13.3
Impact Factor
Theranostics 2017; 7(17):4118-4134. doi:10.7150/thno.20112 This issue Cite
Research Paper
DNA Methylation of miR-7 is a Mechanism Involved in Platinum Response through MAFG Overexpression in Cancer Cells
1. Cancer Epigenetics Laboratory, INGEMM, La Paz University Hospital, Madrid, Spain;
2. Biomarkers and Experimental Therapeutics in Cancer, IdiPAZ, Madrid, Spain;
3. Bioinformatics Unit, Centro Nacional de Investigaciones Cardiovasculares Madrid, Spain;
4. Department of Obstetrics and Gynecology, La Paz University Hospital, Madrid, Spain;
5. Department of Pathology, La Paz University Hospital, Madrid, Spain;
6. Genomics Unit, Centro Nacional de Investigaciones Cardiovasculares, Madrid, Spain;
7. Department of Pathology, University Hospital Fundación Jiménez Díaz, Madrid, Spain;
8. Department of Oncology, HM Hospitales, Madrid, Spain;
9. Translational Research in pediatric oncology Hematopoietic transplantation and cell Therapy, IdiPAZ, Madrid, Spain;
10. Human Genetics Group, Spanish National Cancer Research Center (CNIO);
11. Spanish Network on Rare Diseases (CIBERER), Madrid, Spain;
12. Department of Animal Models for Human Diseases, Institute for Biomedical Research CSIC/UAM, CIBER for Rare Diseases, Madrid, Spain.
* Both authors are first authors.
Abstract
One of the major limitations associated with platinum use is the resistance that almost invariably develops in different tumor types. In the current study, we sought to identify epigenetically regulated microRNAs as novel biomarkers of platinum resistance in lung and ovarian cancers, the ones with highest ratios of associated chemo-resistance.
Methods: We combined transcriptomic data from microRNA and mRNA under the influence of an epigenetic reactivation treatment in a panel of four paired cisplatin -sensitive and -resistant cell lines, followed by real-time expression and epigenetic validations for accurate candidate selection in 19 human cancer cell lines. To identify specific candidate genes under miRNA regulation, we assembled “in silico” miRNAs and mRNAs sequences by using ten different algorithms followed by qRT-PCR validation. Functional assays of site-directed mutagenesis and luciferase activity, miRNAs precursor overexpression, silencing by antago-miR and cell viability were performed to confirm their specificity in gene regulation. Results were further explored in 187 primary samples obtained from ovarian tumors and controls.
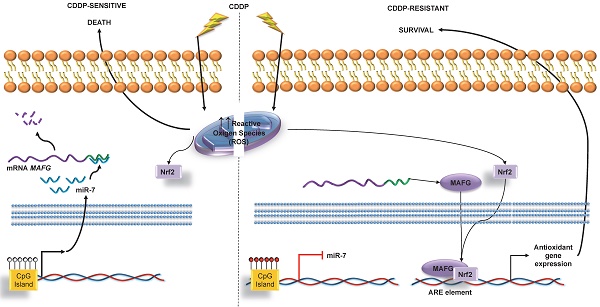
Results: We identified 4 candidates, miR-7, miR-132, miR-335 and miR-148a, which deregulation seems to be a common event in the development of resistance to cisplatin in both tumor types. miR-7 presented specific methylation in resistant cell lines, and was associated with poorer prognosis in ovarian cancer patients. Our experimental results strongly support the direct regulation of MAFG through miR-7 and their involvement in the development of CDDP resistance in human tumor cells.
Conclusion: The basal methylation status of miR-7 before treatment may be a potential clinical epigenetic biomarker, predictor of the chemotherapy outcome to CDDP in ovarian cancer patients. To the best of our knowledge, this is the first report linking the regulation of MAFG by miRNA-7 and its role in chemotherapy response to CDDP. Furthermore, this data highlights the possible role of MAFG as a novel therapeutic target for platinum resistant tumors.
Keywords: miR-7, MAFG, DNA methylation, Cisplatin-resistance, cancer.
 Global reach, higher impact
Global reach, higher impact