Impact Factor
Theranostics 2018; 8(16):4520-4534. doi:10.7150/thno.25130 This issue Cite
Research Paper
Rewiring of cisplatin-resistant bladder cancer cells through epigenetic regulation of genes involved in amino acid metabolism
1. Departments of Surgery, Cedars-Sinai Medical Center, Los Angeles, CA, USA
2. National Institute of Dental and Craniofacial Research, National Institutes of Health Bethesda, MD, USA
3. Department of Biochemistry and Molecular Medicine, USC Norris Comprehensive Cancer Center, University of Southern California
4. Department of Urology, Keck School of Medicine, University of Southern California, Los Angeles, CA, USA
5. Departments of Biomedical Sciences, Cedars-Sinai Medical Center, Los Angeles, CA, USA
6. Samuel Oschin Comprehensive Cancer Institute, Cedars-Sinai Medical Center, Los Angeles, CA, USA
7. Department of Medicine, University of California Los Angeles, CA, USA
8. Department of Urology, Ga Cheon University College of Medicine, Incheon, Republic of Korea
Abstract
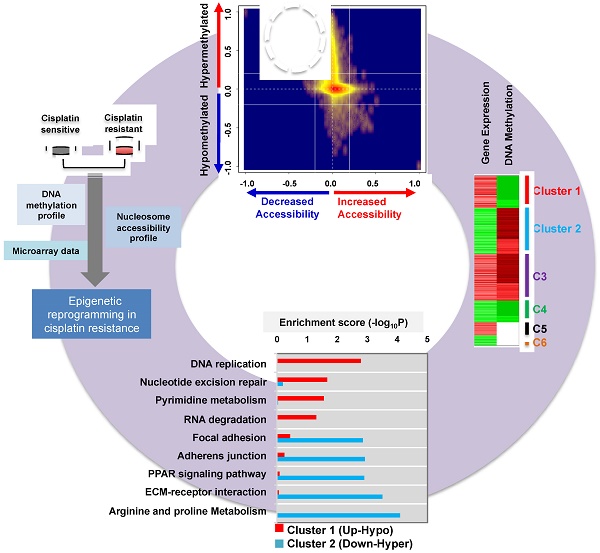
Alterations in DNA methylation are important epigenetic markers in bladder cancer (BC). These epigenome modifications may drive the mechanisms of aggressive chemo-resistant BC. Clinicopathological biomarkers that indicate chemotherapeutic resistance are critical for better assessing treatment strategies for individual patients. Thus, in this study, we aimed to determine whether DNA methylation of certain metabolic enzymes is significantly altered in cisplatin-resistant BC cells.
Methods: To characterize CpG methylation and nucleosome accessibility in cisplatin-resistant BC cells, the Illumina Infinium HM450 DNA methylation assay was performed. Perturbed gene expression was found to be associated with cisplatin resistance, and the biological roles of spermidine/spermine N1-acetyltransferase (SAT1) and argininosuccinate synthase 1 (ASS1) were further studied using qRT-PCR analysis and various cell biology assays, including western blot.
Results: ASS1 and SAT1, genes for amino acid and polyamine metabolism catalysts, respectively, were found to be vastly hypermethylated, resulting in greatly downregulated expression. ASS1 expression is of particular interest because prior studies have demonstrated its potential association with BC stage and recurrence. In regard to chemoresistance, we found that aberrant expression or induced stimulation of SAT1 restored cisplatin sensitivity in the cell culture system. We also found that the addition of exogenous arginine deiminase through administration of ADI-PEG 20 (pegylated arginine deiminase) increased ASS1 expression and enhanced cisplatin's apoptotic effects.
Conclusions: Our study demonstrates a novel mechanistic link between the epigenetic perturbation of SAT1 and ASS1 and cancer metabolism in cisplatin-resistant bladder cancer cells. These findings suggest potential utility of SAT1 and ASS1 as predictive biomarkers in re-sensitizing bladder cancer to chemotherapy and personalizing therapy.
Keywords: SAT1, ASS1, cancer metabolism, metabolomics, DNA methylation, chromatin accessibility, cisplatin resistance
 Global reach, higher impact
Global reach, higher impact