hortobagyi GN, Stemmer SM, Burris HA, Yap YS, Sonke GS, Paluch-Shimon S. et al. Ribociclib as First-Line Therapy for HR-Positive, Advanced Breast Cancer. N Engl J Med. 2016;375:1738-48
99. Sledge GW Jr, Toi M, Neven P, Sohn J, Inoue K, Pivot X. et al. MONARCH 2: Abemaciclib in Combination With Fulvestrant in Women With HR+/HER2- Advanced Breast Cancer Who Had Progressed While Receiving Endocrine Therapy. J Clin Oncol. 2017;35:2875-84
100. Quereda V, Bayle S, Vena F, Frydman SM, Monastyrskyi A, Roush WR. et al. Therapeutic Targeting of CDK12/CDK13 in Triple-Negative Breast Cancer. Cancer Cell. 2019;36:545-58 e7
101. Spring LM, Wander SA, Andre F, Moy B, Turner NC, Bardia A. Cyclin-dependent kinase 4 and 6 inhibitors for hormone receptor-positive breast cancer: past, present, and future. Lancet. 2020;395:817-27
102. Dean JL, Thangavel C, McClendon AK, Reed CA, Knudsen ES. Therapeutic CDK4/6 inhibition in breast cancer: key mechanisms of response and failure. Oncogene. 2010;29:4018-32
103. Caldon CE, Sergio CM, Kang J, Muthukaruppan A, Boersma MN, Stone A. et al. Cyclin E2 overexpression is associated with endocrine resistance but not insensitivity to CDK2 inhibition in human breast cancer cells. Mol Cancer Ther. 2012;11:1488-99
104. Feng T, Shao F, Wu Q, Zhang X, Xu D, Qian K. et al. miR-124 downregulation leads to breast cancer progression via LncRNA-MALAT1 regulation and CDK4/E2F1 signal activation. Oncotarget. 2016;7:16205-16
105. Fan S, Yang Z, Ke Z, Huang K, Liu N, Fang X. et al. Downregulation of the long non-coding RNA TUG1 is associated with cell proliferation, migration, and invasion in breast cancer. Biomed Pharmacother. 2017;95:1636-43
106. Deng X, Zhao Y, Wu X, Song G. Upregulation of CCAT2 promotes cell proliferation by repressing the P15 in breast cancer. Biomed Pharmacother. 2017;91:1160-6
107. Yuan H, Qin Y, Zeng B, Feng Y, Li Y, Xiang T. et al. Long noncoding RNA LINC01089 predicts clinical prognosis and inhibits cell proliferation and invasion through the Wnt/beta-catenin signaling pathway in breast cancer. Onco Targets Ther. 2019;12:4883-95
108. Farmer H, McCabe N, Lord CJ, Tutt AN, Johnson DA, Richardson TB. et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature. 2005;434:917-21
109. Noordermeer SM, van Attikum H. PARP Inhibitor Resistance: A Tug-of-War in BRCA-Mutated Cells. Trends Cell Biol. 2019;29:820-34
110. Wan G, Hu X, Liu Y, Han C, Sood AK, Calin GA. et al. A novel non-coding RNA lncRNA-JADE connects DNA damage signalling to histone H4 acetylation. EMBO J. 2013;32:2833-47
111. Shahabi S, Kumaran V, Castillo J, Cong Z, Nandagopal G, Mullen DJ. et al. LINC00261 Is an Epigenetically Regulated Tumor Suppressor Essential for Activation of the DNA Damage Response. Cancer Res. 2019;79:3050-62
112. Tracy KM, Tye CE, Ghule PN, Malaby HLH, Stumpff J, Stein JL. et al. Mitotically-Associated lncRNA (MANCR) Affects Genomic Stability and Cell Division in Aggressive Breast Cancer. Mol Cancer Res. 2018;16:587-98
113. Gazy I, Zeevi DA, Renbaum P, Zeligson S, Eini L, Bashari D. et al. TODRA, a lncRNA at the RAD51 Locus, Is Oppositely Regulated to RAD51, and Enhances RAD51-Dependent DSB (Double Strand Break) Repair. PLoS One. 2015;10:e0134120
114. Vousden KH, Lane DP. p53 in health and disease. Nat Rev Mol Cell Biol. 2007;8:275-83
115. Hu WL, Jin L, Xu A, Wang YF, Thorne RF, Zhang XD. et al. GUARDIN is a p53-responsive long non-coding RNA that is essential for genomic stability. Nat Cell Biol. 2018;20:492-502
116. Zhang Y, He Q, Hu Z, Feng Y, Fan L, Tang Z. et al. Long noncoding RNA LINP1 regulates repair of DNA double-strand breaks in triple-negative breast cancer. Nat Struct Mol Biol. 2016;23:522-30
117. Chu W, Zhang X, Qi L, Fu Y, Wang P, Zhao W. et al. The EZH2-PHACTR2-AS1-Ribosome Axis induces Genomic Instability and Promotes Growth and Metastasis in Breast Cancer. Cancer Res. 2020;80:2737-50
118. Momparler RL, Karon M, Siegel SE, Avila F. Effect of adriamycin on DNA, RNA, and protein synthesis in cell-free systems and intact cells. Cancer Res. 1976;36:2891-5
119. Pang B, Qiao X, Janssen L, Velds A, Groothuis T, Kerkhoven R. et al. Drug-induced histone eviction from open chromatin contributes to the chemotherapeutic effects of doxorubicin. Nat Commun. 2013;4:1908
120. Yvon AM, Wadsworth P, Jordan MA. Taxol suppresses dynamics of individual microtubules in living human tumor cells. Mol Biol Cell. 1999;10:947-59
121. Brito DA, Yang Z, Rieder CL. Microtubules do not promote mitotic slippage when the spindle assembly checkpoint cannot be satisfied. J Cell Biol. 2008;182:623-9
122. Slamon DJ, Leyland-Jones B, Shak S, Fuchs H, Paton V, Bajamonde A. et al. Use of chemotherapy plus a monoclonal antibody against HER2 for metastatic breast cancer that overexpresses HER2. N Engl J Med. 2001;344:783-92
123. Sosnik A. Reversal of multidrug resistance by the inhibition of ATP-binding cassette pumps employing “Generally Recognized as Safe” (GRAS) nanopharmaceuticals: A review. Adv Drug Deliv Rev. 2013;65:1828-51
124. Wang R, Zhang T, Yang Z, Jiang C, Seng J. Long non-coding RNA FTH1P3 activates paclitaxel resistance in breast cancer through miR-206/ABCB1. J Cell Mol Med. 2018;22:4068-75
125. Chang L, Hu Z, Zhou Z, Zhang H. Linc00518 Contributes to Multidrug Resistance Through Regulating the MiR-199a/MRP1 Axis in Breast Cancer. Cell Physiol Biochem. 2018;48:16-28
126. Yao N, Fu Y, Chen L, Liu Z, He J, Zhu Y. et al. Long non-coding RNA NONHSAT101069 promotes epirubicin resistance, migration, and invasion of breast cancer cells through NONHSAT101069/miR-129-5p/Twist1 axis. Oncogene. 2019;38:7216-33
127. Zheng P, Dong L, Zhang B, Dai J, Zhang Y, Wang Y. et al. Long noncoding RNA CASC2 promotes paclitaxel resistance in breast cancer through regulation of miR-18a-5p/CDK19. Histochem Cell Biol. 2019;152:281-91
128. Pan Y, Pan Y, Cheng Y, Yang F, Yao Z, Wang O. Knockdown of LncRNA MAPT-AS1 inhibites proliferation and migration and sensitizes cancer cells to paclitaxel by regulating MAPT expression in ER-negative breast cancers. Cell Biosci. 2018;8:7
129. Gu M, Zheng W, Zhang M, Dong X, Zhao Y, Wang S. et al. LncRNA NONHSAT141924 promotes paclitaxel chemotherapy resistance through p-CREB/Bcl-2 apoptosis signaling pathway in breast cancer. J Cancer. 2020;11:3645-54
130. Wang X, Pei X, Guo G, Qian X, Dou D, Zhang Z. et al. Exosome-mediated transfer of long noncoding RNA H19 induces doxorubicin resistance in breast cancer. J Cell Physiol. 2020
131. Wang Y, Zhou P, Li P, Yang F, Gao XQ. Long non-coding RNA H19 regulates proliferation and doxorubicin resistance in MCF-7 cells by targeting PARP1. Bioengineered. 2020;11:536-46
132. Xiu DH, Liu GF, Yu SN, Li LY, Zhao GQ, Liu L. et al. Long non-coding RNA LINC00968 attenuates drug resistance of breast cancer cells through inhibiting the Wnt2/beta-catenin signaling pathway by regulating WNT2. J Exp Clin Cancer Res. 2019;38:94
133. Chen Z, Pan T, Jiang D, Jin L, Geng Y, Feng X. et al. The lncRNA-GAS5/miR-221-3p/DKK2 Axis Modulates ABCB1-Mediated Adriamycin Resistance of Breast Cancer via the Wnt/beta-Catenin Signaling Pathway. Mol Ther Nucleic Acids. 2020;19:1434-48
134. Wang YY, Yan L, Yang S, Xu HN, Chen TT, Dong ZY. et al. Long noncoding RNA AC073284.4 suppresses epithelial-mesenchymal transition by sponging miR-18b-5p in paclitaxel-resistant breast cancer cells. J Cell Physiol. 2019;234:23202-15
135. Adams S, Gatti-Mays ME, Kalinsky K, Korde LA, Sharon E, Amiri-Kordestani L. et al. Current Landscape of Immunotherapy in Breast Cancer: A Review. JAMA Oncol. 2019
136. Hulpke S, Tampe R. The MHC I loading complex: a multitasking machinery in adaptive immunity. Trends Biochem Sci. 2013;38:412-20
137. Salama EA, Adbeltawab RE, El Tayebi HM. XIST and TSIX: Novel Cancer Immune Biomarkers in PD-L1-Overexpressing Breast Cancer Patients. Front Oncol. 2019;9:1459
138. Xing F, Liu Y, Wu SY, Wu K, Sharma S, Mo YY. et al. Loss of XIST in Breast Cancer Activates MSN-c-Met and Reprograms Microglia via Exosomal miRNA to Promote Brain Metastasis. Cancer Res. 2018;78:4316-30
139. Xu N, Wang F, Lv M, Cheng L. Microarray expression profile analysis of long non-coding RNAs in human breast cancer: a study of Chinese women. Biomed Pharmacother. 2015;69:221-7
140. Zhang Y, Li Z, Chen M, Chen H, Zhong Q, Liang L. et al. lncRNA TCL6 correlates with immune cell infiltration and indicates worse survival in breast cancer. Breast Cancer. 2020
141. Thomas PB, Seim I, Jeffery PL, Gahete MD, Maugham M, Crisp GJ. et al. The long non-coding RNA GHSROS facilitates breast cancer cell migration and orthotopic xenograft tumour growth. Int J Oncol. 2019;55:1223-36
142. Xu S, Wan L, Yin H, Xu H, Zheng W, Shen M. et al. Long Noncoding RNA Linc00152 Functions as a Tumor Propellant in Pan-Cancer. Cell Physiol Biochem. 2017;44:2476-90
143. Saadatmand S, Geuzinge HA, Rutgers EJT, Mann RM, de Roy van Zuidewijn DBW, Zonderland HM. et al. MRI versus mammography for breast cancer screening in women with familial risk (FaMRIsc): a multicentre, randomised, controlled trial. Lancet Oncol. 2019;20:1136-47
144. McKinney SM, Sieniek M, Godbole V, Godwin J, Antropova N, Ashrafian H. et al. International evaluation of an AI system for breast cancer screening. Nature. 2020;577:89-94
145. Zackrisson S, Lang K, Rosso A, Johnson K, Dustler M, Fornvik D. et al. One-view breast tomosynthesis versus two-view mammography in the Malmo Breast Tomosynthesis Screening Trial (MBTST): a prospective, population-based, diagnostic accuracy study. Lancet Oncol. 2018;19:1493-503
146. Xu N, Chen F, Wang F, Lu X, Wang X, Lv M. et al. Clinical significance of high expression of circulating serum lncRNA RP11-445H22.4 in breast cancer patients: a Chinese population-based study. Tumour Biol. 2015;36:7659-65
147. Liu YR, Jiang YZ, Xu XE, Yu KD, Jin X, Hu X. et al. Comprehensive transcriptome analysis identifies novel molecular subtypes and subtype-specific RNAs of triple-negative breast cancer. Breast Cancer Res. 2016;18:33
148. Collina F, Aquino G, Brogna M, Cipolletta S, Buonfanti G, De Laurentiis M. et al. LncRNA HOTAIR up-regulation is strongly related with lymph nodes metastasis and LAR subtype of Triple Negative Breast Cancer. J Cancer. 2019;10:2018-24
149. Milevskiy MJ, Al-Ejeh F, Saunus JM, Northwood KS, Bailey PJ, Betts JA. et al. Long-range regulators of the lncRNA HOTAIR enhance its prognostic potential in breast cancer. Hum Mol Genet. 2016;25:3269-83
150. Salvador MA, Wicinski J, Cabaud O, Toiron Y, Finetti P, Josselin E. et al. The histone deacetylase inhibitor abexinostat induces cancer stem cells differentiation in breast cancer with low Xist expression. Clin Cancer Res. 2013;19:6520-31
151. Liu YR, Jiang YZ, Xu XE, Hu X, Yu KD, Shao ZM. Comprehensive Transcriptome Profiling Reveals Multigene Signatures in Triple-Negative Breast Cancer. Clin Cancer Res. 2016;22:1653-62
152. Jiang YZ, Liu YR, Xu XE, Jin X, Hu X, Yu KD. et al. Transcriptome Analysis of Triple-Negative Breast Cancer Reveals an Integrated mRNA-lncRNA Signature with Predictive and Prognostic Value. Cancer Res. 2016;76:2105-14
153. Ingle JN, Xie F, Ellis MJ, Goss PE, Shepherd LE, Chapman JW. et al. Genetic Polymorphisms in the Long Noncoding RNA MIR2052HG Offer a Pharmacogenomic Basis for the Response of Breast Cancer Patients to Aromatase Inhibitor Therapy. Cancer Res. 2016;76:7012-23
154. Botti G, Cantile M. Circulating long non-coding RNAs: could they be a useful tool for cancer therapy monitoring? Expert Rev Anticancer Ther. 2018;18:1167-8
155. Zhang K, Shi H, Xi H, Wu X, Cui J, Gao Y. et al. Genome-Wide lncRNA Microarray Profiling Identifies Novel Circulating lncRNAs for Detection of Gastric Cancer. Theranostics. 2017;7:213-27
156. Zhou Z, Wu Q, Yan Z, Zheng H, Chen CJ, Liu Y. et al. Extracellular RNA in a single droplet of human serum reflects physiologic and disease states. Proc Natl Acad Sci U S A. 2019;116:19200-8
157. Matsui M, Corey DR. Non-coding RNAs as drug targets. Nat Rev Drug Discov. 2017;16:167-79
158. Lennox KA, Behlke MA. Cellular localization of long non-coding RNAs affects silencing by RNAi more than by antisense oligonucleotides. Nucleic Acids Res. 2016;44:863-77
159. Arun G, Diermeier S, Akerman M, Chang KC, Wilkinson JE, Hearn S. et al. Differentiation of mammary tumors and reduction in metastasis upon Malat1 lncRNA loss. Genes Dev. 2016;30:34-51
160. Xiu B, Chi Y, Liu L, Chi W, Zhang Q, Chen J. et al. LINC02273 drives breast cancer metastasis by epigenetically increasing AGR2 transcription. Mol Cancer. 2019;18:187
161. Qiao K, Ning S, Wan L, Wu H, Wang Q, Zhang X. et al. LINC00673 is activated by YY1 and promotes the proliferation of breast cancer cells via the miR-515-5p/MARK4/Hippo signaling pathway. J Exp Clin Cancer Res. 2019;38:418
162. Li M, Li X, Zhuang Y, Flemington EK, Lin Z, Shan B. Induction of a novel isoform of the lncRNA HOTAIR in Claudin-low breast cancer cells attached to extracellular matrix. Mol Oncol. 2017;11:1698-710
163. Wang S, Liang K, Hu Q, Li P, Song J, Yang Y. et al. JAK2-binding long noncoding RNA promotes breast cancer brain metastasis. J Clin Invest. 2017;127:4498-515
164. Kaur H, Arora A, Wengel J, Maiti S. Thermodynamic, counterion, and hydration effects for the incorporation of locked nucleic acid nucleotides into DNA duplexes. Biochemistry. 2006;45:7347-55
165. Owczarzy R, You Y, Groth CL, Tataurov AV. Stability and mismatch discrimination of locked nucleic acid-DNA duplexes. Biochemistry. 2011;50:9352-67
166. Veedu RN, Wengel J. Locked nucleic acids: promising nucleic acid analogs for therapeutic applications. Chem Biodivers. 2010;7:536-42
167. Xing Z, Lin A, Li C, Liang K, Wang S, Liu Y. et al. lncRNA directs cooperative epigenetic regulation downstream of chemokine signals. Cell. 2014;159:1110-25
168. Zheng X, Han H, Liu GP, Ma YX, Pan RL, Sang LJ. et al. LncRNA wires up Hippo and Hedgehog signaling to reprogramme glucose metabolism. EMBO J. 2017;36:3325-35
169. Juan V, Crain C, Wilson C. Evidence for evolutionarily conserved secondary structure in the H19 tumor suppressor RNA. Nucleic Acids Res. 2000;28:1221-7
Author contact
Corresponding authors: Na Xie, Tel.: +86-15884555293, E-mail: naxieedu.cn; Yuan-Yuan Zhang, Tel.: +86-18728495257, E-mail: sarahyyzhangcom.
 Global reach, higher impact
Global reach, higher impact