13.3
Impact Factor
Theranostics 2018; 8(1):61-77. doi:10.7150/thno.20893 This issue Cite
Research Paper
Epigenomic characterization of a p53-regulated 3p22.2 tumor suppressor that inhibits STAT3 phosphorylation via protein docking and is frequently methylated in esophageal and other carcinomas
1. Cancer Epigenetics Laboratory, Department of Clinical Oncology, State Key Laboratory of Oncology in South China, Sir YK Pao Center for Cancer and Li Ka Shing Institute of Health Sciences, The Chinese University of Hong Kong, Hong Kong;
2. Shenzhen Second People's Hospital, the First Affiliated Hospital of Shenzhen University, Shenzhen, China;
3. Johns Hopkins Singapore, Singapore;
4. Department of Pathology, National Cancer Center/Cancer Hospital, Chinese Academy of Medical Sciences & Peking Union Medical College, Beijing, China;
5. Chongqing Key Laboratory of Molecular Oncology and Epigenetics, The First Affiliated Hospital of Chongqing Medical University, Chongqing, China;
6. Department of Pathology, Queen Mary Hospital, The University of Hong Kong;
7. State Key Laboratory of Oncology in South China, Sun Yat-sen University Cancer Center, Guangzhou, China;
8. Institute of Digestive Disease and State Key Laboratory of Digestive Diseases, Department of Medicine and Therapeutics, The Chinese University of Hong Kong;
9. Sidney Kimmel Comprehensive Cancer Center, Johns Hopkins School of Medicine, Baltimore
*Equal Contribution
Abstract
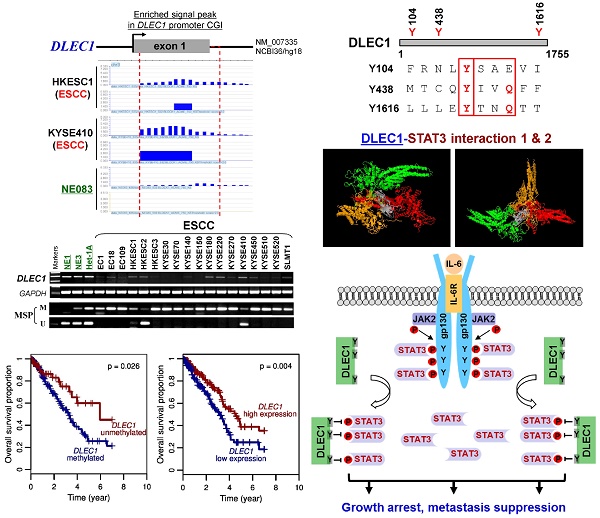
Rationale: Oncogenic STAT3 signaling activation and 3p22-21.3 locus alteration are common in multiple tumors, especially carcinomas of the nasopharynx, esophagus and lung. Whether these two events are linked remains unclear. Our CpG methylome analysis identified a 3p22.2 gene, DLEC1, as a methylated target in esophageal squamous cell (ESCC), nasopharyngeal (NPC) and lung carcinomas. Thus, we further characterized its epigenetic abnormalities and functions.
Methods: CpG methylomes were established by methylated DNA immunoprecipitation. Promoter methylation was analyzed by methylation-specific PCR and bisulfite genomic sequencing. DLEC1 expression and clinical significance were analyzed using TCGA database. DLEC1 functions were analyzed by transfections followed by various cell biology assays. Protein-protein interaction was assessed by docking, Western blot and immunoprecipitation analyses.
Results: We defined the DLEC1 promoter within a CpG island and p53-regulated. DLEC1 was frequently downregulated in ESCC, lung and NPC cell lines and primary tumors, but was readily expressed in normal tissues and immortalized normal epithelial cells, with mutations rarely detected. DLEC1 methylation was frequently detected in ESCC tumors and correlated with lymph node metastasis, tumor recurrence and progression, with DLEC1 as the most frequently methylated among the established 3p22.2 tumor suppressors (RASSF1A, PLCD1 and ZMYND10/BLU). DLEC1 inhibits carcinoma cell growth through inducing cell cycle arrest and apoptosis, and also suppresses cell metastasis by reversing epithelial-mesenchymal transition (EMT) and cell stemness. Moreover, DLEC1 represses oncogenic signaling including JAK/STAT3, MAPK/ERK, Wnt/β-catenin and AKT pathways in multiple carcinoma types. Particularly, DLEC1 inhibits IL-6-induced STAT3 phosphorylation in a dose-dependent manner. DLEC1 contains three YXXQ motifs and forms a protein complex with STAT3 via protein docking, which blocks STAT3-JAK2 interaction and STAT3 phosphorylation. IL-6 stimulation enhances the binding of DLEC1 with STAT3, which diminishes their interaction with JAK2 and further leads to decreased STAT3 phosphorylation. The YXXQ motifs of DLEC1 are crucial for its inhibition of STAT3 phosphorylation, and disruption of these motifs restores STAT3 phosphorylation through abolishing DLEC1 binding to STAT3.
Conclusions: Our study demonstrates, for the first time, predominant epigenetic silencing of DLEC1 in ESCC, and a novel mechanistic link of epigenetic DLEC1 disruption with oncogenic STAT3 signaling in multiple carcinomas.
Keywords: DLEC1, 3p22, STAT3, methylation, carcinoma
 Global reach, higher impact
Global reach, higher impact