13.3
Impact Factor
Theranostics 2019; 9(24):7122-7139. doi:10.7150/thno.35729 This issue Cite
Research Paper
Aldehyde dehydrogenase 1A1 confers erlotinib resistance via facilitating the reactive oxygen species-reactive carbonyl species metabolic pathway in lung adenocarcinomas
1. Department of Pharmacology and Chemical Biology, Shanghai Jiao Tong University School of Medicine, Shanghai, 200025, China.
2. Shanghai Collaborative Innovation Center for Translational Medicine, Shanghai, 200025, China
3. State Key Laboratory of Oncogenes and Related Genes, Ren Ji Hospital, School of Medicine, Shanghai Jiao Tong University, Shanghai, 200127, China.
4. Central laboratory, Shanghai Chest Hospital, Shanghai Jiao Tong University, Shanghai, 200030, China; Shanghai Institute of Immunology, Shanghai Jiao Tong University School of Medicine, Shanghai, 200025, China.
5. Department of Pathophysiology, Key Laboratory of Cell Differentiation and Apoptosis of National Ministry of Education, Shanghai Key Laboratory of Tumor Microenvironment and Inflammation, Shanghai Jiao-Tong University School of Medicine, Shanghai 200025, China.
* These authors contributed equally to this study.
Abstract
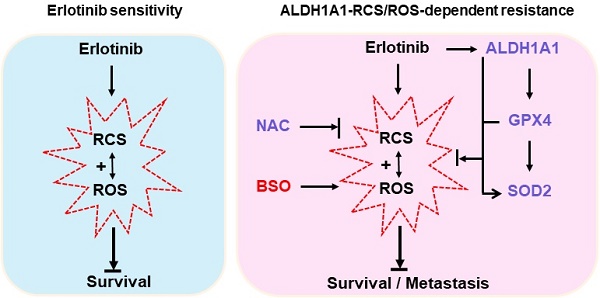
Background: Acquired resistance to epidermal growth factor receptor (EGFR)-tyrosine kinase inhibitors (TKIs) such as erlotinib is a major challenge to achieve an overall clinical benefit of the targeted therapy. Recently, aldehyde dehydrogenase 1 (ALDH1) induction has been found to render lung adenocarcinomas resistant to EGFR-TKIs, and targeting ALDH1A1 becomes a novel strategy to overcome resistance. However, the molecular mechanism underlying such effect remains poorly understood. Methods: Comprehensive assays were performed in a panel of lung adenocarcinoma cell lines and xenografts that acquired resistance to erlotinib. Cancer phenotype was evaluated by cell viability, apoptosis, migration, and epithelial-mesenchymal transition analysis in vitro, tumorsphere formation analysis ex vivo, and tumor growth and dissemination analysis in vivo. Reactive oxygen species (ROS) and reactive carbonyl species (RCS) were detected based on fluorescent oxidation indicator and liquid chromatography coupled to mass spectrometry, respectively. Protein target was suppressed by RNA interference and pharmacological inhibition or ecto-overexpressed by lentivirus-based cloning. Gene promoter activity was measured by dual-luciferase reporting assay. Results: Knockdown or pharmacological inhibition of ALDH1A1 overcame erlotinib resistance in vitro and in vivo. ALDH1A1 overexpression was sufficient to induce erlotinib resistance. Metabolomic analysis demonstrated lower ROS-RCS levels in ALDH1A1-addicted, erlotinib-resistant cells; in line with this, key enzymes for metabolizing ROS and RCS, SOD2 and GPX4, respectively, were upregulated in these cells. Knockdown of SOD2 or GPX4 re-sensitized the resistant cells to erlotinib and the effect was abrogated by ROS-RCS scavenging and mimicked by ROS-RCS induction. The ALDH1A1 overexpressed cells, though resisted erlotinib, were more sensitive to SOD2 or GPX4 knockdown. The ALDH1A1 effect on erlotinib resistance was abrogated by ROS-RCS induction and mimicked by ROS-RCS scavenging. Detection of GPX4 and SOD2 expression and analysis of promoter activities of GPX4 and SOD2 under the condition of suppression or overexpression of ALDH1A1 demonstrated that the RCS-ROS-metabolic pathway was controlled by the ALDH1A1-GPX4-SOD2 axis. The ROS-RCS metabolic dependence mechanism in ALDH1A1-induced resistance was confirmed in vivo. Analysis of public databases showed that in patients undergoing chemotherapy, those with high co-expression of ALDH1A1, GPX4, and SOD2 had a lower probability of survival. Conclusions: ALDH1A1 confers erlotinib resistance by facilitating the ROS-RCS metabolic pathway. ALDH1A1-induced upregulation of SOD2 and GPX4, as well as ALDH1A1 itself, mitigated erlotinib-induced oxidative and carbonyl stress, and imparted the TKI resistance. The elucidation of previously unrecognized metabolic mechanism underlying erlotinib resistance provides new insight into the biology of molecular targeted therapies and help to design improved pharmacological strategies to overcome the drug resistance.
Keywords: ALDH, erlotinib resistance, reactive oxygen species, reactive carbonyl species, lung cancer.
 Global reach, higher impact
Global reach, higher impact