Impact Factor
Theranostics 2017; 7(17):4183-4191. doi:10.7150/thno.21299 This issue Cite
Review
Identifying and Characterizing circRNA-Protein Interaction
1. Sunnybrook Research Institute, Sunnybrook Health Sciences Centre, Toronto;
2. Department of Laboratory Medicine and Pathobiology, University of Toronto, Toronto;
3. State Key Laboratory of Applied Microbiology Southern China, Guangdong Provincial Key Laboratory of Microbial Culture Collection and Application, Guangdong Institute of Microbiology, Guangzhou, 510070, China;
4. Yuewei Edible Fungi Technology Co. Ltd., Guangzhou, 510070, China;
5. Atta-ur-Rahman School of Applied Biosciences (ASAB), National University of Sciences and Technology (NUST), H-12 Islamabad, Pakistan;
6. Institute of Medical Science, University of Toronto, Toronto, Canada.
Abstract
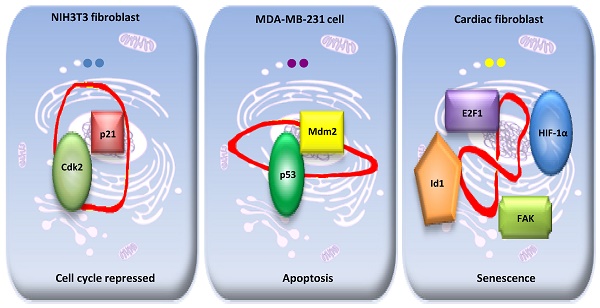
Circular RNAs have been identified as naturally occurring RNAs that are highly represented in the eukaryotic transcriptome. Although a large number of circRNAs have been reported, circRNA functions remain largely unknown. CircRNAs can function as miRNA sponges, thereby reducing their ability to target mRNAs. We hypothesize that circRNAs may bind, store, sort, and sequester proteins to particular subcellular locations, and act as dynamic scaffolding molecules that modulate protein-protein interactions. Here, we review the biological implication and function of circRNA-protein interaction, and reveal a dynamic model of the interaction in various tissues, development stages and physiological conditions. Improved techniques to identify and characterize the dynamic RNA-protein interactions may elucidate the molecular mechanisms associated with the expression and functional diversity of circRNAs.
 Global reach, higher impact
Global reach, higher impact