Impact Factor
Theranostics 2019; 9(4):1115-1124. doi:10.7150/thno.29622 This issue Cite
Research Paper
Genome-wide profiling of Epstein-Barr virus integration by targeted sequencing in Epstein-Barr virus associated malignancies
1. State Key Laboratory of Oncology in South China, Collaborative Innovation Center for Cancer Medicine, Sun Yat-sen University Cancer Center, Guangzhou, China.
2. State Key Lab of Molecular Oncology, National Cancer Center/Cancer Hospital, Chinese Academy of Medical Sciences and Peking Union Medical College; Collaborative Innovation Center for Cancer Medicine, Beijing, China.
3. Department of Otolaryngology/Head and Neck Surgery, First Affiliated Hospital of Guangxi Medical University, Nanning, China.
4. Department of Medical Microbiology, Qingdao University Medical College, Qingdao, China.
5. State Key Laboratory of Medical Genomics, Shanghai Institute of Hematology, Rui Jin Hospital, Shanghai Jiao Tong University (SJTU) School of Medicine and Collaborative Innovation Center of Systems Biomedicine, Shanghai, China.
* These authors contributed equally to the work.
Abstract
Rationale: Epstein-Barr virus (EBV) is associated with multiple malignancies with expression of viral oncogenic proteins and chronic inflammation as major mechanisms contributing to tumor development. A less well-studied mechanism is the integration of EBV into the human genome possibly at sites which may disrupt gene expression or genome stability.
Methods: We sequenced tumor DNA to profile the EBV sequences by hybridization-based enrichment. Bioinformatic analysis was used to detect the breakpoints of EBV integrations in the genome of cancer cells.
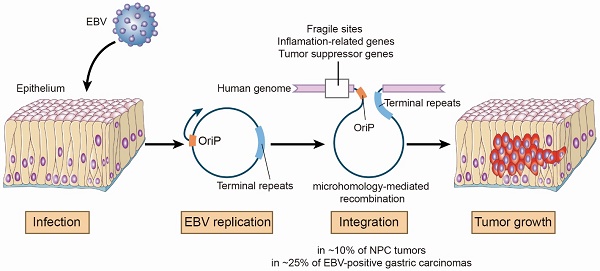
Results: We identified 197 breakpoints in nasopharyngeal carcinomas and other EBV-associated malignancies. EBV integrations were enriched at vulnerable regions of the human genome and were close to tumor suppressor and inflammation-related genes. We found that EBV integrations into the introns could decrease the expression of the inflammation-related genes, TNFAIP3, PARK2, and CDK15, in NPC tumors. In the EBV genome, the breakpoints were frequently at oriP or terminal repeats. These breakpoints were surrounded by microhomology sequences, consistent with a mechanism for integration involving viral genome replication and microhomology-mediated recombination.
Conclusion: Our finding provides insight into the potential of EBV integration as an additional mechanism mediating tumorigenesis in EBV associated malignancies.
Keywords: Epstein-Barr virus (EBV), nasopharyngeal carcinoma (NPC), DNA integration
 Global reach, higher impact
Global reach, higher impact