Impact Factor
Theranostics 2014; 4(2):119-133. doi:10.7150/thno.7552 This issue Cite
Review
MicroRNA-206: a Promising Theranostic Marker
1. International Clinical Research Center, Department of Cardiovascular Diseases, St. Anne's University Hospital Brno, Brno, Czech Republic
2. Department of Physiology, Faculty of Medicine, Masaryk University, Brno, Czech Republic
3. Department of Pathological Physiology, Faculty of Medicine, Masaryk University, Brno, Czech Republic
4. Molecular Oncology II, Central European Institute of Technology, Brno, Czech Republic
Received 2013-9-1; Accepted 2013-10-14; Published 2014-1-2
Abstract
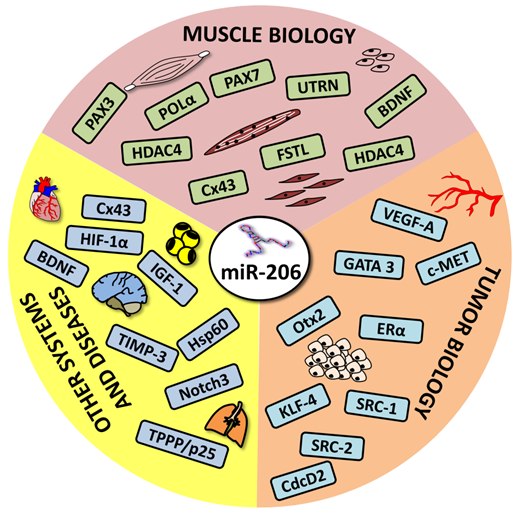
MicroRNAs (miRs) are small non-coding RNAs that negatively regulate gene expression by binding to the 3` untranslated regions (3`UTR) of their target mRNAs. MiRs were shown to play pivotal roles in tissue development and function and are also involved in the pathogenesis of various diseases including cancer.
MicroRNA-206, which belongs to the group of so-called “myomiRs”, is one of the most studied miRs thus far. In addition to being involved in skeletal muscle development and pathology, it has also been established that it is involved in the pathogenesis of numerous diseases including heart failure, chronic obstructive pulmonary disease, Alzheimer's disease and various types of cancers.
The aim of this review is to provide a complex overview of microRNA-206, including regulating its expression, a brief description of its known functions in skeletal muscle and a complex overview of its roles in the biology and pathology of other tissues, emphasizing its significant diagnostic and therapeutic potential.
Keywords: microRNA-206, Theranostic Marker
1. Introduction
MicroRNAs (miRNAs) are small non-coding RNAs, first discovered in the C. elegans nematode worm in 1993 [1]. According to the latest release of miRBase [2] 20 years later, there are now over 30,000 known mature miRNA sequences described in 206 species.
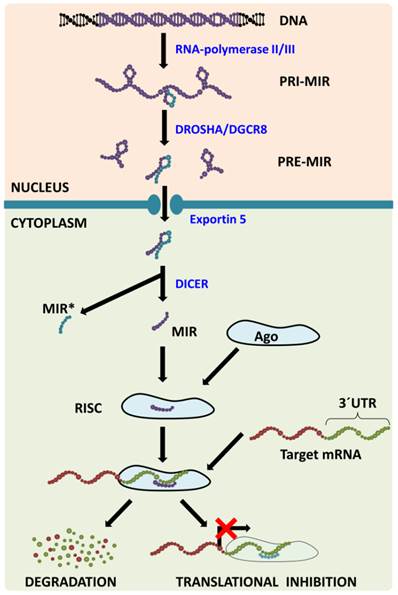
MiRNA Biogenesis (Figure 1) consists of several consecutive steps with the first one being transcribing the miRNA from its corresponding gene using RNA-polymerase II/III into the molecule called primary miRNA (pri-miR). In the nucleus, pri-miR is cleaved with Drosha/DGCR8 complex to form a precursor miRNA (pre-miR) that is transported from the nucleus with exportin‑5 protein. In the cytoplasm, nuclease Dicer cleaves pre-miR, which leads to the generation of miRNA-miRNA* duplex. MiRNA* is usually degraded even though some miRNA* has been observed to function in a similar way to to miRNAs, i.e. they create a complex with Argonaut (Ago) proteins called (mi)RNA-induced silencing complex (RISC or miRISC). The miRNAs in this complex function as negative regulators of gene expression: they bind to the 3´-untranslated regions (3´-UTR) of their target mRNAs (in accordance with base complementarity) and this then leads either to target mRNA degradation or to translational inhibition [3]. One miRNA usually has more than one target mRNA and one mRNA is usually targeted by more than one miRNA [4], which leads to the formation of a huge regulatory network which plays a pivotal role in numerous processes such as embryonic development, normal tissue functions and the pathogenesis of various diseases.
MicroRNA biogenesis pathway. MicroRNAs are transcribed from corresponding genes with RNA-polymerase II/III; the primary transcript is called primary-miRNA (pri-miR). Already in the nucleus, microprocessing complex Drosha/DGCR8 cleaves pri-miR to precursor miR (pre-miR) that is transported from the nucleus with Exportin 5. In the cytoplasm, the Dicer claves pre-miR: the passenger strand (miR*) is consecutively degraded and the mature strand (miR) is incorporated into the RNA-induced silencing complex (RISC). Within the RISC, base pairing between miR and target mRNA 3´untranslated region (3´UTR) occurs leading to target mRNA degradation or translational inhibition.
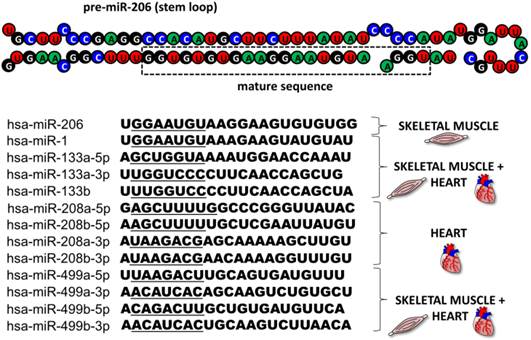
This review focuses on microRNA-206 (miR-206), a vertebrate specific miRNA, which is referred to as a skeletal muscle specific myomiR [5]. This miRNA shares the same seed sequence and at least some targets with miR-1 [2], which also belongs to the myomiR family along with miR-133a, miR-133b, miR-208a, miR-208b and miR-499 ([6], Figure 2); however miR-1 is expressed in both skeletal and cardiac muscles [7]. MiR-206 is being intensively studied in various contexts: skeletal muscle development [8, 9], function and pathology [5, 9], heart failure [10], various types of cancers [11-13] and other diseases [14-16]. The aim of this review is to introduce miR-206, provide a short description of its relevance in skeletal muscle biology as well as offering a more detailed description of this miRNA's role in other conditions while placing special emphasis on its theranostic potential.
MyomiR family. The figure depicts the structure of a pre-miR-206 stem loop with the mature sequence indicated in a rectangle. Under the structure, there are sequences of all miRNAs from the human myomiR family with their seed sequences underlined. Next to the sequences, the miRNAs' tissue specificity is visualized.
2. Regulation of microRNA-206 expression
The gene for human miR-206 (hsa-miR-206) is localized on chromosome 6 in a bicistronic cluster together with the gene for miR-133b [2]. The expression of these microRNAs can be regulated on different levels including chromatin remodeling, transcriptional and posttranscriptional regulation and is also under endocrinological and environmental control.
The chromatin remodeling enzyme, high mobility group box 1 protein (HMGB1), has previously been shown to increase miR-206 levels when injected into the chronically failing hearts of experimental animals [10]. In another cell type, the Th17-lymphocyte, it was reported that miR-206 and miR-133b are co-transcribed together with interleukin (IL)-17 when IL-23 is used [17]. This co-transcription is specific for T-lymphocytes (IL-17 is not detectable in skeletal muscle even after IL-23 stimulation) and may be affected both by chromatin remodeling, since the IL-17 gene is located near the genes for miR-206/133b, and transcription factors (TFs) specific for IL-17, e.g. STAT3 [17].
Various TFs important for skeletal muscle development have been shown to regulate miR-206 expression during skeletal muscle development; likewise, the proper spatiotemporal expression of these TFs is critical to successfully complete myogenesis [18]. One of the strongest inducers of myogenesis, and therefore also of mir-206, is Myf5 [19], which acts through MyoD [20-22] and myogenin [19] with MEF2C enhancement [23]. Negative regulation of miR-206 expression during myogenesis is carried out via myostatin, since myostatin knockout caused upregulation of all myomiRs in experimental animals [24]. Whether this regulation is direct or indirect still remains to be determined. The importance of proper myostatin-mediated negative myogenesis regulation was previously documented on Texel sheep where just one SNP in the 3'UTR of myostatin created a binding site for miR-1/206, resulting in these animals' muscular phenotype [25]. Various autoregulatory loops which add robustness to this TF network have also been described, one of them involving Notch3 protein [23]. Notch3 expression is increased by MyoD at the beginning of muscle differentiation, thus ensuring MEF2C inhibition, which is important in the final stages of myogenesis. When cells are ready to finish the differentiation process, miR-1/206 targets Notch3, thereby inhibiting its function [23].
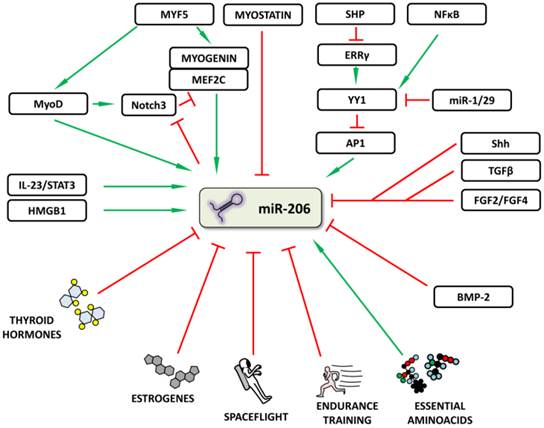
In addition to the above mentioned TFs, other molecules and various signal pathways have also been shown to increase or decrease miR-206 expression in various tissues (Figure 3). Song et al. [26] described a “dual inhibitory mechanism”, in which small heterodimer partner (SHP), a known nuclear transcriptional co-repressor, plays a role. In this model, SHP inhibits the expression of estrogen-related receptor γ (ERRγ) which is responsible for Ying Yang 1 (YY1) TF expression. A decrease in YY1 leads to the activation of TF activator protein 1 (AP1) and to an increase in miR-206 expression [26]. A study by Lu et al. [27] lists other significant players involved in this regulatory mechanism: NFκB, miR-29 and miR-1. NFκB stimulates and miR-1/29 inhibits YY1 signaling, thus inhibiting or activating miR-206 expression, respectively [27]. Other signals inhibiting miR-206 expression include e.g. Sonic Hedgehog (Shh) [28], fibroblast growth factors 2 and 4 (FGF2/FGF4) [29] signaling pathways and tumor growth factor β (TGFβ) [30].
Regulation of microRNA-206 expression. The figure summarizes how miR-206 expression may be regulated or influenced. Red (blunt) arrows indicate inhibition/downregulation and green (sharp) arrows indicate activation/upregulation. Abbreviations are explained in the text.
The participation of thyroid hormones in the miR-206 regulation was described in mouse liver and corresponding cell line models [31] and in human skeletal muscle in patients with hypothyroidism [32]. Thyroid hormones and their substituents decrease miR-206 levels and hypothyroidism increases its expression [31, 32]; accordingly, miR-206 is also proposed to serve as an important mediator of thyroid hormone function since some of the miR-206 targets (e.g. Mup1 and Gpd2) were previously shown to be regulated by thyroid hormones [31]. Additional hormones to be considered in miR-206 regulation are estrogens, as shown in human breast cancer [33], where miR-206 further targets estrogen receptor α (ERα), i.e. one of the factors responsible for cancer cells' ER-negative phenotype and an element affecting responsiveness to hormonal therapy [33].
Focusing on posttranscriptional regulation, bone morphogenetic protein 2 (BMP-2) was shown to inhibit pri-miR-206 to mature miR-206 processing [34]. BMP-2 has previously been shown to cause C2C12 myoblast cell lineage differentiation into osteoblasts when supplemented [35]; this was explained by its inhibition of MyoD and myogenin activity [36]. This study therefore further expands our understanding of the lineage switch process with BMP-2 inhibiting miR-206, myoD and myogenin activity, resulting in the inhibition of myogenic differentiation.
Last but not least, the effect of environmental factors on miR-206 expression needs to be mentioned. Essential aminoacids are known to be important for muscle growth and as such stimulate muscle protein synthesis - along with pri-miR-206 levels [37]. Physical exercise was repeatedly shown not to the affect the circulating miR-206 levels [38, 39], which may become an advantage when monitoring the course of muscle dystrophies because circulating levels of currently used markers (e.g. creatinkinase) are affected [40, 41]. Endurance training did not significantly affect the levels of circulating miR-206 either; however, the basal expression of miR-206 in muscle tissue was significantly lower after 12 weeks and it returned to the pre-training values following 2 weeks of withdrawn training [42]. Interestingly, spaceflight has been shown to decrease miR-206 levels [43]. These results indicate that miR-206 is probably not involved in short-term muscle adaptation to physical exercise; however, everyday muscle activity is needed for its sustained expression and this basal expression may be reduced if these stimuli are either withdrawn (spaceflight) or chronically increased (endurance training).
3. ROLES FOR MicroRNA-206 IN SKELETAL MUSCLE
3.1 Myogenesis and muscle regeneration (Figure 4)
Myogenesis is a complex process during which muscle progenitor cells leave the somites and migrate to final body locations where they proliferate and differentiate in order to create skeletal muscle [18]. MiR-206 is currently considered an important part of the myogenesis process in humans [44], mice [45], zebrafish [46], pigs [47, 48] and even common carp [49]. It can be specifically detected in the somites, with the highest expression in mouse embryos being detected 11.5-12.5 days post conception and tending to decrease from the 3rd day after birth [50]. If miR-206 and other miRNAs levels decrease due to Dicer knockout (this knockout results in global downregulation of most miRNAs), mice die prenatally [51]. If muscle-specific Dicer knockout is performed, it results in skeletal muscle hypoplasia and defective fiber morphology indicating the important role of microRNAs in muscle development [52].
Roles of microRNA-206 in muscle development and regeneration. MiR-206 involvement in muscle development with putative targets shown above black arrows. A comprehensive explanation may be found in the text.
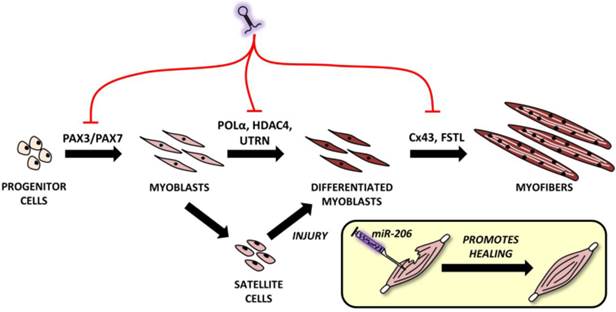
The mouse myoblast cell line (C2C12) is the most commonly used cell line for the study of myogenesis. When C2C12 cells are cultured under high serum conditions, they remain in a proliferating state; however, if they are switched to low serum conditions, they begin to differentiate and fuse into myofibers [53]. The importance of miR-206 in myogenesis is highlighted by a study by Kim et al. [45]. This study indicated that even in stable serum conditions, adding miR-206 induces C2C12 cell differentiation and vice versa - if miR-206 is blocked and serum conditions are changed, no differentiation occurs [45].
Proper miR-206 downregulation and upregulation during different stages of myogenesis is crucial since it targets key molecules like paired box 3 (Pax3) and Pax7, both of which are important TFs for muscle progenitor cell migration and proliferation, respectively [18]. At the right moment, miR-206 needs to be upregulated (via the regulatory pathways described above and including Myf5, MyoD and others') in order to initiate the muscle differentiation program. Inhibiting miR-206 expression using antagomiRs results in a delayed differentiation onset and improper migration of muscle progenitors [54]. Later, during myogenesis, miR-206 targets various genes enabling early and - subsequently - terminal differentiation [9], e.g. DNA-polymerase α (POL α) [45], histon deacetylase 4 (HDAC4) [30], utrophin [21] and connexin 43 (Cx43) genes [55]. Inhibiting POLα is important to terminate the proliferation program [45] at the beginning of differentiation, which is promoted by removing the inhibitory effects of HDAC4 on MEF2C [30]. The downregulation of utrophin seems to be important later in the myogenesis process [21] while follistatin [21] and Cx43 downregulation [55] play an important role in myocyte fusion into myofibers. This process naturally involves additional miRNAs, but a detailed description is clearly outside the scope of this review.
During myogenesis, part of the myoblasts enter quiescence, locate themselves between the basal membrane and sarcolemma [56] and remain in a self-renewing state which maintains their ability to enter the muscle differentiation program in the future if the muscle is injured [18]. In this case, miR-206 levels in SCs are downregulated, thus enabling Pax7 to increase while causing SCs proliferation [57]. After a short time, miR-206 levels return to the pre-injury state and SCs differentiate to replace the damaged rhabdomyocytes. When miR-206 is artificially added to injured skeletal muscle, it promotes muscle healing as shown in rat models in a study conducted by Nakasa et al. [58].
3.2 Innervation, reinnervation and amyotrophic lateral sclerosis (ALS)
In order to properly perform its function, skeletal muscle must be properly innervated; during innervation, skeletal muscle tissue cooperates with axons in order to create neuromuscular junctions (NMJ) (Figure 5). MiR-206 levels were found to be increased in NMJ [59], facilitating mutual communication between muscle fibers and motoneuron axons by targeting various molecules including brain-derived neurotropic factor (BDNF) and its receptor (p75NTR) [60], HDAC4 [59] and MEF2 [61].
MicroRNA-206 involvement in neuromuscular junctions function. The figure depicts neuromuscular junction (NMJ) with the neural cell marked in blue and muscle cell in red. During transmission (marked as yellow thunderbolts), myoD expression is enhanced, leading to the expression of miR-206, which further affects mutual communication between muscle and neural cell and affects reinnervation after denervation.
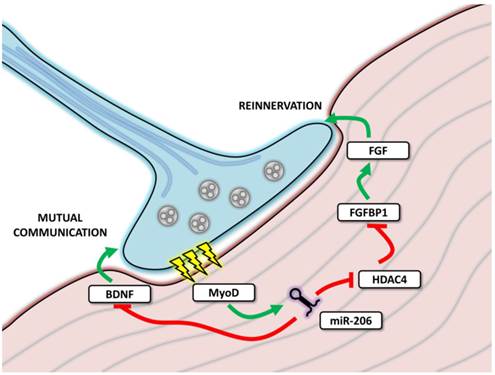
The experiments focused on reinnervation after a sciatic nerve injury showed that miR-206 expression remained unchanged immediately after denervation; however, it increased during the reinnervation process [61]. By targeting MEF2, miR-206 was proposed to affect muscle composition (fiber type) and to constitute one of the sources of the differences observed between healthy and reinnervated muscle [61]. However, these results were not confirmed by a subsequent study where miR-206 levels remained unchanged both after denervation and during reinnervation; the role of other myomiRs was thus emphasized [62].
Reinnervation has also been intensely studied in amyotrophic lateral sclerosis (ALS) [59, 63]. ALS is a neurodegenerative disease which causes a loss of motoneurons, resulting in the denervation of skeletal muscle, consequent paralysis and death [59, 63]. MiR-206 levels were shown to be upregulated in both mice [59] and humans [63]. In the mouse model, the effects of miR-206 were analyzed in detail using miR-206 knockdown animals. Although the ALS onset in these animals was the same as wild type animals, the progression of ALS was faster, resulting in a more severe phenotype, delayed reinnervation and subsequent over-innervation with motoric axons sprouting beyond the original NMJ area [59]. HDAC4, a miR-206 target was suggested to play a role in these effects and the entire signaling pathway, including miR-206/HDAC4/FGFBP1/FGF, was suggested as an explanation of these processes [59]. In human ALS samples, miR-206 was upregulated in both slow and rapid progression patients and only levels of HDAC4 served to distinguish between these groups, indicating that other signals are also necessary for the reinnervation process [63].
3.3 Muscular dystrophies
Muscular dystrophies (MD) represent a group of heritable diseases whose common sign is muscle weakness, often in a progressive form [64]. The most common X-linked and autosomal dominant MDs are Duchene MD and myotonic dystrophy type 1 (MDT1) respectively [64, 65]. The first study to report the dysregulation of myomiRs in Duchene MD was conducted by McCarthy et al. using mdx mice [66]. The study showed that miR-206 is downregulated in the plantaris muscle, but is increased in the diaphragm, most likely due to the upregulation of MyoD and possibly due to the functional denervation of the regenerating muscle [66]. Roberts et al. further confirmed that miR-206 levels also differ between muscle groups in an experimental animal [67]. It was also shown that miR-206 increase occurs during muscle regeneration [68, 69] and, since the diaphragm is the most affected muscle in mdx mice [66], high expression of miR-206 may indicate a high regenerative capacity of this muscle. Interestingly, Yuasa et al. reported that in CXMDJ dogs, which display a more severe phenotype than mdx mice, miR-206 levels are not increased, which may reflect insufficient regeneration [68]. All of the above mentioned information correlates with a subsequently reported observation, i.e. that miR-206 may be found mostly in satellite cells where it stimulates their differentiation in order to initiate muscle regeneration [70].
In MDT1, the miR-206 dysregulation and other muscle-specific miRNAs were described several years later [71, 72]. Whether miR-206 levels are upregulated [72] or unchanged [71] still remains to be elucidated by more extensive studies; however, in MDT1 patients, miR-206 is mislocalized within the cell compared to normal muscle tissue [71, 72].
The above mentioned studies clearly indicate that miR-206 is extensively involved in the pathophysiology of various muscle dystrophies. From a theranostic point of view, miR-206 levels (along with miR-1 and miR-133) were shown to be good serum biomarkers for DMD, correlating even with the severity of the disease [40, 67]. Since their levels are not affected by physical exercise [38-40] they may serve as more precise and valuable biomarkers than the currently used creatinkinase. Current studies also suggest that miR-206 may be used as a therapeutic tool in muscular dystrophy thanks to its reported pro-differentiating and disease-progression slowing effect [67, 73] (Figure 6).
Roles of microRNA-206 in muscular dystrophies. The figure represents miR-206 involvement in muscle dystrophies (MD). The upper left circle emphasizes its role in MD pathophysiology, i.e. involvement in muscle regeneration and differentiation and being differentially expressed in distinct muscle groups. The upper right circle shows its possible use as a circulating biomarker of MD and its correlation with patient prognosis. The lower circle shows the potential to use miR-206 mimics in the treatment of MD.
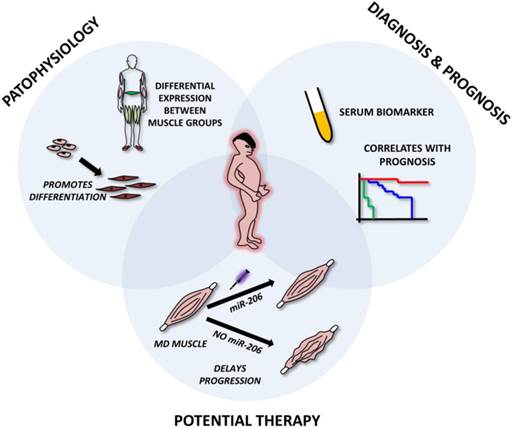
4. MIR-206 IN TUMOR BIOLOGY (Figure 7)
4.1 Rhabdomyosarcoma
Rhabdomyosarcoma represents the most common soft tissue sarcoma with embryonal RMS (ERMS) being the most common type, followed by alveolar RMS (ARMS) [74]. Both of these tumors express markers specific to skeletal muscle; however, their cells are much less differentiated in comparison with muscle tissue [75]. Although the downregulation of miR-206 (and other myomiRs) has been observed repeatedly in both types, it is more prominent in ARMS, thus partially explaining its less differentiated phenotype and more aggressive clinical behavior [76-78].
MicroRNA-206 in tumors. The figure summarizes various cancer types where miR-206 levels and roles were investigated. The colors of the boxes on both sides correspond to the colors of the organs indicated in the human outline. The arrows indicate whether miR-206 levels are upregulated or downregulated.
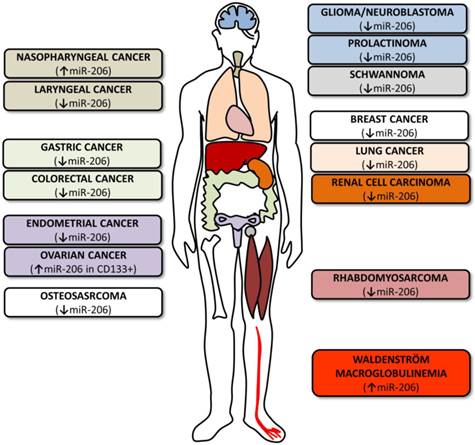
Experiments reinducing miR-206 in RMS cell lines have led to decreased cell proliferation and migration properties and increased myogenic differentiation [76, 77]. Expression profiling in the RD18 ERMS cell line identified more than 700 mRNA levels to be affected by miR-206 transfection [77]. Cell line experiments were further extended by injecting miR-206 directly into RMS mouse models which caused a significant decrease in tumor mass and an increase in the expression of differentiation markers implicating miR-206 as a RMS therapy [76, 77].
Clinical trials focusing on the association between miR-206 and RMS have also been conducted, resulting in circulating miR-206 being identified as a potent diagnostic marker: the levels of this microRNA are muscle-specific even if this microRNA is downregulated in RMS tissue and its circulating levels are higher in RMS patients than in healthy controls [79]. MiR-206 also provides information about the prognosis of a patient: tumors expressing lower miR-206 levels are considered more aggressive with a poorer prognosis [80].
4.2 Breast cancer
Breast cancer is the most common cancer in women [81]. In most cases, ERα is present and provides an enormous theranostic potential: its presence facilitates both disease classification and the tailoring of individual therapy. MicroRNA array analysis identified microRNA deregulations in this cancer [82] with miR-206 being increased in ERα negative and decreased in ERα positive breast cancer [83, 84]. ERα was repeatedly shown to be a target for miR-206 [33, 83, 85, 86] and two miR-206 binding sites were found to be localized in its 3´UTR, along with binding sites for a number of additional miRNAs [33]. Moreover, miR-206 also targets steroid receptor coactivators (SRC)-1 and SRC-3 and TF GATA-3, all of which are part of estrogen-mediated signaling pathways; miR-206-mediated inhibition thus disrupts estrogen signaling on more levels than simply affecting ERα expression [87].
MiR-206 levels and activity in breast cancer are subject to the influence of numerous factors. Estradiol activation of ERα causes miR-206 downregulation [33]; however, this regulatory circuitry may be distracted by epidermal growth factor (EGF) signaling (applying both estradiol and EGF does not decrease miR-206) partly through the EGF-receptor/MAPK pathway [87]. On the contrary, miR-206 activity may be increased when ellagic acid is used [88] or when a single nucleotide polymorphism (rs9341070) in ERα 3´UTR is present - as it increases miR-206 binding [33], or when Ago-2 protein levels are increased - as observed in breast cancer cells [89]. The factors contributing to increased miR-206 activity also contribute to ERα negativity [89] and consequently to a poorer prognosis.
Thus far, applying miR-206 to breast cancer cell lines has been shown to decrease cell growth [83, 85] due to cell cycle arrest [11, 83, 86] and to decrease cell proliferation following estradiol stimulation [86]. Underlying cell growth decrease mechanisms include the cyclinD2 downregulation, an important molecule inducing cell cycle entry [11] and hepatocyte growth factor receptor (MET) downregulation [85]. These results indicate miR-206's future therapeutic potential in breast cancer therapy.
MiR-206 has also been identified as a metastasis-preventing microRNA [90] and its decreased levels have been associated with advanced clinical stages and the presence of lymph node metastasis [91]. Even if its levels do not correlate with prognostic parameters, pre-miR-206 SNP (rs6920648) presence affects patient survival (HR 0.77, p = 0.02), with stronger associations being observed in Caucasians and women with ER+ tumors than in African Americans [92]. However, how exactly this SNP affects miR-206 biogenesis still remains to be clarified.
4.3 Other cancers
As miR-206's potential roles in other cancers are currently being intensely studied, only a limited number of studies are currently available (Table 1). Some of these studies focus on the microRNA profiling of specific cancers [93-95], others evaluate the potential link of miR-206 expression levels to the clinical outcomes of individual patients [96, 97] and still others evaluate the miR-206's potential use in treatment [12, 98].
Overview of miR-206 roles in other cancers
| Cancer | Significance of miR-206 | Ref. |
|---|---|---|
| Endometrial cancer | In ERα+ endometrial endometroid carcinoma, miR-206 levels are decreased and correlate negatively with ERα levels. | [101] |
| Introduction of miR-206 causes cell cycle arrest and affects invasiveness. | ||
| Colorectal carcinoma | Lower expression of miR-206 indicates worse prognosis. | [96] |
| Prognostic signature of miR-206 and additional four miRs (let-7a, miR-21, miR-135a, miR-335) may be used to detect metastases. | ||
| Downregulation of miR-206 dysregulates miR-206/KLF-4 feedback loop which is involved in cancer development. | [102, 103] | |
| Gastric cancer | Low expression of miR-206 indicates presence of metastases, venous invasion and higher probability of hematogenous recurrence along with shortened overall survival. | [12, 97] |
| MiR-206 targeting of cyclinD2 is partially responsible for decrease in cell growth and impaired colony growth of gastric cancer cell lines. | [12] | |
| Lung cancer | Low expression of miR-206 was detected in samples with metastasis. | [13] |
| Transfection of miR-206 caused decrease in cell proliferation, migration and invasion and induced apoptosis. | ||
| Lung carcinogen 4-(methylnitrosamino)-1-(3-pyridyl)-1-butanone increases levels of miR-206 (and miR-133b) in plasma of exposed rats. | [104] | |
| Osteosarcoma | MiR-206 is one of the downregulated microRNAs in osteosarcoma. | [105] |
| Naso-pharyngeal carcinoma | MiR-206 is one of 13 identified microRNAs suggested to play important roles in nasopharyngeal carcinoma development. | [106] |
| Levels of miR-206 increased when lymph node metastasis were present. | ||
| Laryngeal cancer | Levels of miR-206 are downregulated in laryngeal carcinoma patients. | [99] |
| Induction of miR-206 in cancer cell lines causes decrease in viability and invasive potential and increase in apoptosis. | ||
| Application of miR-206 into mouse model leads to significant decrease in tumor mass. | ||
| Ovarian cancer | MiR-206 is higher in CD133+ cells (ovarian cancer stem cells) compared to CD133- cells. | [93] |
| Prostate cancer | MiR-206 expression does not change during the progression of prostate cancer from intraepithelial neoplasia to metastases. | [107] |
| Renal cell carcinoma | MiR-206 is downregulated in clear cell renal cell carcinoma and inhibits cancer cell proliferation. | [94, 108] |
| Prolactinoma | Bromocriptine treatment increases levels of miR-206 in prolatinomas which may partly explain bromocriptine-induced apoptosis. | [98] |
| Vestibular schwannoma | MiR-206 is significantly downregulated in schwannoma cells. | [95] |
| Glioma and neuroblastoma | MiR-206 is downregulated in tumor cell lines and its transfection leads to apoptosis, in part by targeting Otx2. | [109] |
| Waldenström macro-globulinemia | MiR-206 levels are upregulated in Waldenström macroglubulinemia cells accounting for modulation of histon deacetylation in part by targeting HAT (Myst3) protein. | [110] |
The studies reviewed in Table 1 clearly indicate that the downregulation of miR-206 is associated with poorer patient prognosis, most notably due to the increased metastatic potential of the listed cancers [13, 96, 97]. A potential explanation for this phenomenon may be found in the miR-206 targeting of VEGFA, an important angiogenesis factor, as shown in laryngeal cancer [99] or during zebrafish development [100]. In the latter study, miR-1/206 knockdown zebrafish exhibited larger intersegmental vessel diameter and an increased number of endothelial cells [100]. While this finding needs to be further confirmed on different tumors and in larger cohorts, it may be hypothesized that miR-206 downregulation is a general sign of tumorigenicity which enables neoangiogenesis and the proliferation of tumor cells.
5. ROLES FOR MIR-206 IN OTHER TISSUES AND DISEASES
5.1 Myocardial infarction, heart failure and dilated cardiomyopathy
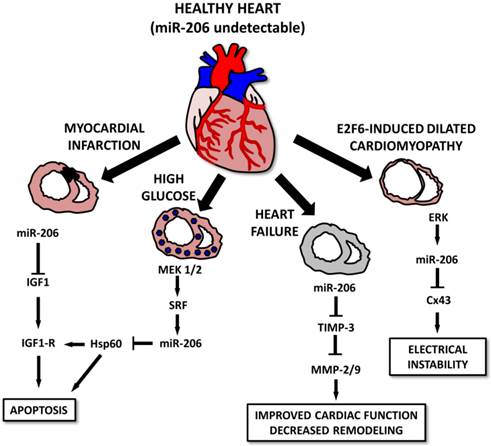
Various studies indicate that in spite of the muscle-specific character of miR-206, it may also play a pivotal role in the heart muscle [10, 111, 112] and associated conditions. The first study connecting miR-206 to the heart was conducted by Shan et al. [112] and showed that in a mouse model of myocardial infarction, levels of miR-1 and miR-206 are increased in infarcted tissue [112]. The authors suggest that both selected miRs may be involved in apoptotic signaling via IGF-1, leading to cardiomyocyte dysfunction [112]. The same group also showed that myocardial injury is further promoted by miR-206 targeting heat-shock protein 60 (Hsp60) in in vitro and in vivo models of diabetes. An increase in miR-1 and miR-206 was probably induced by a MEK1/2 increase which caused a serum response factor (SRF) increase which induced the expression of the above mentioned miRNAs by binding to specific response elements (RE 2,3,4 for miR-1 and RE5 for miR-206) [111]. A miR-206 increase was also observed in a rat model of heart failure and was even more prominent in mice treated with HMGB1 [10]. Administering HMGB1 improved cardiac function along with collagenolytic activity and decreased cardiac remodeling in experimental animals. These effects may be at least in part explained by an miR-206-induced decrease in tissue inhibitor of metalloproteinase 3 (TIMP-3) levels, thus enabling an increase in matrix metalloproteinase (MMP)2 and MMP9 [10].
Last but not least, miR-206 is involved in E2F6-induced dilated cardiomyopathy (DCM) [113]. E2F6 represses the EF2/Rb signaling pathway; transgenic mice in which E2F6 expression was sustained until adulthood generally present with DCM leading to heart failure. E2F6 dysregulation in Tg mice was associated with ERK pathway activation leading to an increase in miR-206 levels and Cx43 downregulation leading to DCM and electrical instability [113] (Figure 8).
Roles for microRNA-206 in cardiac diseases. In a healthy heart, miR-206 levels are barely detectable. In diseased conditions, its levels increase, leading to the apoptosis of the cells after myocardial infarction or in some cases, diabetic cardiomyopathy. In E2F6-induced dilated cardiomyopathy cases, increased miR-206 levels lead to electrical instability. However, heart failure cases, miR-206 overexpression seems to improve cardiac function and positively affect cardiac remodeling.
5.2 Pulmonary hypertension, COPD and infectious respiratory diseases
It has been suggested previously that the development of pulmonary artery hypertension (PAH) resembles tumor growth to a certain degree, especially due to the devastating proliferation of smooth muscle cells (SMC) [114]. As in the case of tumors, mouse models of pulmonary hypertension have likewise shown downregulated miR-206 levels in examined tissue [14, 115] and upregulated in serum [14]. However, other recently conducted studies present contradictory results. Jalali et al. showed that transfecting miR-206 to pulmonary artery SMCs (PASMCs) caused a decrease in proliferation and cellular migration and an increase in apoptosis, probably due to a decrease in Notch3 signaling. Notch3 is a known repressor of SMC differentiation and miR-206-mediated inhibition results in an increase of SMC differentiation markers, such as calponin and SM specific α-actin. Notch3 is also known to be involved in the pathogenesis of PAH and its inhibition by miR-206 may serve as a potential treatment for PAH in the future [115]. Contradictory results, namely a miR-206-mediated increase in proliferation, has been observed in a study by Yue et al.; miR-206 was shown to target HIF-1a, thus affecting the HIF-1a/Fhl-1 pathway; however, miR-206 transfection paradoxically led to an increase in HIF-1a levels [14]. Future studies are therefore needed to shed more light on the role of miR-206 in PAH pathogenesis.
Furthermore, it is known that PAH often develops as a result of COPD [116]. Increased levels of miR-206 and other muscle-specific miRs were established in COPD patients sera samples; these levels negatively correlated with the function of the quadriceps muscle [16] in which miRNA expression analysis previously showed altered miRNA expression [117]. An increase in serum levels of miR-206 is probably the result of increased muscle turnover, fiber type shift and muscle wasting in COPD patients; circulating miR levels may thus serve as biochemical markers of muscle performance [16].
MicroRNAs have also been shown to play a role in infectious respiratory diseases as they participate in agent-host interactions [118]. Using porcine models, miRNA expression profiles were found to be changed and miR-206 levels being upregulated in both porcine reproductive and respiratory syndrome virus (PRRSV) [118] and influenza virus [119] infections. Targeting viral RNAs and mRNAs of various inflammatory molecules (determined using computational prediction) indicates potential roles for miR-206 in antiviral response [118, 119].
In humans, miR-206 serum levels have been proven to be differentially expressed in pertussis patients in comparison with healthy controls [120]. Using a panel of five microRNAs (miR-202, miR-342-5p, miR-206, miR-487b, miR-576-5p), the authors were able to diagnose pertussis with a sensitivity of 97.4 % and a specificity of 94.3 % (cut off 0.707) [120].
5.3 Central nervous system
The brain is the most complex organ in the human body. Various miRNAs were shown to be upregulated or downregulated in distinct parts of the brain, which therefore supports their possible area-specific functions: e.g., miR-206 is detected at higher levels in the cerebellum than in the amygdala, the hippocampus or substantia nigra of adult male rats [121]. In a healthy brain, however, miR-206 can be detected only at very low levels, which may be important for proper neural system development: introducing miR-206 into the oligodendrocyte cell line CG-4 causes a decrease in tubulin polymerization-promoting protein (TPPP)/p25, which results in inhibiting oligodendrocyte differentiation [122].
An increase in miR-206 levels has been observed in various pathological conditions and has been shown to be of theranostic significance [123-125]. During ischemia/reperfusion injury of the brain [123], miR-206 levels correlate with infarct size in a rat embolic stroke model, i.e. its levels increase during the first two days, followed by a continuous decrease [15]. Similar correlations were observed in mRNA levels of genes controlled by miR-206 (BDNF, Notch3, Cx43 and Otx2) [15]. MiR-206 levels are also upregulated in Alzheimer's disease; downregulation reached by the delivery of antagomiRs (either directly intraventricularly or indirectly intranasally) caused a prominent increase in memory function, synaptogenesis and neurogenesis in the hippocampus [124]. This may serve as a promising strategy for future Alzheimer's disease management. Last but not least, with respect to schizophrenia, miR-206 SNPs (rs17578796 and rs1700) were associated with the occurrence of this disease [125]. MiR-206 was shown to target genes previously associated with schizophrenia [125, 126] while also targeting the RNAs of several viruses (e.g. coxsackievirus B2, poliovirus 1) which were shown to affect neural cell function [126].
5.4 Other roles of microRNA-206
MicroRNA-206 has already been found to be involved in posttranscriptional regulation of numerous processes throughout the body and seems to be potentially useful in clinical practice. Other organs, diseases and conditions where miR-206 was shown to play a role are summarized in Table 2 and it may be expected that more roles for miR-206 will be provided in the near future.
Roles of miR-206 in other diseases
| Tissue | MiR-206 significance | Ref |
|---|---|---|
| Scleroderma | MiR-206 along with miR-21 may serve as a potential diagnostic tool for scleroderma. | [127] |
| Pain control | MiR-206 inhibits the expression of substance P from its mRNA in human mesenchymal-stem-cells-derived neuronal cells. | [128] |
| MiR-206 levels are changed in dorsal root ganglion and spinal cord dorsal horn in CFA-induced inflammatory response or during sciatic nerve ligation. | [129] | |
| Biorhythms | MiR-206 is a part of the MyoD-miR-206-CLOCK-MyoD regulatory loop and thus participates in circadian rhythms regulation. | [130] |
| Brown adipose tissue | MiR-206 expression is abundant in brown adipocytes and is missing in white adipocytes. This supports the idea that brown adipocytes and myocytes originate from a common cell lineage. | [131] |
| Liver steatosis | MiR-206 suppresses liver X receptor α (LXRα) levels, thus inhibiting lipogenesis and serving as a potential therapeutic tool for future treatment of lipogenesis-associated diseases such as liver steatosis. | [132] |
6. Conclusions
Under physiological conditions, miR-206 may be skeletal-muscle-specific [5, 7]; however, under pathological conditions, its pleiotropic character is shown to be involved in the pathogenesis of various diseases, including heart failure [10], different types of tumors such as rhabdomyosarcoma [22, 76, 77] and breast cancer [11, 33, 83, 91] or neurological disorders such as Alzheimer's disease [124]. In most of the above mentioned diseases, miR-206 levels are downregulated, which leads to the recognition of this miRNA as “disease-preventing”. In muscle dystrophies, where miR-206 is upregulated, this upregulation seems to be of “self-treating” significance [66, 73]. Alzheimer's disease is an exception from this “disease-preventing rule”, since in this case miR-206 upregulation does more harm than good by decreasing levels of BDNF, a molecule involved in neuronal plasticity and other brain functions [124].
MiR-206's importance is further highlighted by the many factors known to be involved in its regulation. The complex network of regulatory mechanisms results not only in miR-206's tissue-specific expression [7] but also in area- or locality-specific expression as shown in various types of muscle [66, 67] and distinct parts of the brain [121]. Whether these slight differences affect the precise functions of the above mentioned tissues (e.g. by affecting fiber type or neurotransmission), must be determined by future studies.
Using many examples, miR-206 was shown to have great theranostic potential for future clinical practice, serving as a disease biomarker, a disease progression and patient prognosis marker and in some cases even serving as a treatment tool.
To diagnose a disease, more than one miRNA is generally necessary and more miRNAs create a specific pattern which facilitates the ability to distinguish different diseases. While MiR-206 is very often part of these patterns, its elevated circulating levels on their own may also be enough to identify patients with rhabdomyosarcoma [79] or to assess the severity of muscle wasting in patients with COPD [16].
With respect to the therapeutic potential of miR-206, both currently suggested methods using miR mimics and antagomiRs may be used [133]. Using miR-206 mimics should lead to its level restoration in diseased tissue and the restoration of many target protein levels and thus to an improvement of the disease or its cure. This has already been suggested for rhabdomyosarcoma [76, 77], muscle dystrophies [73] and pulmonary hypertension [134]. AntagomiR against miR-206 may - on the other hand - offer a promising strategy to deal with Alzheimer's disease [124].
Using miRNAs as treatment tools is currently becoming a reality since there are biotech companies focusing on the use of miRNAs in diagnostics as well as pharmaceutical companies finalizing preclinical research phases and proceeding to clinical trials. In the case of liver cancer, one of the miRNAs, miR-34a, has already entered Phase I (NCT01829971). In the case of miR-206, MiRagen therapeutics is now in the “lead optimization” phase of research for using this miRNA as treatment for neuromuscular disorders [131].
In conclusion, miRNAs, and especially miR-206, have great theranostic potential; its real significance will most likely be determined in the very near future.
Abbreviations
miRNA/miR: microRNA; mRNA: messenger RNA; DGCR8: DiGeorge syndrome critical region 8; UTR: untranslated region; Ago: Argounat; hsa: homo sapiens; HMGB1: high mobility group box 1 protein; IL: interleukin; TF: transcription factor; MyoD: myocyte differentiation; MEF2C: myocyte enhancer factor 2C; SHP: small hetodimer partner; ERRγ: estrogen-related receptor γ; YY1: Ying Yang 1; AP1: activator protein 1; NFκB: nuclear factor κ B; Shh: Sonic Hedgehog; TGFβ: tumor growth factor β; FGF2: fibroblast growth factor 2; FGF4: fibroblast growth factor 4; ERα: estrogen receptor α; BMP-2: bone morphogenetic protein 2; Pax3: paired box 3; Pax7: paired box 7; POLα: DNA-polymerase α; HDAC4: histon deacetylase 4; Cx43: connexin 43; SCs: satellite cells; NMJ: neuromuscular junction; BDNF: brain-derived neurotropic factor; ALS: amyotrophic lateral sclerosis; FGFBP1: fibroblast growth factor binding protein 1; FGF: fibroblast growth factor; MD: muscular dystrophy; mdx: muscular dystrophy; DMD: Duchene muscular dystrophy; MDT1: myotonic dystrophy type 1; CXMDJ: canine X-linked muscular dystrophy; RMS: rhabdomyosarcoma; ERMS: embryonal rhabdomyosarcoma; ARMS: alveolar rhabdomyosarcoma; SRC1: steroid receptor coactivator 1; SRC3: steroid receptor coactivator 3; EGF: epidermal growth factor; MAPK: mitogen activated protein kinase; MET: hepatocyte growth factor receptor; SNP: single nucleotide polymorphism; VEGF: vascular endothelial growth factor; KLF-4: Krüppel Like-Factor 4; CD: cluster of differentiation; Hsp: heat-shock protein; RE: response elements; TIMP-3: tissue inhibitor of metalloproteinase 3; MMP: matrix metalloproteinase; DCM: dilated cardiomyopathy; PAH: pulmonary artery hypertension; SMC: smooth muscle cells; PASMC: pulmonary artery SMC; HIF: hypoxia-inducible factor; COPD: chronic obstructive pulmonary disease; PRRSV: porcine reproductive and respiratory syndrome virus; TPPP: tubulin polymerization-promoting protein; LXRα: liver X receptor α
Acknowledgements
This work was supported by the European Regional Development Fund - Project FNUSA-ICRC (No. CZ.1.05/1.1.00/02.0123) and by the MUNI/A/0951/2012 grant. We would like to gratefully thank Mr. Matthew Smith for careful proofreading of the manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Lee RC, Feinbaum RL, Ambros V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell. 1993;75:843-54
2. Kozomara A, Griffiths-Jones S. miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic acids research. 2011;39:D152-7 doi:10.1093/nar/gkq1027
3. Huang Y, Shen XJ, Zou Q, Wang SP, Tang SM, Zhang GZ. Biological functions of microRNAs: a review. Journal of physiology and biochemistry. 2011;67:129-39 doi:10.1007/s13105-010-0050-6
4. Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM, Castle J. et al. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature. 2005;433:769-73 doi:10.1038/nature03315
5. McCarthy JJ. MicroRNA-206: the skeletal muscle-specific myomiR. Biochimica et biophysica acta. 2008;1779:682-91 doi:10.1016/j.bbagrm.2008.03.001
6. van Rooij E, Quiat D, Johnson BA, Sutherland LB, Qi X, Richardson JA. et al. A family of microRNAs encoded by myosin genes governs myosin expression and muscle performance. Developmental cell. 2009;17:662-73 doi:10.1016/j.devcel.2009.10.013
7. Lagos-Quintana M, Rauhut R, Yalcin A, Meyer J, Lendeckel W, Tuschl T. Identification of tissue-specific microRNAs from mouse. Current biology: CB. 2002;12:735-9
8. Townley-Tilson WH, Callis TE, Wang D. MicroRNAs 1, 133, and 206: critical factors of skeletal and cardiac muscle development, function, and disease. The international journal of biochemistry & cell biology. 2010;42:1252-5 doi:10.1016/j.biocel.2009.03.002
9. Novak J, Vinklarek J, Bienertova-Vasku J, Slaby O. MicroRNAs involved in skeletal muscle development and their roles in rhabdomyosarcoma pathogenesis. Pediatric blood & cancer. 2013 doi:10.1002/pbc.24664
10. Limana F, Esposito G, D'Arcangelo D, Di Carlo A, Romani S, Melillo G. et al. HMGB1 attenuates cardiac remodelling in the failing heart via enhanced cardiac regeneration and miR-206-mediated inhibition of TIMP-3. PloS one. 2011;6:e19845. doi:10.1371/journal.pone.0019845
11. Zhou J, Tian Y, Li J, Lu B, Sun M, Zou Y. et al. miR-206 is down-regulated in breast cancer and inhibits cell proliferation through the up-regulation of cyclinD2. Biochemical and biophysical research communications. 2013;433:207-12 doi:10.1016/j.bbrc.2013.02.084
12. Zhang L, Liu X, Jin H, Guo X, Xia L, Chen Z. et al. miR-206 inhibits gastric cancer proliferation in part by repressing cyclinD2. Cancer Lett. 2013;332:94-101 doi:10.1016/j.canlet.2013.01.023
13. Wang X, Ling C, Bai Y, Zhao J. MicroRNA-206 is associated with invasion and metastasis of lung cancer. Anatomical record (Hoboken, NJ: 2007). 2011;294:88-92 doi:10.1002/ar.21287
14. Yue J, Guan J, Wang X, Zhang L, Yang Z, Ao Q. et al. MicroRNA-206 is involved in hypoxia-induced pulmonary hypertension through targeting of the HIF-1alpha/Fhl-1 pathway. Lab Invest. 2013;93:748-59 doi:10.1038/labinvest.2013.63
15. Liu FJ, Lim KY, Kaur P, Sepramaniam S, Armugam A, Wong PT. et al. microRNAs Involved in Regulating Spontaneous Recovery in Embolic Stroke Model. PloS one. 2013;8:e66393. doi:10.1371/journal.pone.0066393
16. Donaldson A, Natanek SA, Lewis A, Man WD, Hopkinson NS, Polkey MI. et al. Increased skeletal muscle-specific microRNA in the blood of patients with COPD. Thorax. 2013 doi:10.1136/thoraxjnl-2012-203129
17. Haas JD, Nistala K, Petermann F, Saran N, Chennupati V, Schmitz S. et al. Expression of miRNAs miR-133b and miR-206 in the Il17a/f locus is co-regulated with IL-17 production in alphabeta and gammadelta T cells. PloS one. 2011;6:e20171. doi:10.1371/journal.pone.0020171
18. Bentzinger CF, Wang YX, Rudnicki MA. Building muscle: molecular regulation of myogenesis. Cold Spring Harbor perspectives in biology. 2012 4. doi:10.1101/cshperspect.a008342
19. Sweetman D, Goljanek K, Rathjen T, Oustanina S, Braun T, Dalmay T. et al. Specific requirements of MRFs for the expression of muscle specific microRNAs, miR-1, miR-206 and miR-133. Developmental biology. 2008;321:491-9 doi:10.1016/j.ydbio.2008.06.019
20. Hirai H, Verma M, Watanabe S, Tastad C, Asakura Y, Asakura A. MyoD regulates apoptosis of myoblasts through microRNA-mediated down-regulation of Pax3. The Journal of cell biology. 2010;191:347-65 doi:10.1083/jcb.201006025
21. Rosenberg MI, Georges SA, Asawachaicharn A, Analau E, Tapscott SJ. MyoD inhibits Fstl1 and Utrn expression by inducing transcription of miR-206. The Journal of cell biology. 2006;175:77-85 doi:10.1083/jcb.200603039
22. Macquarrie KL, Yao Z, Young JM, Cao Y, Tapscott SJ. miR-206 integrates multiple components of differentiation pathways to control the transition from growth to differentiation in rhabdomyosarcoma cells. Skelet Muscle. 2012;2:7. doi:10.1186/2044-5040-2-7
23. Gagan J, Dey BK, Layer R, Yan Z, Dutta A. Notch3 and Mef2c proteins are mutually antagonistic via Mkp1 protein and miR-1/206 microRNAs in differentiating myoblasts. The Journal of biological chemistry. 2012;287:40360-70 doi:10.1074/jbc.M112.378414
24. Rachagani S, Cheng Y, Reecy JM. Myostatin genotype regulates muscle-specific miRNA expression in mouse pectoralis muscle. BMC research notes. 2010;3:297. doi:10.1186/1756-0500-3-297
25. Georges M, Clop A, Marcq F, Takeda H, Pirottin D, Hiard S. et al. Polymorphic microRNA-target interactions: a novel source of phenotypic variation. Cold Spring Harbor symposia on quantitative biology. 2006;71:343-50 doi:10.1101/sqb.2006.71.056
26. Song G, Wang L. Nuclear receptor SHP activates miR-206 expression via a cascade dual inhibitory mechanism. PloS one. 2009;4:e6880. doi:10.1371/journal.pone.0006880
27. Lu L, Zhou L, Chen EZ, Sun K, Jiang P, Wang L. et al. A Novel YY1-miR-1 regulatory circuit in skeletal myogenesis revealed by genome-wide prediction of YY1-miRNA network. PloS one. 2012;7:e27596. doi:10.1371/journal.pone.0027596
28. Radzikinas K, Aven L, Jiang Z, Tran T, Paez-Cortez J, Boppidi K. et al. A Shh/miR-206/BDNF cascade coordinates innervation and formation of airway smooth muscle. J Neurosci. 2011;31:15407-15 doi:10.1523/jneurosci.2745-11.2011
29. Sweetman D, Rathjen T, Jefferson M, Wheeler G, Smith TG, Wheeler GN. et al. FGF-4 signaling is involved in mir-206 expression in developing somites of chicken embryos. Developmental dynamics: an official publication of the American Association of Anatomists. 2006;235:2185-91 doi:10.1002/dvdy.20881
30. Winbanks CE, Wang B, Beyer C, Koh P, White L, Kantharidis P. et al. TGF-beta regulates miR-206 and miR-29 to control myogenic differentiation through regulation of HDAC4. The Journal of biological chemistry. 2011;286:13805-14 doi:10.1074/jbc.M110.192625
31. Dong H, Paquette M, Williams A, Zoeller RT, Wade M, Yauk C. Thyroid hormone may regulate mRNA abundance in liver by acting on microRNAs. PloS one. 2010;5:e12136. doi:10.1371/journal.pone.0012136
32. Visser WE, Heemstra KA, Swagemakers SM, Ozgur Z, Corssmit EP, Burggraaf J. et al. Physiological thyroid hormone levels regulate numerous skeletal muscle transcripts. The Journal of clinical endocrinology and metabolism. 2009;94:3487-96 doi:10.1210/jc.2009-0782
33. Adams BD, Furneaux H, White BA. The micro-ribonucleic acid (miRNA) miR-206 targets the human estrogen receptor-alpha (ERalpha) and represses ERalpha messenger RNA and protein expression in breast cancer cell lines. Molecular endocrinology (Baltimore, Md). 2007;21:1132-47 doi:10.1210/me.2007-0022
34. Sato MM, Nashimoto M, Katagiri T, Yawaka Y, Tamura M. Bone morphogenetic protein-2 down-regulates miR-206 expression by blocking its maturation process. Biochemical and biophysical research communications. 2009;383:125-9 doi:10.1016/j.bbrc.2009.03.142
35. Katagiri T, Yamaguchi A, Komaki M, Abe E, Takahashi N, Ikeda T. et al. Bone morphogenetic protein-2 converts the differentiation pathway of C2C12 myoblasts into the osteoblast lineage. The Journal of cell biology. 1994;127:1755-66
36. Katagiri T, Akiyama S, Namiki M, Komaki M, Yamaguchi A, Rosen V. et al. Bone morphogenetic protein-2 inhibits terminal differentiation of myogenic cells by suppressing the transcriptional activity of MyoD and myogenin. Experimental cell research. 1997;230:342-51 doi:10.1006/excr.1996.3432
37. Drummond MJ, Glynn EL, Fry CS, Dhanani S, Volpi E, Rasmussen BB. Essential amino acids increase microRNA-499, -208b, and -23a and downregulate myostatin and myocyte enhancer factor 2C mRNA expression in human skeletal muscle. The Journal of nutrition. 2009;139:2279-84 doi:10.3945/jn.109.112797
38. Mizuno H, Nakamura A, Aoki Y, Ito N, Kishi S, Yamamoto K. et al. Identification of muscle-specific microRNAs in serum of muscular dystrophy animal models: promising novel blood-based markers for muscular dystrophy. PloS one. 2011;6:e18388. doi:10.1371/journal.pone.0018388
39. Aoi W, Ichikawa H, Mune K, Tanimura Y, Mizushima K, Naito Y. et al. Muscle-enriched microRNA miR-486 decreases in circulation in response to exercise in young men. Front Physiol. 2013;4:80. doi:10.3389/fphys.2013.00080
40. Cacchiarelli D, Legnini I, Martone J, Cazzella V, D'Amico A, Bertini E. et al. miRNAs as serum biomarkers for Duchenne muscular dystrophy. EMBO Mol Med. 2011;3:258-65 doi:10.1002/emmm.201100133
41. Nicholson GA, Morgan GJ, Meerkin M, Strauss ER, McLeod JG. The effect of aerobic exercise on serum creatine kinase activities. Muscle & nerve. 1986;9:820-4 doi:10.1002/mus.880090905
42. Nielsen S, Scheele C, Yfanti C, Akerstrom T, Nielsen AR, Pedersen BK. et al. Muscle specific microRNAs are regulated by endurance exercise in human skeletal muscle. The Journal of physiology. 2010;588:4029-37 doi:10.1113/jphysiol.2010.189860
43. Allen DL, Bandstra ER, Harrison BC, Thorng S, Stodieck LS, Kostenuik PJ. et al. Effects of spaceflight on murine skeletal muscle gene expression. Journal of applied physiology (Bethesda, Md: 1985). 2009;106:582-95 doi:10.1152/japplphysiol.90780.2008
44. Koutsoulidou A, Mastroyiannopoulos NP, Furling D, Uney JB, Phylactou LA. Expression of miR-1, miR-133a, miR-133b and miR-206 increases during development of human skeletal muscle. BMC Dev Biol. 2011;11:34. doi:10.1186/1471-213x-11-34
45. Kim HK, Lee YS, Sivaprasad U, Malhotra A, Dutta A. Muscle-specific microRNA miR-206 promotes muscle differentiation. The Journal of cell biology. 2006;174:677-87 doi:10.1083/jcb.200603008
46. Shkumatava A, Stark A, Sive H, Bartel DP. Coherent but overlapping expression of microRNAs and their targets during vertebrate development. Genes & development. 2009;23:466-81 doi:10.1101/gad.1745709
47. Hou X, Tang Z, Liu H, Wang N, Ju H, Li K. Discovery of MicroRNAs associated with myogenesis by deep sequencing of serial developmental skeletal muscles in pigs. PloS one. 2012;7:e52123. doi:10.1371/journal.pone.0052123
48. Xie SS, Huang TH, Shen Y, Li XY, Zhang XX, Zhu MJ. et al. Identification and characterization of microRNAs from porcine skeletal muscle. Animal genetics. 2010;41:179-90 doi:10.1111/j.1365-2052.2009.01991.x
49. Yan X, Ding L, Li Y, Zhang X, Liang Y, Sun X. et al. Identification and profiling of microRNAs from skeletal muscle of the common carp. PloS one. 2012;7:e30925. doi:10.1371/journal.pone.0030925
50. Takada S, Berezikov E, Yamashita Y, Lagos-Quintana M, Kloosterman WP, Enomoto M. et al. Mouse microRNA profiles determined with a new and sensitive cloning method. Nucleic acids research. 2006;34:e115. doi:10.1093/nar/gkl653
51. Bernstein E, Kim SY, Carmell MA, Murchison EP, Alcorn H, Li MZ. et al. Dicer is essential for mouse development. Nature genetics. 2003;35:215-7 doi:10.1038/ng1253
52. O'Rourke JR, Georges SA, Seay HR, Tapscott SJ, McManus MT, Goldhamer DJ. et al. Essential role for Dicer during skeletal muscle development. Developmental biology. 2007;311:359-68 doi:10.1016/j.ydbio.2007.08.032
53. Nakajima N, Takahashi T, Kitamura R, Isodono K, Asada S, Ueyama T. et al. MicroRNA-1 facilitates skeletal myogenic differentiation without affecting osteoblastic and adipogenic differentiation. Biochemical and biophysical research communications. 2006;350:1006-12 doi:10.1016/j.bbrc.2006.09.153
54. Goljanek-Whysall K, Sweetman D, Abu-Elmagd M, Chapnik E, Dalmay T, Hornstein E. et al. MicroRNA regulation of the paired-box transcription factor Pax3 confers robustness to developmental timing of myogenesis. Proceedings of the National Academy of Sciences of the United States of America. 2011;108:11936-41 doi:10.1073/pnas.1105362108
55. Anderson C, Catoe H, Werner R. MIR-206 regulates connexin43 expression during skeletal muscle development. Nucleic acids research. 2006;34:5863-71 doi:10.1093/nar/gkl743
56. Cerletti M, Shadrach JL, Jurga S, Sherwood R, Wagers AJ. Regulation and function of skeletal muscle stem cells. Cold Spring Harbor symposia on quantitative biology. 2008;73:317-22 doi:10.1101/sqb.2008.73.054
57. Chen JF, Tao Y, Li J, Deng Z, Yan Z, Xiao X. et al. microRNA-1 and microRNA-206 regulate skeletal muscle satellite cell proliferation and differentiation by repressing Pax7. The Journal of cell biology. 2010;190:867-79 doi:10.1083/jcb.200911036
58. Nakasa T, Ishikawa M, Shi M, Shibuya H, Adachi N, Ochi M. Acceleration of muscle regeneration by local injection of muscle-specific microRNAs in rat skeletal muscle injury model. Journal of cellular and molecular medicine. 2010;14:2495-505 doi:10.1111/j.1582-4934.2009.00898.x
59. Williams AH, Valdez G, Moresi V, Qi X, McAnally J, Elliott JL. et al. MicroRNA-206 delays ALS progression and promotes regeneration of neuromuscular synapses in mice. Science (New York, NY). 2009;326:1549-54 doi:10.1126/science.1181046
60. Miura P, Amirouche A, Clow C, Belanger G, Jasmin BJ. Brain-derived neurotrophic factor expression is repressed during myogenic differentiation by miR-206. J Neurochem. 2012;120:230-8 doi:10.1111/j.1471-4159.2011.07583.x
61. Jeng SF, Rau CS, Liliang PC, Wu CJ, Lu TH, Chen YC. et al. Profiling muscle-specific microRNA expression after peripheral denervation and reinnervation in a rat model. Journal of neurotrauma. 2009;26:2345-53 doi:10.1089/neu.2009.0960
62. Rau CS, Jeng JC, Jeng SF, Lu TH, Chen YC, Liliang PC. et al. Entrapment neuropathy results in different microRNA expression patterns from denervation injury in rats. BMC musculoskeletal disorders. 2010;11:181. doi:10.1186/1471-2474-11-181
63. Bruneteau G, Simonet T, Bauche S, Mandjee N, Malfatti E, Girard E. et al. Muscle histone deacetylase 4 upregulation in amyotrophic lateral sclerosis: potential role in reinnervation ability and disease progression. Brain: a journal of neurology. 2013;136:2359-68 doi:10.1093/brain/awt164
64. Hoffman EP, Fischbeck KH, Brown RH, Johnson M, Medori R, Loike JD. et al. Characterization of dystrophin in muscle-biopsy specimens from patients with Duchenne's or Becker's muscular dystrophy. The New England journal of medicine. 1988;318:1363-8 doi:10.1056/nejm198805263182104
65. Turner C, Hilton-Jones D. The myotonic dystrophies: diagnosis and management. Journal of neurology, neurosurgery, and psychiatry. 2010;81:358-67 doi:10.1136/jnnp.2008.158261
66. McCarthy JJ, Esser KA, Andrade FH. MicroRNA-206 is overexpressed in the diaphragm but not the hindlimb muscle of mdx mouse. American journal of physiology Cell physiology. 2007;293:C451-7 doi:10.1152/ajpcell.00077.2007
67. Roberts TC, Blomberg KE, McClorey G, El Andaloussi S, Godfrey C, Betts C. et al. Expression analysis in multiple muscle groups and serum reveals complexity in the microRNA transcriptome of the mdx mouse with implications for therapy. Mol Ther Nucleic Acids. 2012;1:e39. doi:10.1038/mtna.2012.26
68. Yuasa K, Hagiwara Y, Ando M, Nakamura A, Takeda S, Hijikata T. MicroRNA-206 is highly expressed in newly formed muscle fibers: implications regarding potential for muscle regeneration and maturation in muscular dystrophy. Cell structure and function. 2008;33:163-9
69. Greco S, De Simone M, Colussi C, Zaccagnini G, Fasanaro P, Pescatori M. et al. Common micro-RNA signature in skeletal muscle damage and regeneration induced by Duchenne muscular dystrophy and acute ischemia. FASEB journal: official publication of the Federation of American Societies for Experimental Biology. 2009;23:3335-46 doi:10.1096/fj.08-128579
70. Cacchiarelli D, Martone J, Girardi E, Cesana M, Incitti T, Morlando M. et al. MicroRNAs involved in molecular circuitries relevant for the Duchenne muscular dystrophy pathogenesis are controlled by the dystrophin/nNOS pathway. Cell metabolism. 2010;12:341-51 doi:10.1016/j.cmet.2010.07.008
71. Perbellini R, Greco S, Sarra-Ferraris G, Cardani R, Capogrossi MC, Meola G. et al. Dysregulation and cellular mislocalization of specific miRNAs in myotonic dystrophy type 1. Neuromuscular disorders: NMD. 2011;21:81-8 doi:10.1016/j.nmd.2010.11.012
72. Gambardella S, Rinaldi F, Lepore SM, Viola A, Loro E, Angelini C. et al. Overexpression of microRNA-206 in the skeletal muscle from myotonic dystrophy type 1 patients. Journal of translational medicine. 2010;8:48. doi:10.1186/1479-5876-8-48
73. Liu N, Williams AH, Maxeiner JM, Bezprozvannaya S, Shelton JM, Richardson JA. et al. microRNA-206 promotes skeletal muscle regeneration and delays progression of Duchenne muscular dystrophy in mice. The Journal of clinical investigation. 2012;122:2054-65 doi:10.1172/jci62656
74. Arndt CA, Rose PS, Folpe AL, Laack NN. Common musculoskeletal tumors of childhood and adolescence. Mayo Clinic proceedings Mayo Clinic. 2012;87:475-87 doi:10.1016/j.mayocp.2012.01.015
75. Davicioni E, Anderson MJ, Finckenstein FG, Lynch JC, Qualman SJ, Shimada H. et al. Molecular classification of rhabdomyosarcoma--genotypic and phenotypic determinants of diagnosis: a report from the Children's Oncology Group. The American journal of pathology. 2009;174:550-64 doi:10.2353/ajpath.2009.080631
76. Yan D, Dong Xda E, Chen X, Wang L, Lu C, Wang J. et al. MicroRNA-1/206 targets c-Met and inhibits rhabdomyosarcoma development. The Journal of biological chemistry. 2009;284:29596-604 doi:10.1074/jbc.M109.020511
77. Taulli R, Bersani F, Foglizzo V, Linari A, Vigna E, Ladanyi M. et al. The muscle-specific microRNA miR-206 blocks human rhabdomyosarcoma growth in xenotransplanted mice by promoting myogenic differentiation. The Journal of clinical investigation. 2009;119:2366-78 doi:10.1172/jci38075
78. Taulli R, Foglizzo V, Morena D, Coda DM, Ala U, Bersani F. et al. Failure to downregulate the BAF53a subunit of the SWI/SNF chromatin remodeling complex contributes to the differentiation block in rhabdomyosarcoma. Oncogene. 2013 doi:10.1038/onc.2013.188
79. Miyachi M, Tsuchiya K, Yoshida H, Yagyu S, Kikuchi K, Misawa A. et al. Circulating muscle-specific microRNA, miR-206, as a potential diagnostic marker for rhabdomyosarcoma. Biochemical and biophysical research communications. 2010;400:89-93 doi:10.1016/j.bbrc.2010.08.015
80. Missiaglia E, Shepherd CJ, Patel S, Thway K, Pierron G, Pritchard-Jones K. et al. MicroRNA-206 expression levels correlate with clinical behaviour of rhabdomyosarcomas. British journal of cancer. 2010;102:1769-77 doi:10.1038/sj.bjc.6605684
81. Jemal A, Siegel R, Ward E, Hao Y, Xu J, Murray T. et al. Cancer statistics, 2008. CA: a cancer journal for clinicians. 2008;58:71-96 doi:10.3322/ca.2007.0010
82. Iorio MV, Ferracin M, Liu CG, Veronese A, Spizzo R, Sabbioni S. et al. MicroRNA gene expression deregulation in human breast cancer. Cancer research. 2005;65:7065-70 doi:10.1158/0008-5472.can-05-1783
83. Kondo N, Toyama T, Sugiura H, Fujii Y, Yamashita H. miR-206 Expression is down-regulated in estrogen receptor alpha-positive human breast cancer. Cancer research. 2008;68:5004-8 doi:10.1158/0008-5472.can-08-0180
84. Yoshimoto N, Toyama T, Takahashi S, Sugiura H, Endo Y, Iwasa M. et al. Distinct expressions of microRNAs that directly target estrogen receptor alpha in human breast cancer. Breast Cancer Res Treat. 2011;130:331-9 doi:10.1007/s10549-011-1672-2
85. Di Leva G, Gasparini P, Piovan C, Ngankeu A, Garofalo M, Taccioli C. et al. MicroRNA cluster 221-222 and estrogen receptor alpha interactions in breast cancer. Journal of the National Cancer Institute. 2010;102:706-21 doi:10.1093/jnci/djq102
86. Leivonen SK, Makela R, Ostling P, Kohonen P, Haapa-Paananen S, Kleivi K. et al. Protein lysate microarray analysis to identify microRNAs regulating estrogen receptor signaling in breast cancer cell lines. Oncogene. 2009;28:3926-36 doi:10.1038/onc.2009.241
87. Adams BD, Cowee DM, White BA. The role of miR-206 in the epidermal growth factor (EGF) induced repression of estrogen receptor-alpha (ERalpha) signaling and a luminal phenotype in MCF-7 breast cancer cells. Molecular endocrinology (Baltimore, Md). 2009;23:1215-30 doi:10.1210/me.2009-0062
88. Munagala R, Aqil F, Vadhanam MV, Gupta RC. MicroRNA 'signature' during estrogen-mediated mammary carcinogenesis and its reversal by ellagic acid intervention. Cancer Lett. 2013 doi:10.1016/j.canlet.2013.06.012
89. Adams BD, Claffey KP, White BA. Argonaute-2 expression is regulated by epidermal growth factor receptor and mitogen-activated protein kinase signaling and correlates with a transformed phenotype in breast cancer cells. Endocrinology. 2009;150:14-23 doi:10.1210/en.2008-0984
90. Tavazoie SF, Alarcon C, Oskarsson T, Padua D, Wang Q, Bos PD. et al. Endogenous human microRNAs that suppress breast cancer metastasis. Nature. 2008;451:147-52 doi:10.1038/nature06487
91. Li Y, Hong F, Yu Z. Decreased expression of microRNA-206 in breast cancer and its association with disease characteristics and patient survival. J Int Med Res. 2013;41:596-602 doi:10.1177/0300060513485856
92. Bensen JT, Tse CK, Nyante SJ, Barnholtz-Sloan JS, Cole SR, Millikan RC. Association of germline microRNA SNPs in pre-miRNA flanking region and breast cancer risk and survival: the Carolina Breast Cancer Study. Cancer Causes Control. 2013;24:1099-109 doi:10.1007/s10552-013-0187-z
93. Guo R, Wu Q, Liu F, Wang Y. Description of the CD133+ subpopulation of the human ovarian cancer cell line OVCAR3. Oncology reports. 2011;25:141-6
94. Zhou L, Chen J, Li Z, Li X, Hu X, Huang Y. et al. Integrated profiling of microRNAs and mRNAs: microRNAs located on Xq27.3 associate with clear cell renal cell carcinoma. PloS one. 2010;5:e15224. doi:10.1371/journal.pone.0015224
95. Torres-Martin M, Lassaletta L, de Campos JM, Isla A, Gavilan J, Pinto GR. et al. Global Profiling in Vestibular Schwannomas Shows Critical Deregulation of MicroRNAs and Upregulation in Those Included in Chromosomal Region 14q32. PloS one. 2013;8:e65868. doi:10.1371/journal.pone.0065868
96. Vickers MM, Bar J, Gorn-Hondermann I, Yarom N, Daneshmand M, Hanson JE. et al. Stage-dependent differential expression of microRNAs in colorectal cancer: potential role as markers of metastatic disease. Clin Exp Metastasis. 2012;29:123-32 doi:10.1007/s10585-011-9435-3
97. Yang Q, Zhang C, Huang B, Li H, Zhang R, Huang Y. et al. Downregulation of microRNA-206 is a potent prognostic marker for patients with gastric cancer. Eur J Gastroenterol Hepatol. 2013;25:953-7 doi:10.1097/MEG.0b013e32835ed691
98. Wang C, Su Z, Sanai N, Xue X, Lu L, Chen Y. et al. microRNA expression profile and differentially-expressed genes in prolactinomas following bromocriptine treatment. Oncology reports. 2012;27:1312-20 doi:10.3892/or.2012.1690
99. Zhang T, Liu M, Wang C, Lin C, Sun Y, Jin D. Down-regulation of MiR-206 promotes proliferation and invasion of laryngeal cancer by regulating VEGF expression. Anticancer Res. 2011;31:3859-63
100. Stahlhut C, Suarez Y, Lu J, Mishima Y, Giraldez AJ. miR-1 and miR-206 regulate angiogenesis by modulating VegfA expression in zebrafish. Development. 2012;139:4356-64 doi:10.1242/dev.083774
101. Chen X, Yan Q, Li S, Zhou L, Yang H, Yang Y. et al. Expression of the tumor suppressor miR-206 is associated with cellular proliferative inhibition and impairs invasion in ERalpha-positive endometrioid adenocarcinoma. Cancer Lett. 2012;314:41-53 doi:10.1016/j.canlet.2011.09.014
102. Lin CC, Liu LZ, Addison JB, Wonderlin WF, Ivanov AV, Ruppert JM. A KLF4-miRNA-206 autoregulatory feedback loop can promote or inhibit protein translation depending upon cell context. Molecular and cellular biology. 2011;31:2513-27 doi:10.1128/mcb.01189-10
103. Parasramka MA, Dashwood WM, Wang R, Saeed HH, Williams DE, Ho E. et al. A role for low-abundance miRNAs in colon cancer: the miR-206/Kruppel-like factor 4 (KLF4) axis. Clin Epigenetics. 2012;4:16. doi:10.1186/1868-7083-4-16
104. Wu J, Yang T, Li X, Yang Q, Liu R, Huang J. et al. Alteration of serum miR-206 and miR-133b is associated with lung carcinogenesis induced by 4-(methylnitrosamino)-1-(3-pyridyl)-1-butanone. Toxicol Appl Pharmacol. 2013;267:238-46 doi:10.1016/j.taap.2013.01.002
105. Namlos HM, Meza-Zepeda LA, Baroy T, Ostensen IH, Kresse SH, Kuijjer ML. et al. Modulation of the osteosarcoma expression phenotype by microRNAs. PloS one. 2012;7:e48086. doi:10.1371/journal.pone.0048086
106. Luo Z, Zhang L, Li Z, Li X, Li G, Yu H. et al. An in silico analysis of dynamic changes in microRNA expression profiles in stepwise development of nasopharyngeal carcinoma. BMC medical genomics. 2012;5:3. doi:10.1186/1755-8794-5-3
107. Leite KR, Tomiyama A, Reis ST, Sousa-Canavez JM, Sanudo A, Camara-Lopes LH. et al. MicroRNA expression profiles in the progression of prostate cancer-from high-grade prostate intraepithelial neoplasia to metastasis. Urol Oncol. 2011 doi:10.1016/j.urolonc.2011.07.002
108. Hidaka H, Seki N, Yoshino H, Yamasaki T, Yamada Y, Nohata N. et al. Tumor suppressive microRNA-1285 regulates novel molecular targets: aberrant expression and functional significance in renal cell carcinoma. Oncotarget. 2012;3:44-57
109. Wang R, Hu Y, Song G, Hao CJ, Cui Y, Xia HF. et al. MiR-206 regulates neural cells proliferation and apoptosis via Otx2. Cell Physiol Biochem. 2012;29:381-90 doi:10.1159/000338493
110. Roccaro AM, Sacco A, Jia X, Azab AK, Maiso P, Ngo HT. et al. microRNA-dependent modulation of histone acetylation in Waldenstrom macroglobulinemia. Blood. 2010;116:1506-14 doi:10.1182/blood-2010-01-265686
111. Shan ZX, Lin QX, Deng CY, Zhu JN, Mai LP, Liu JL. et al. miR-1/miR-206 regulate Hsp60 expression contributing to glucose-mediated apoptosis in cardiomyocytes. FEBS letters. 2010;584:3592-600 doi:10.1016/j.febslet.2010.07.027
112. Shan ZX, Lin QX, Fu YH, Deng CY, Zhou ZL, Zhu JN. et al. Upregulated expression of miR-1/miR-206 in a rat model of myocardial infarction. Biochemical and biophysical research communications. 2009;381:597-601 doi:10.1016/j.bbrc.2009.02.097
113. Westendorp B, Major JL, Nader M, Salih M, Leenen FH, Tuana BS. The E2F6 repressor activates gene expression in myocardium resulting in dilated cardiomyopathy. FASEB journal: official publication of the Federation of American Societies for Experimental Biology. 2012;26:2569-79 doi:10.1096/fj.11-203174
114. Song G, Zhang Y, Wang L. MicroRNA-206 targets notch3, activates apoptosis, and inhibits tumor cell migration and focus formation. The Journal of biological chemistry. 2009;284:31921-7 doi:10.1074/jbc.M109.046862
115. Jalali S, Ramanathan GK, Parthasarathy PT, Aljubran S, Galam L, Yunus A. et al. Mir-206 regulates pulmonary artery smooth muscle cell proliferation and differentiation. PloS one. 2012;7:e46808. doi:10.1371/journal.pone.0046808
116. Farber HW, Loscalzo J. Pulmonary arterial hypertension. The New England journal of medicine. 2004;351:1655-65 doi:10.1056/NEJMra035488
117. Lewis A, Riddoch-Contreras J, Natanek SA, Donaldson A, Man WD, Moxham J. et al. Downregulation of the serum response factor/miR-1 axis in the quadriceps of patients with COPD. Thorax. 2012;67:26-34 doi:10.1136/thoraxjnl-2011-200309
118. Guo XK, Zhang Q, Gao L, Li N, Chen XX, Feng WH. Increasing expression of microRNA 181 inhibits porcine reproductive and respiratory syndrome virus replication and has implications for controlling virus infection. J Virol. 2013;87:1159-71 doi:10.1128/jvi.02386-12
119. Skovgaard K, Cirera S, Vasby D, Podolska A, Breum SO, Durrwald R. et al. Expression of innate immune genes, proteins and microRNAs in lung tissue of pigs infected experimentally with influenza virus (H1N2). Innate Immun. 2013 doi:10.1177/1753425912473668
120. Ge Y, Zhao K, Qi Y, Min X, Shi Z, Qi X. et al. Serum microRNA expression profile as a biomarker for the diagnosis of pertussis. Mol Biol Rep. 2013;40:1325-32 doi:10.1007/s11033-012-2176-9
121. Olsen L, Klausen M, Helboe L, Nielsen FC, Werge T. MicroRNAs show mutually exclusive expression patterns in the brain of adult male rats. PloS one. 2009;4:e7225. doi:10.1371/journal.pone.0007225
122. Lehotzky A, Lau P, Tokesi N, Muja N, Hudson LD, Ovadi J. Tubulin polymerization-promoting protein (TPPP/p25) is critical for oligodendrocyte differentiation. Glia. 2010;58:157-68 doi:10.1002/glia.20909
123. Jeyaseelan K, Lim KY, Armugam A. MicroRNA expression in the blood and brain of rats subjected to transient focal ischemia by middle cerebral artery occlusion. Stroke, a journal of cerebral circulation. 2008;39:959-66 doi:10.1161/strokeaha.107.500736
124. Lee ST, Chu K, Jung KH, Kim JH, Huh JY, Yoon H. et al. miR-206 regulates brain-derived neurotrophic factor in Alzheimer disease model. Ann Neurol. 2012;72:269-77 doi:10.1002/ana.23588
125. Hansen T, Olsen L, Lindow M, Jakobsen KD, Ullum H, Jonsson E. et al. Brain expressed microRNAs implicated in schizophrenia etiology. PloS one. 2007;2:e873. doi:10.1371/journal.pone.0000873
126. Carter CJ. Schizophrenia susceptibility genes directly implicated in the life cycles of pathogens: cytomegalovirus, influenza, herpes simplex, rubella, and Toxoplasma gondii. Schizophrenia bulletin. 2009;35:1163-82 doi:10.1093/schbul/sbn054
127. Koba S, Jinnin M, Inoue K, Nakayama W, Honda N, Makino K. et al. Expression analysis of multiple microRNAs in each patient with scleroderma. Exp Dermatol. 2013;22:489-91 doi:10.1111/exd.12173
128. Greco SJ, Rameshwar P. MicroRNAs regulate synthesis of the neurotransmitter substance P in human mesenchymal stem cell-derived neuronal cells. Proceedings of the National Academy of Sciences of the United States of America. 2007;104:15484-9 doi:10.1073/pnas.0703037104
129. Kusuda R, Cadetti F, Ravanelli MI, Sousa TA, Zanon S, De Lucca FL. et al. Differential expression of microRNAs in mouse pain models. Molecular pain. 2011;7:17. doi:10.1186/1744-8069-7-17
130. Zhou W, Li Y, Wang X, Wu L, Wang Y. MiR-206-mediated dynamic mechanism of the mammalian circadian clock. BMC Syst Biol. 2011;5:141. doi:10.1186/1752-0509-5-141
131. Walden TB, Timmons JA, Keller P, Nedergaard J, Cannon B. Distinct expression of muscle-specific microRNAs (myomirs) in brown adipocytes. Journal of cellular physiology. 2009;218:444-9 doi:10.1002/jcp.21621
132. Zhong D, Huang G, Zhang Y, Zeng Y, Xu Z, Zhao Y. et al. MicroRNA-1 and microRNA-206 suppress LXRalpha-induced lipogenesis in hepatocytes. Cell Signal. 2013;25:1429-37 doi:10.1016/j.cellsig.2013.03.003
133. Soriano A, Jubierre L, Almazan-Moga A, Molist C, Roma J, de Toledo JS. et al. microRNAs as pharmacological targets in cancer. Pharmacological research: the official journal of the Italian Pharmacological Society. 2013;75:3-14 doi:10.1016/j.phrs.2013.03.006
134. Meloche J, Paulin R, Provencher S, Bonnet S. Therapeutic Potential of microRNA Modulation in Pulmonary Arterial Hypertension. Curr Vasc Pharmacol. 2013
Author contact
Corresponding author: Jan Novák, Department of Physiology, Faculty of Medicine, Masaryk University, Kamenice 5, 625 00, Brno, Czech Republic. Email: 324158muni.cz; Phone: +420 725 431 052
 Global reach, higher impact
Global reach, higher impact