Impact Factor
Theranostics 2017; 7(18):4445-4469. doi:10.7150/thno.18456 This issue Cite
Review
CRISPR Genome Engineering for Human Pluripotent Stem Cell Research
1. Department of Computer Science, Purdue University, West Lafayette, IN 47907, USA;
2. Department of Pathology, University of Washington, Seattle, WA 98195, USA;
3. Department of Bioengineering, University of Washington, Seattle, WA 98195, USA;
4. Institute for Stem Cell and Regenerative Medicine, University of Washington, Seattle, WA 98109, USA;
5. Center for Cardiovascular Biology, University of Washington, Seattle, WA 98109, USA.
Abstract
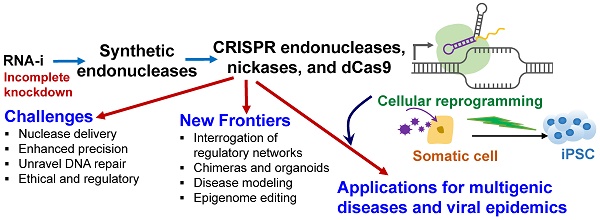
The emergence of targeted and efficient genome editing technologies, such as repurposed bacterial programmable nucleases (e.g., CRISPR-Cas systems), has abetted the development of cell engineering approaches. Lessons learned from the development of RNA-interference (RNA-i) therapies can spur the translation of genome editing, such as those enabling the translation of human pluripotent stem cell engineering. In this review, we discuss the opportunities and the challenges of repurposing bacterial nucleases for genome editing, while appreciating their roles, primarily at the epigenomic granularity. First, we discuss the evolution of high-precision, genome editing technologies, highlighting CRISPR-Cas9. They exist in the form of programmable nucleases, engineered with sequence-specific localizing domains, and with the ability to revolutionize human stem cell technologies through precision targeting with greater on-target activities. Next, we highlight the major challenges that need to be met prior to bench-to-bedside translation, often learning from the path-to-clinic of complementary technologies, such as RNA-i. Finally, we suggest potential bioinformatics developments and CRISPR delivery vehicles that can be deployed to circumvent some of the challenges confronting genome editing technologies en route to the clinic.
 Global reach, higher impact
Global reach, higher impact