13.3
Impact Factor
Theranostics 2017; 7(14):3478-3488. doi:10.7150/thno.20717 This issue Cite
Research Paper
CBX8 Suppresses Tumor Metastasis via Repressing Snail in Esophageal Squamous Cell Carcinoma
1. State Key Laboratory of Oncology in South China, Collaborative Innovation Center for Cancer Medicine, Sun Yat-Sen University Cancer Center, Guangzhou 510060, P. R. China;
2. Guangdong Esophageal Cancer Research Institute, Guangzhou 510060, Guangdong, P. R. China;
3. Department of Gastroenterology, Jiangxi Cancer Hospital, Nanchang 330029, P.R. China.
* These authors contributed equally to this work.
Received 2017-4-24; Accepted 2017-6-15; Published 2017-8-15
Abstract
The poor clinical outcome and prognosis of esophageal squamous cell carcinoma (ESCC) is mainly attributed to its highly invasive and metastatic nature, making it urgent to further elicit the molecular mechanisms of the metastasis of ESCC. The function of each polycomb chromobox (CBX) protein in cancer is cell-type-dependent. Although CBX8 has been reported to promote the esophageal squamous cell carcinoma (ESCC) tumorigenesis, its role in ESCC metastasis has not been explored yet. In this study, we report that the inhibition of cell migration, invasion, and metastasis in ESCC requires CBX8-mediated repression of Snail, a key transcription factor that induces epithelial-to-mesenchymal transition (EMT), and that CBX8 inversely correlated with Snail in the ESCC tissues. Moreover, this novel function of CBX8 is dependent on its binding with the Snail promoter, which in turn suppresses the transcription of Snail. Collectively, CBX8 may play paradoxical roles in ESCC, inhibiting metastasis while promoting cell proliferation.
Keywords: CBX8, Snail, EMT, Tumor metastasis, Esophageal squamous cell carcinoma.
Introduction
Esophageal squamous cell carcinoma (ESCC), the predominant pathological type of esophageal cancer in the East Asian, is one of the most frequent malignant cancers and the fourth leading cause of cancer-related death in China [1, 2]. Despite considerable advances achieved in diagnosis and multimodality therapies, the prognosis of ESCC is still poor with a dismal 5-year survival rate of around 30% [3-5]. The high incidence of lymphatic metastasis remains a major challenge in the management of ESCC [6, 7]. However, the precise mechanisms underlying the metastasis of ESCC remain to be elucidated. Therefore, it is imperative to identify potential molecular biomarkers for the diagnosis and treatment of metastatic ESCC.
Polycomb group (PcG) proteins as major epigenetic regulators are assembled into two complexes, PRC1 and PRC2, which are involved in gene silencing via modifying the chromatin [8-12]. The polycomb chromobox proteins (CBXs), including five members (CBX2, CBX4, CBX6, CBX7 and CBX8), have been shown to participate in the PRC1 complex and provide PRC1 distinguish functions, suggesting that the roles of CBX proteins in cancer may be context-dependent [13-15]. For instance, CBX4 promotes the transcription activity of HIF-1α, thus inducing VEGF expression and angiogenesis by increasing the SUMOylation of HIF-1α in hepatocellular carcinoma (HCC) [16], whereas CBX4 is a tumor suppressor in colorectal carcinoma (CRC) via recruiting HDAC3 to the Runx2 promoter to impede Runx2 expression [17]; CBX7 acts as a tumor suppressor in lung carcinoma by recruiting HDAC2 to the CCNE1 promoter to suppress CCNE1 expression [18], while acts as an oncogene in gastric cancer and lymphoma [19, 20]. Therefore, the function of each CBX protein should be assessed separately in any cancer type.
As a transcriptional repressor, CBX8, also known as human polycomb 3 (HPC3), has been reported to have non-canonical functions [14]. For examples, CBX8 is required for MLL-AF9 induced leukemogenesis through its interactions with oncogenes [8, 21], whereas its interaction with WD repeat domain 5 (WDR5) promotes mammary tumorigenesis [22]. Recent reports have indicated that CBX8 may promote tumorigenesis in ESCC [23, 24], but its potential role in ESCC metastasis remains unknown. Given that CBX8 exerts paradoxical effects, promoting proliferation while suppressing metastasis in CRC [25], we were very curious to determine the role of CBX8 in ESCC metastasis. As shown in this report, CBX8 may serve as a tumor suppressor in ESCC metastasis by directly inhibiting the Snail promoter activity, even though it can promote cell proliferation in ESCC.
Materials and Methods
Cell lines and culture
All cells were incubated in humidified air at 37°C with 5% CO2. Human ESCC cell line TE-1 was acquired from Cell Bank of Chinese Academy of Sciences (Shanghai, China). Four cell lines (KYSE30, KYSE140, KYSE180, KYSE410) were kindly provided by Prof. Guan [26]. The Chinese ESCC cell line HKESC1 and an immortalized esophageal epithelial cell line NE-1 were gifts from Prof. Tsao (University of Hong Kong). All ESCC cells were maintained in RPMI-1640 medium (Gibco, USA) supplemented with 10% fetal bovine serum (Gibco, USA), 100 units/ml penicillin and 100 mg/ml streptomycin (Beyotime Biotech, China). An embryonic kidney cell line 293T was obtained from the American Type Culture Collection (ATCC) and cultured according to its instructions. All cell lines used in this study were regularly authenticated by morphological observation and haven't been in culture for more than 2 months.
Clinical samples
Those patients recruited in our study haven't received radiotherapy or chemotherapy before surgery. All ESCC primary tumor tissues from surgical resection were obtained from Sun Yat-Sen University Cancer Center (Guangzhou, China). All processes were approved by the Ethics Committees of Sun Yat-Sen University Cancer Center and written informed consent was obtained from each patient prior to sample collection.
RNA extraction and Quantitative Real-time Polymerase Chain Reaction (qRT-PCR)
These procedures were performed as described previously [25]. Briefly, total RNA was extracted from frozen clinical tissues or cultured cell lines using Trizol reagent (Invitrogen, USA), and 1μg of the total RNA was reverse transcribed with the PrimeScript™ II 1st Strand cDNA Synthesis Kit (Takara, Japan). qRT-PCR was performed using a SYBR Green PCR Kit (Takara, Japan) and an CFX96 Touch Real-Time PCR Detection System (Bio-Rad, USA). The primers used for amplifying CBX8, Snail, Slug, E-Cadherin, N-Cadherin, Fibronectin, Vimentin and GAPDH are listed in Table S1.
Western blotting and antibodies
Total protein was extracted from cultured cells and analyzed as previously described [27, 28]. A total of 50 μg of protein extracts were loaded to 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to PVDF membranes (Millipore, USA). After incubated with the primary antibodies overnight at 4 ℃ and then with HRP conjugated secondary antibodies (Promega, USA) for 1 h at room temperature, the membranes were subjected to the ECL chemiluminescence system (Pierce, USA). All of the experiments were repeated thrice. Primary antibodies used: CBX8 from Sigma Aldrich; Snail, Slug and Vimentin from BD Biosciences; E-cadherin and N-cadherin from Cell Signaling Technology; CBX6, Fibronectin and GAPDH from Santa Cruz.
Plasmids
The CBX8 and Snail short hairpin RNA (shRNA) pLKO1 lentiviral constructs were purchased from Sigma Addrich. The following oligonucleotides for human CBX8 and Snail: shCBX8 #1, 5'-GCGTGAGCTTGGCATAGTGAT-3'; shCBX8 #2, 5'-GCATGGAATACCTCGTGAAAT-3'; shSnail #1, 5'-GCAAATACTGCAACAAGGAAT-3'; shSnail #2, 5'-TGCTCCACAAGCACCAAGAGT-3'. The full-length cDNAs of human CBX8 was cloned into pMSCV retroviral vector. Snail and flag-tagged CBXs (2, 4, 6, 7, 8) were cloned into the pcDNA3.1 vector. The promoter region of Snail was cloned into pGL3.0 basic vector.
Establishment of stable cell lines
The indicated viral constructs were packaged in 293T cells. Cell lines stably expressing negative control shRNA (NC), CBX8-shRNA, Snail-shRNA or both were established by the Sigma Addrich shRNA System according to the manufacturer's instructions. CBX8- expressing retrovirus was used to stably transfect HKESC1 to establish CBX8-overexpressing cells. Empty vector-transfected cells were established as controls. The efficiency of gene silencing or overexpression was confirmed by qRT-PCR and western blotting.
Wound-healing assay and cell migration assay and invasion assays
As described previously, cell motility was assessed by measuring the movement of cells into a cellular area wounded with a 10 μl pipette tip, and the spread of wound gap closure was observed at 0 h, 36 h and 72 h after scratching and photographed under a microscope. The cell migration and invasion assays were performed using chambers (BD Biosciences, USA) containing polyethylene terephthalate membranes of 8 μm pore size. Cells were suspended in serum-free medium and added to the upper chamber, which was coated with (invasion assay) or without (migration assay) the matrigel mix. And culture medium with 20% fetal bovine serum was placed into the lower chamber. After 36 h of incubation, the migrated or invaded cells on the bottom of the membrane were fixed, stained, and counted from six random fields under 10 × objective lens using an inverted microscope (Olympus, Japan). All experiments were performed in duplicate and repeated three times.
MTT assay
The 3-(4, 5-dimethyl-2-thiazolyl)-2, 5-diphenyl-2-H-tetrazolium bromide (MTT) assay was conducted to examine the viability of ESCC cells. The viability of the transfected cells was assessed at five time points (on days 1, 2, 3, 4, and 5) after seeding into 96-well culture plates at the density of 1000 cells/well. 20 µl of MTT (Sigma-Aldrich, USA) reagent (5 mg/mL) was added into each well, and the plates were incubated for 4 h. After discarding the supernatant, 150 µl DMSO was added to dissolve the formazan product. Then the optical density (OD) value was measured by a microplate reader at 490 nm. Each group was comprised of six duplicated wells, and the assay was performed three times independently.
The tail vein-lung metastasis model
Animal experiments were performed in accordance with protocols approved by the Ethics Committees of Sun Yat-Sen University Cancer Center. For the tail vein- lung metastasis model, the 4-week-old SPF grade nude mice were obtained from the SLAC Laboratory Animal Center (Shanghai, China) and kept in certified vivarium facility at constant temperature (23 ± 2 °C) and humidity (50-70%) with a 12-h light-dark cycle. Briefly, 2× 106 cells suspended in 100μl normal saline were injected intravenously through the tail vein with an insulin syringe. After 6 weeks, the mice were sacrificed and lung metastatic nodules were carefully examined and counted.
Immunofluorescence
Briefly, cells were fixed with 4% paraformaldehyde for 15 min, permeabilized with 0.5% Triton X-100 for 15 min and blocked with 3% bovine serum albumin for 1 h at room temperature. Then, cells were incubated with phalloidin conjugated with rhodamine (Sigma-Aldrich, USA) for 30 min to stain the F-Actin, followed by incubation for 2 min with 4,6-diamidino-2-phenylindole (DAPI) to label the nuclei. Images (1000 ×) were captured using a confocal scanning microscope (Olympus, Japan).
Luciferase reporter and chromatin immunoprecipitation (ChIP) assays
The luciferase reporter assay was carried out as described previously [28, 29]. Cells were plated in 12-well plates and transfected with 0.8 μg of promoter-luciferase plasmid. In the same time, the cells were co-transfected with 8 ng of renilla luciferase. The expression levels of firefly and renilla luciferases were analyzed 48 h after transfection using the Dual-Luciferase Assay kit (Promega, USA). All experiments were performed in triplicate.
For the ChIP assay, the EZ-Magna ChIP™ A/G One-Day Chromatin Immunoprecipitation Kit was used. According to the manufacturer's instructions, adherent cells at 80-90 % confluence in a 150 mm culture dish were fixed with formaldehyde, lysed and sonicated to generate DNA fragments of 200-1000 bp (confirmed by agarose gel electrophoresis). Then, the cell lysate were subjected to immunoprecipitation overnight using anti-CBX8 antibody. Purified DNA was resuspended in TE buffer for qRT-PCR. The primers are listed in Table S2.
Statistical analysis
All data were expressed as mean ± SD of at least three independent experiments. Student's t-test was used when data were normally distributed and the Mann-Whitney test was applied when the data were independently distributed. The value of p < 0.05 was considered statistically significant.
Results
CBX8 suppresses cell migration, invasion and metastasis, but promotes cell proliferation in ESCC
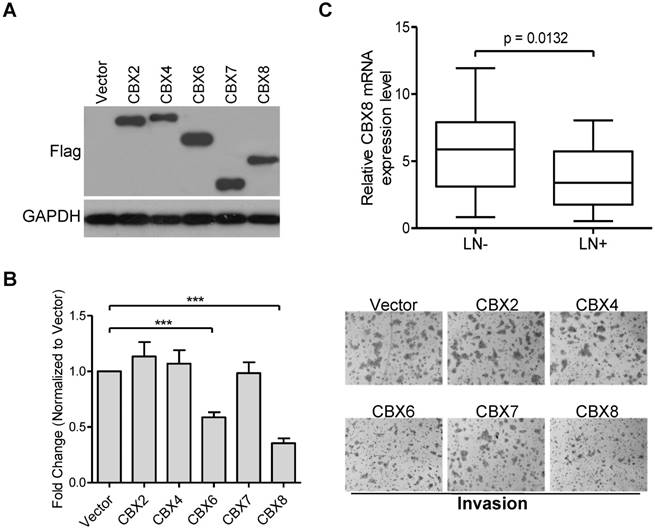
CBX proteins may act as an oncogene or tumor suppressor in a cell-type dependent manner [13], and we have recently reported that CBX8 exerts paradoxical effects in CRC, promoting proliferation while suppressing metastasis [25]. However, the potential role of each CBX protein in ESCC progression remains to be determined. Initially, we ectopically expressed Flag-tagged CBX2, CBX4, CBX6, CBX7 or CBX8 in HKESC1 cells (Fig. 1A), and found that both CBX6 and CBX8, but not other CBX members, repressed cell invasion in HKESC1 cells (Fig. 1B). Compared with CBX6, CBX8 showed a more significant inhibition of HKESC1 cell invasion, we were focusing on the roles of CBX8 in ESCC metastasis in this study. More interestingly, as shown in Fig. 1C, using qRT-PCR, CBX8 was significantly down-regulated in the ESCC primary tissues with lymph node metastasis (55%, 27/49) compared in those without lymph node metastasis (45%, 22/49). These results suggest that CBX8 may play a role in the ESCC metastasis.
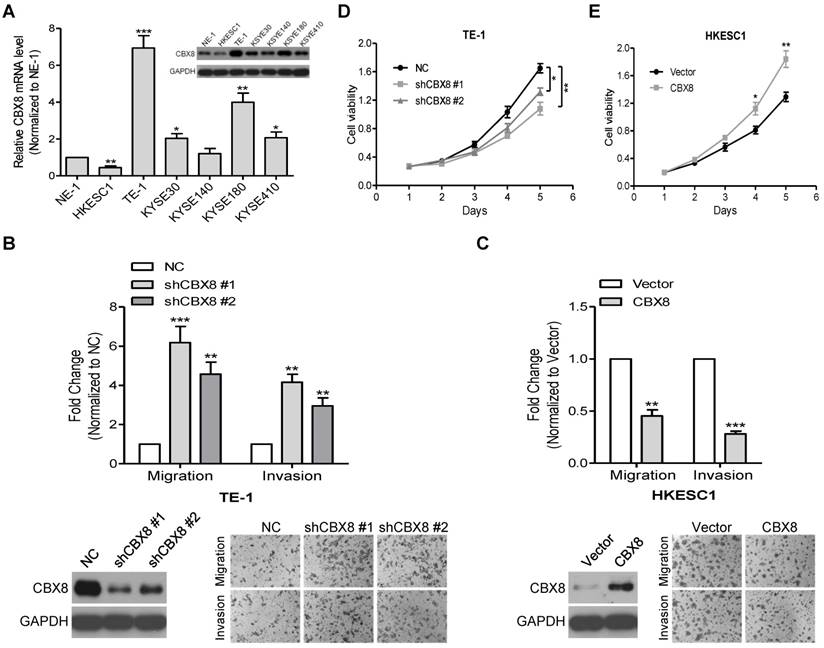
Indeed, CBX8 were differently detected at both mRNA and protein levels among the tested ESCC cell lines (Fig. 2A). Next, we used two pairs of small hairpin RNAs (shRNAs) to dramatically reduce endogenous CBX8 expression in TE-1 cells, which endogenously express a high level of CBX8, and this knockdown significantly enhanced the migration and invasion of TE-1 cells (Fig. S1 and Fig. 2B). Conversely, the stably ectopic expression of CBX8 dramatically inhibited the migration and invasion in HKESC1 cells, which endogenously express a low level of CBX8 (Fig. 2C). However, as demonstrated by the MTT assay, the knockdown of CBX8 impaired the viability of TE-1 cells (Fig. 2D), whereas the stable ectopic expression of CBX8 increased the viability of HKESC1 cells (Fig. 2E), which are consistent with previous reports showing that CBX8 may promote the ESCC tumorigenesis [23, 24].
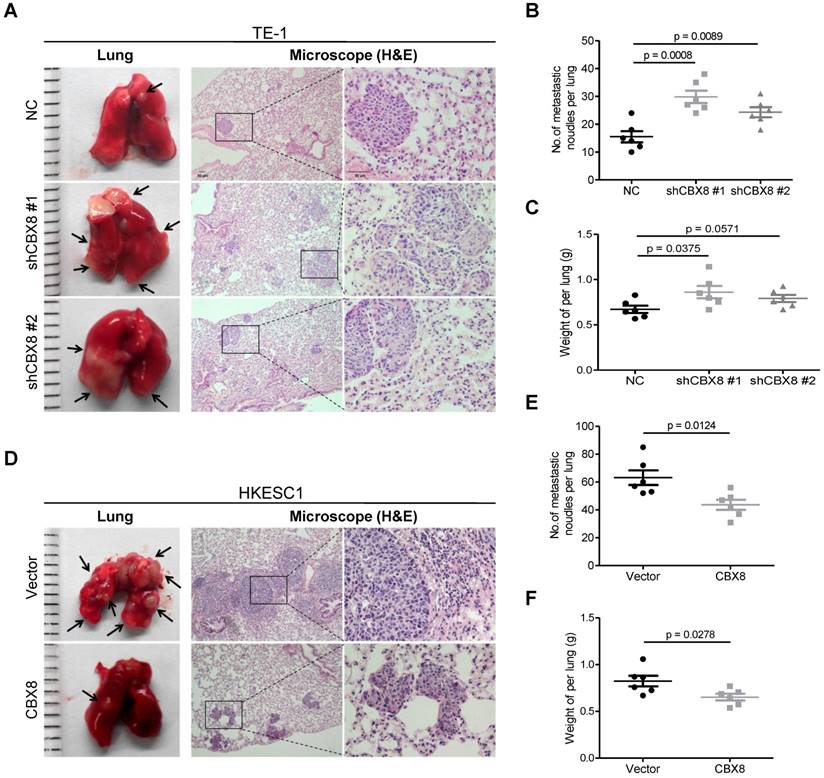
Furthermore, knocking down CBX8 increased the tumor size, the number of metastatic nodules and the lung weight (Fig. 3A-C) when TE-1 cells were injected into the tail veins of the 4-week-old nude mice. Consistently, overexpression of CBX8 decreased these parameters (Fig. 3D-F) when HKESC1 cells were injected into the tail veins of mice. Taken together, CBX8 may play paradoxical roles in ESCC, promoting cell proliferation while repressing metastasis.
Ectopic expression of CBX6 or CBX8, but not of other CBX members, represses HKESC1 cells invasion ability, and CBX8 expression is reduced in patients with lymph node metastasis. (A) HKESC1 cells were transfected with Vector or Flag-tagged CBXs for 36 h and then subjected to Western blotting. GAPDH was the loading control. (B) Cell invasion assay was performed in the indicated cells as described in the Materials and methods. Representative images (100 ×) are presented (right panel). The results are expressed as the mean ± SD of three independent experiments (left panel). *** p < 0.001 (Student's t-test). (C) The relative mRNA levels of CBX8 in 27 and 22 patients with (LN+) and without (LN-) lymph node metastasis was detected by qRT-PCR, respectively. p = 0.0132 (Mann-Whitney test).
CBX8 plays paradoxical roles in ESCC, suppressing cell migration and invasion while promoting proliferation. (A) The expression of CBX8 in ESCC cell lines was detected at mRNA and protein levels by qRT-PCR and Western blotting (insertion panel), respectively. An immortalized esophageal epithelial cell line NE-1 was used as control. Data from three independent experiments are expressed as mean ± SD. ** p < 0.01 and *** p < 0.001, Student's t-test. (B, C) Both the cell migration and invasion assays were performed in the indicated stable cell lines as described in the Materials and methods. Representative images (100 ×) are shown. The columns, mean of three independent experiments; the bars, SD. ** p < 0.01 and *** p < 0.001 using Student's t-test. (D, E) The cell viability of the indicated stable cell lines was measured in different time points as indicated by MTT assay. Data from three independent experiments are expressed as mean ± SD. * p < 0.05 and ** p < 0.01, Student's t-test.
CBX8 impedes epithelial-mesenchymal transition (EMT) in ESCC cells
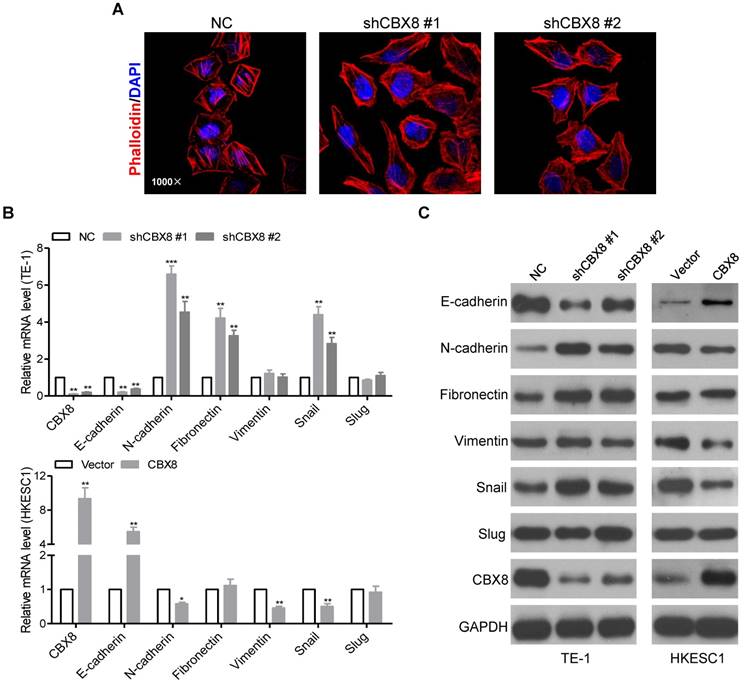
We noticed that the morphology of stable CBX8-shRNAs TE-1 cells have been changed from the tight cell-to-cell adhesion into the spindle-like and fibroblastic phenotype, suggesting that TE-1 cells knockdown of CBX8 occur EMT [30]. The morphological changes were further supported by phalloidin staining, indicating that the cytoplasmic bundles of Actin fibers were markedly reduced in shCBX8-treated TE-1 cells, instead showing cortical localization of F-Actin (Fig. 4A). Moreover, both the epithelial cell marker (E-cadherin) and the mesenchymal cell markers (N-cadherin, Fibronectin, and Vimentin) were tested in the cells stably knockdown or overexpressed of CBX8. As shown in Fig. 4B-C, E-cadherin was dramatically reduced, while N-cadherin and Fibronectin were up-regulated in CBX8-shRNAs TE-1 cells at both the mRNA and the protein levels. On the contrary, overexpression of CBX8 increased E-cadherin while inhibited N-cadherin and Vimentin in HKESC1 cells at both mRNA and protein levels. Collectively, these results suggest that CBX8 negatively modulates metastasis probably through regulating the EMT program in ESCC cells.
CBX8, but not other CBX members, specifically binds to the Snail promoter and suppresses its expression in ESCC cells
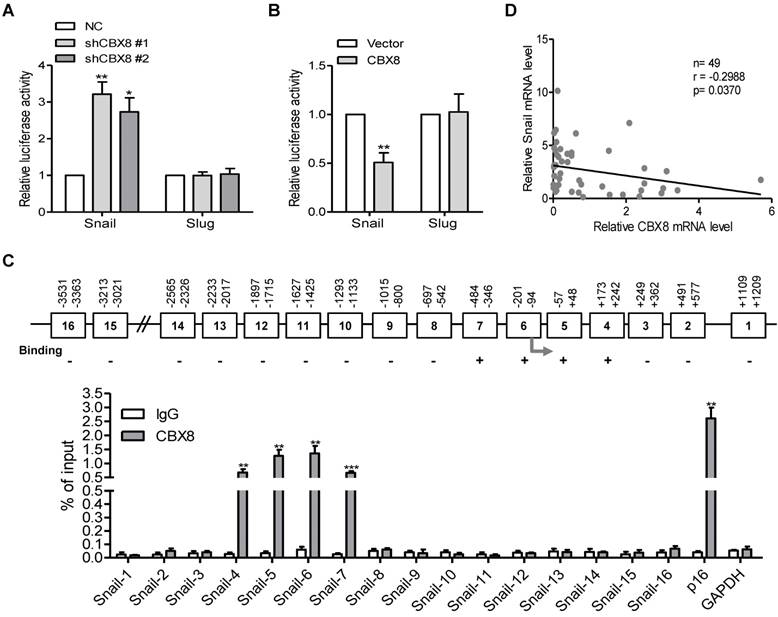
Our previous report showed that CBX8 affects the metastasis of colorectal carcinoma in part through direct up-regulation of integrin β4 [25], we tested if this effect was the case in ESCC cells. Unfortunately, knockdown or overexpression of CBX8 did not affect the expression of integrin β4 (data not shown). Given that Snail and Slug are two major transcription factors for EMT [31-33], their mRNA and protein levels were then tested among these established stable cell lines. Strikingly, Snail, but not Slug, was increased in CBX8-silenced TE-1 cells and was reduced in HKESC1 cells stably overexpressing CBX8 (Fig. 4B-C), indicating that CBX8 may repress Snail to regulate the EMT program. This promoted us to test if CBX8 modulate Snail at transcriptional level, as CBX8 generally functions through the PRC1. To this end, the luciferase reporter assays for both Snail and Slug promoters were performed in these established cell lines. Interestingly, the promoter luciferase activity of Snail, but not Slug, was increased in the CBX8-shRNAs TE-1 cells while was impaired in the HKESC1 cells stably expressing CBX8 (Fig. 5A-B). Furthermore, using chromatin immunoprecipitation (ChIP) assay, we found that CBX8 was clearly associated with several regions of the Snail promoter (Fig. 5C).
CBX8 suppresses the metastasis of ESCC cells in nude mice. (A, D) The effects of CBX8 on tumor metastasis were evaluated by the tail vein-lung metastasis model. Left panel, representative images of lungs derived from mice injected with the indicated stable cell lines; right panel, H&E staining (Scale bars = 50 μm) of pulmonary sections from the tumor-bearing mice. (B, C, E, F) Numbers (B, E) and wet weights (C, F) of metastatic nodules per lung in the nude mice were summarized (n = 6). The bars indicate the SD. A two-tailed Student's t-test was used for statistical analysis.
CBX8 inhibits EMT in ESCC cells. (A) TE-1 cells stably expressing NC or shRNA-CBX8 as indicated were analyzed by immunofluorescence assay for phalloidin (red) with DAPI (blue) counterstaining. (B, C) Relative expressions of the indicated molecules were detected by qRT-PCR (B) and Western blotting (C) in the indicated stable cell lines, respectively. The results are expressed as the mean ± SD of three independent experiments. * p < 0.05, ** p < 0.01 and *** p < 0.001, Student's t-test.
Considering that the target genes of CBX8 and its paralogous proteins were largely overlapping in some cases [34, 35], we sought to explore whether other CBX proteins are associated with down-regulation of Snail, and the expression levels of Snail were compared between empty Vector- and Flag-tagged CBXs-transfected cells by Western blotting. The results showed that the protein level of Snail was only significantly decreased in the CBX8-expressing cells, but not in other CBXs-expressing cells (Figure S2A). Moreover, co-transfection of CBX8, but not other family members, dramatically impaired the Snail reporter luciferase activity in HKESC1 cells (Figure S2B). Furthermore, using qRT-PCR, mRNA levels of CBX8 and Snail in the ESCC tumor tissues were inversely correlated (Fig. 5D). Taken together, these results indicate that CBX8 directly binds to the Snail promoter and suppresses its expression in ESCC cells.
Inhibition of cell mobility by CBX8 depends on its down-regulation of Snail in ESCC cells
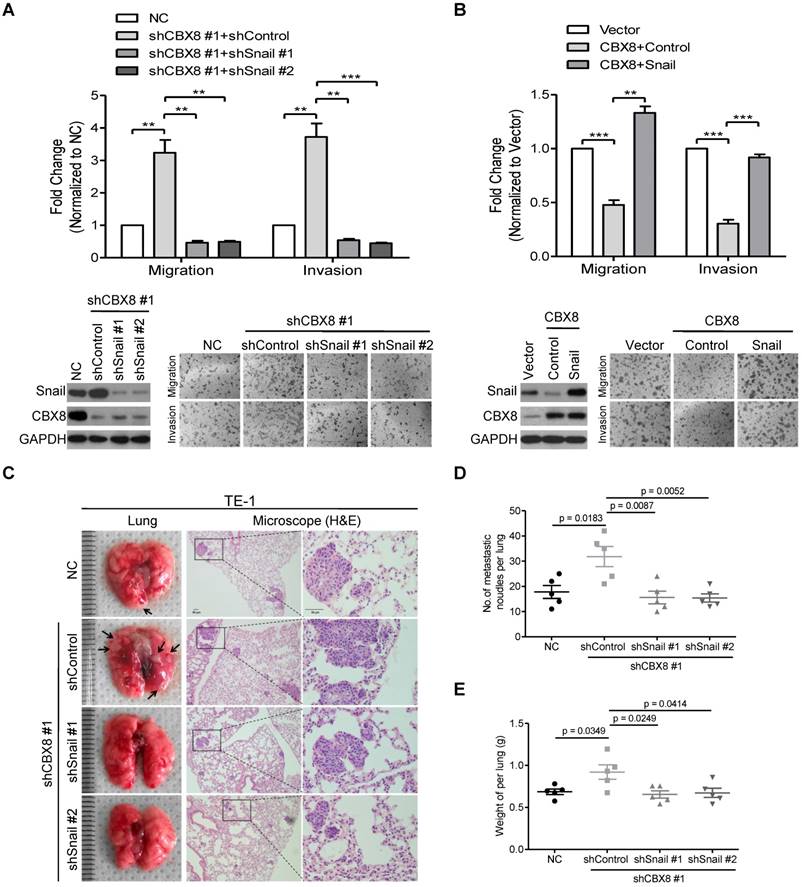
Next, we sought to determine whether CBX8 suppressed cell mobility via its inhibition of Snail. Using two pairs of shRNAs to stably knock down Snail in the CBX8-silenced TE-1 cells, we found that the down-regulation of Snail abrogated the increase of cell migration and invasion in TE-1 cells induced by knocking down CBX8 (Fig. 6A). On the other hand, Snail was transiently transfected into the CBX8-overexpressed HKESC1 cells, and the migratory and invasive abilities were almost rescued to the original levels in the HKESC1 cells up-regulated of both Snail and CBX8 (Fig. 6B). Moreover, using the tail vein-lung metastasis model, the inhibition of ESCC metastasis to the lungs by CBX8 was abolished in the cells bearing stably knocking down of both CBX8 and Snail (Fig. 6C-E). These results demonstrate that CBX8 suppressed cell mobility via its impairment of Snail in ESCC cells.
Discussion
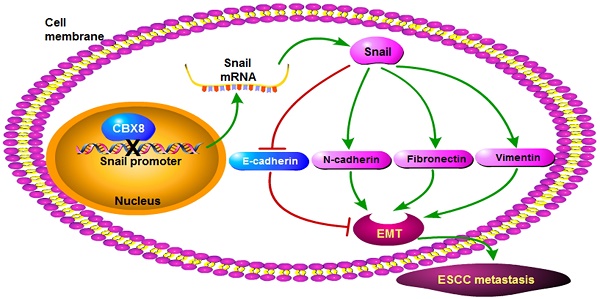
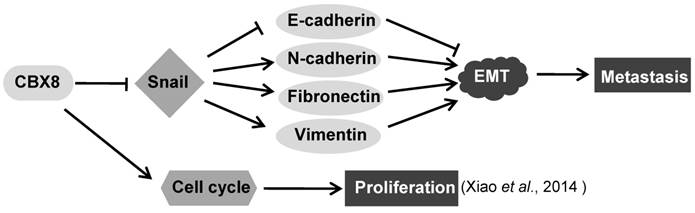
In this report, we have demonstrated that CBX8 had paradoxical effects in ESCC progression, by suppressing tumor metastasis while promoting proliferation (Fig. 7). Overall, CBX8 may function as a tumor suppressor in ESCC, as CBX8 inhibits EMT and metastasis of ESCC. This is the first evidence that CBX8 modulates the EMT progression and tumor metastasis in ESCC, and this new function of CBX8 is dependent on Snail.
All CBX proteins share highly conserved chromodomain and Pc boxes, but their different sizes and motifs confer their different functions, and cells deficient for different CBX proteins show different binding affinities to chromatin [36]. For instance, H3K27me3 contributes significantly to the targeting of CBX7 and CBX8 to chromatin, but less to CBX2, CBX4, and CBX6 [37]. In stem cells, the autoregulatory loop among CBX members is crucial for both self-renewal and differentiation [34, 38, 39]. Studies indicated that CBX7 is down-regulated and CBX2, CBX4, and CBX8 are up-regulated simultaneously during embryonic stem cells differentiation. Nevertheless, in ESCC cells, the inhibition of Snail and subsequent inactivation of EMT are specifically regulated by CBX8, not its paralogous proteins (Fig. S2). Though overexpression of CBX6 or CBX8 impeded cell invasion in HKESC1 cells, knockdown or overexpression of CBX8 doesn't affect the expression of CBX6 in ESCC cell lines (Fig. S3). Thus, the tumor suppressive role for CBX8 in ESCC metastasis may be independent of other paralogous proteins.
CBX8 directly binds to and down-regulates the Snail promoter activity. (A, B) The indicated stable cell lines were transfected with Snail or Slug promoter linked to luciferase for 48 h and then subjected to the luciferase assay as described in Materials and Methods. (C) The upper panel, a schematic illustration of the Snail promoter regions (1-16) with (+) or without (-) binding affinity for CBX8. The arrow indicates the transcriptional start site (TSS). The lower panel, the result of ChIP-quantitative PCR analysis of CBX8 binding to the distinct regions in the Snail promoter, showing enrichment with CBX8 antibody compared with IgG control. The p16 and GAPDH promoters were used as the positive and negative controls, respectively. The results are expressed as the mean ± SD of individual samples from three independent experiments. ** p < 0.01 and *** p < 0.001, Student's t-test. (D) There was a reverse correlation between CBX8 and Snail at mRNA level in ESCC tissue samples. p = 0.0215, linear regression analysis.
The inhibitory effect of CBX8 on cell migration and invasion depends on the down-regulation of Snail. (A, B) Both the cell migration and invasion assays were performed in the indicated stable cell lines as described in the Materials and methods. Representative images (100 ×) are shown. The columns, mean of three independent experiments; The bars indicate the SD. ** p < 0.01 and *** p < 0.001 using Student's t-test. (C-E) An in vivo lung metastasis model was established in nude mice using TE-1 cells stably expressing NC, shControl, shCBX8, shSnail or both, as indicated, as described in Materials and Methods. (C) Representative images of gross and H&E staining (Scale bars = 50 μm) of metastatic lung nodules. Numbers (D) and wet weights (E) of metastatic nodules per lung in the nude mice were summarized (n = 5). The bars indicate the SD. A two-tailed Student's t-test was used for statistical analysis.
CBX8 is one of the five human homologs of the Drosophila Pc protein, and participates in the PRC1 complex and drives the PRC1 complex to the promoter regions of CBX8 target genes to mediate epigenetic gene silencing [8, 40]. However, recent evidences have shown that CBX proteins can associate with proteins other than the PRC1 complex, thereby playing the PRC1-independent roles in regulating the gene transcription. For instance, the interaction of CBX4 with HDAC3 plays crucial roles in its impairment of both the Runx2 transcription and metastasis in CRC [17]; CBX8 associates with the non-PRC1 complexes containing H3K4 methyltransferase complex component WDR5 to regulate Notch gene expression [22]; CBX8 interacts with MLL-AF9 and TIP60, and this interaction is required for MLL-AF9-induced transcriptional activation and leukemogenesis [8]. Herein, we have showed that Snail, a key regulator of the EMT program, is a direct target of CBX8, and that CBX8 decreases the mesenchymal cell markers (N-cadherin, Fibronectin, and Vimentin) and increases the epithelial cell marker E-cadherin, which consequently impair cell migration and invasion and metastasis in ESCC, as proposed in Fig. 7. In fact, the regulation of CBX8 on cancer metastasis have been also reported in CRC, showing that the knockdown of CBX8 releases its inhibition on the ITGB4 promoter, which in turn reduces active RhoA and enhances CRC metastasis [25].
Recently, there are increasing evidences show that CBX8 promotes tumorigenesis in different type of cancer, including leukemia [8], breast cancer [22], colorectal carcinoma [25] and ESCC [23, 24]. However, our current report shows that CBX8 may function as a tumor suppressor in ESCC metastasis. These paradoxical functions in ESCC proliferation and metastasis for CBX8 indicate that CBX8 may play essential role in early stages of ESCC when proliferation is dominant, while CBX8 may inhibit cancer cell invasion and metastasis at late stages of ESCC. These findings are quite similar to our previous report of the paradoxical roles of CBX8 in colorectal carcinoma [25]. Notably, the paradoxical functions in proliferation and metastasis for some molecules have been reported. For instance, MYC, a well-established proliferation marker, suppresses cancer metastasis through the direct transcriptional repression of integrin subunits [41, 42]; Cyclin A2, another proliferation promoter, negatively controls cell motility by promoting RhoA activation and cytoskeletal rearrangements [43].
In summary, our studies shed new light on the metastasis-suppressor role of CBX8 in ESCC. Although the detailed underlying mechanism requires further investigations, the CBX8/Snail axis modulating EMT, as shown here, may provide an explanation for the unexpected correlation between CBX8 and ESCC metastasis.
The proposed model for the functions of CBX8 in ESCC metastasis and proliferation. CBX8 may down-regulate Snail to increase E-cadherin and to decrease N-cadherin, Fibronectin and Vimentin, which in turn impair EMT and consequently repress metastasis in ESCC. In the meanwhile, CBX8 may positively modulate the ESCC cell cycle progression and proliferation as indicated by Xiao et al. [24].
Acknowledgements
We thank the kind gifts of ESCC cell lines from both Prof. Guan (The University of Hong Kong) and Prof. Tsao (The University of Hong Kong). This work was supported by Guangdong Esophageal Cancer Research Institute [M201515 to T. K.], the National Key Research and Development Program of China [2016YFC0904601 and 2016YFA0500304 to T. K.], Natural Science Foundation of Guangdong Province [2016A030310218 to W. Y.], the Fundamental Research Funds for the Central Universities [16ykpy39 to W. Y.] and the National Nature Science Foundation in China (NSFC) [81530081 and 31571395 to T. K.].
Authors' contributions
Kang T conceived the idea. Wang G, Tang J and Zhan W performed the experiments. Wang G, Tang J, Zhan W, Zhang R, Zhang M, Wu Y, Liao D, Wang X analyzed the data. Kang T and Wang G wrote the manuscript. All co-authors have seen and approved this version of the manuscript.
Supplementary Material
Supplementary figures and tables.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Parkin DM, Bray F, Ferlay J, Pisani P. Global Cancer Statistics, 2002. CA: A Cancer Journal for Clinicians. 2005;55:74-108
2. Kamangar F, Dores GM, Anderson WF. Patterns of Cancer Incidence, Mortality, and Prevalence Across Five Continents: Defining Priorities to Reduce Cancer Disparities in Different Geographic Regions of the World. Journal of Clinical Oncology. 2006;24:2137-50
3. Gertler R, Stein HJ, Langer R, Nettelmann M, Schuster T, Hoefler H. et al. Long-term Outcome of 2920 Patients With Cancers of the Esophagus and Esophagogastric Junction. Annals of Surgery. 2011;253:689-98
4. Jemal A, Siegel R, Ward E, Murray T, Xu J, Thun MJ. Cancer Statistics, 2007. CA: A Cancer Journal for Clinicians. 2007;57:43-66
5. Allum WH, Stenning SP, Bancewicz J, Clark PI, Langley RE. Long-Term Results of a Randomized Trial of Surgery With or Without Preoperative Chemotherapy in Esophageal Cancer. Journal of Clinical Oncology. 2009;27:5062-7
6. Enzinger PC, Mayer RJ. Esophageal Cancer. New England Journal of Medicine. 2003;349:2241-52
7. Chen G, Wang Z, Liu X-y, Liu F-y. Recurrence Pattern of Squamous Cell Carcinoma in the Middle Thoracic Esophagus after Modified Ivor-Lewis Esophagectomy. World Journal of Surgery. 2007;31:1108-15
8. Tan J, Jones M, Koseki H, Nakayama M, Muntean AG, Maillard I. et al. CBX8, a Polycomb Group Protein, Is Essential for MLL-AF9-Induced Leukemogenesis. Cancer Cell. 2011;20:563-75
9. Margueron R, Reinberg D. The Polycomb Complex PRC2 and its Mark in Life. Nature. 2011;469:343-9
10. Simon JA, Kingston RE. Mechanisms of Polycomb gene silencing: knowns and unknowns. Nat Rev Mol Cell Biol. 2009;10:697-708
11. Laible G, Wolf A, Dorn R, Reuter G, Nislow C, Lebersorger A. et al. Mammalian homologues of the Polycomb-group gene Enhancer of zeste mediate gene silencing in Drosophila heterochromatin and at S. cerevisiae telomeres. The EMBO Journal. 1997;16:3219-32
12. Morey L, Helin K. Polycomb group protein-mediated repression of transcription. Trends in Biochemical Sciences. 2010;35:323-32
13. Gil J, O'Loghlen A. PRC1 complex diversity: where is it taking us? Trends in Cell Biology. 2014;24:632-41
14. Kerppola TK. Polycomb group complexes - many combinations, many functions. Trends in Cell Biology. 2009;19:692-704
15. Vandamme J, Volkel P, Rosnoblet C, Le Faou P, Angrand PO. Interaction Proteomics Analysis of Polycomb Proteins Defines Distinct PRC1 Complexes in Mammalian Cells. Molecular & Cellular Proteomics. 2011;10:M110.002642-M110
16. Li J, Xu Y, Long X-D, Wang W, Jiao H-K, Mei Z. et al. Cbx4 Governs HIF-1α to Potentiate Angiogenesis of Hepatocellular Carcinoma by Its SUMO E3 Ligase Activity. Cancer Cell. 2014;25:118-31
17. Wang X, Li L, Wu Y, Zhang R, Zhang M, Liao D. et al. CBX4 Suppresses Metastasis via Recruitment of HDAC3 to the Runx2 Promoter in Colorectal Carcinoma. Cancer Research. 2016;76:7277-89
18. Forzati F, Federico A, Pallante P, Abbate A, Esposito F, Malapelle U. et al. CBX7 is a tumor suppressor in mice and humans. The Journal of Clinical Investigation. 2012;122:612-23
19. Zhang X-W, Zhang L, Qin W, Yao X-H, Zheng L-Z, Liu X. et al. Oncogenic role of the chromobox protein CBX7 in gastric cancer. Journal of Experimental & Clinical Cancer Research. 2010;29:114
20. Scott CL, Gil J, Hernando E, Teruya-Feldstein J, Narita M, Martinez D. et al. Role of the chromobox protein CBX7 in lymphomagenesis. Proc Natl Acad Sci USA. 2007;104:5389-94
21. Bardos JI. HPC3 Is a New Human Polycomb Orthologue That Interacts and Associates with RING1 and Bmi1 and Has Transcriptional Repression Properties. Journal of Biological Chemistry. 2000;275:28785-92
22. Chung C-Y, Sun Z, Mullokandov G, Bosch A, Qadeer Zulekha A, Cihan E. et al. Cbx8 Acts Non-canonically with Wdr5 to Promote Mammary Tumorigenesis. Cell Reports. 2016;16:472-86
23. Zhang L, Zhou Y, Cheng C, Cui H, Cheng L, Kong P. et al. Genomic Analyses Reveal Mutational Signatures and Frequently Altered Genes in Esophageal Squamous Cell Carcinoma. The American Journal of Human Genetics. 2015;96:597-611
24. Xiao W, Ou C, Qin J, Xing F, Sun Y, Li Z. et al. CBX8, a novel DNA repair protein, promotes tumorigenesis in human esophageal carcinoma. International Journal of Clinical and Experimental Pathology. 2014;7:4817-26
25. Tang J, Wang G, Zhang M, Li F-y, Sang Y, Wang B. et al. Paradoxical role of CBX8 in proliferation and metastasis of colorectal cancer. Oncotarget. 2014;5:10778-90
26. Zhu Y-H, Fu L, Chen L, Qin Y-R, Liu H, Xie F. et al. Downregulation of the Novel Tumor Suppressor DIRAS1 Predicts Poor Prognosis in Esophageal Squamous Cell Carcinoma. Cancer Research. 2013;73:2298-309
27. Sang Y, Chen M-y, Luo D, Zhang R-H, Wang L, Li M. et al. TEL2 suppresses metastasis by down-regulating SERPINE1 in nasopharyngeal carcinoma. Oncotarget. 2015;6:29240-53
28. Wang L, Sang Y, Tang J, Zhang R-H, Luo D, Chen M. et al. Down-regulation of prostate stem cell antigen (PSCA) by Slug promotes metastasis in nasopharyngeal carcinoma. The Journal of Pathology. 2015;237:411-22
29. Liao D, Zhong L, Duan T, Zhang R-H, Wang X, Wang G. et al. Aspirin Suppresses the Growth and Metastasis of Osteosarcoma through the NF-κB Pathway. Clinical Cancer Research. 2015;21:5349-59
30. Chaffe CL, Weinberg RA. A perspective on cancer cell metastasis. Science. 2011;331:1559-64
31. Craene BD, Berx G. Regulatory networks defining EMT during cancer initiation and progression. Nature Reviews Cancer. 2013;13:97-110
32. Serrano-Gomez SJ, Maziveyi M, Alahari SK. Regulation of epithelial-mesenchymal transition through epigenetic and post-translational modifications. Molecular Cancer. 2016;15:18
33. Villarejo A, Cortés-Cabrera Á, Molina-Ortíz P, Portillo F, Cano A. Differential Role of Snail1 and Snail2 Zinc Fingers in E-cadherin Repression and Epithelial to Mesenchymal Transition. The Journal of Biological Chemistry. 2014;289:930-41
34. Klauke K, Radulovic V, Broekhuis M, Weersing E, Zwart E, Olthof S. et al. Polycomb Cbx family members mediate the balance between haematopoietic stem cell self-renewal and differentiation. Nat Cell Biol. 2013;15:353-62
35. Pemberton H, Anderton E, Patel H, Brookes S, Chandler H, Palermo R. et al. Genome-wide co-localization of Polycomb orthologs and their effects on gene expression in human fibroblasts. Genome biology. 2014;15:R23
36. Whitcomb SJ, Basu A, Allis CD, Bernstein E. Polycomb Group proteins: an evolutionary perspective. Trends in Genetics. 2007;23:494-502
37. Zhen CY, Tatavosian R, Huynh TN, Duc HN, Das R, Kokotovic M. et al. Live-cell single-molecule tracking reveals co-recognition of H3K27me3 and DNA targets polycomb Cbx7-PRC1 to chromatin. eLife. 2016;5:e17667
38. Morey L, Pascual G, Cozzuto L, Roma G, Wutz A, Benitah SA. et al. Nonoverlapping functions of the Polycomb group Cbx family of proteins in embryonic stem cells. Cell stem cell. 2012;10:47-62
39. O'Loghlen A, Muñoz-Cabello Ana M, Gaspar-Maia A, Wu H-A, Banito A, Kunowska N. et al. MicroRNA Regulation of Cbx7 Mediates a Switch of Polycomb Orthologs during ESC Differentiation. Cell stem cell. 2012;10:33-46
40. Valk-Lingbeek ME, Bruggeman SWM, van Lohuizen M. Stem Cells and Cancer. Cell. 2004;118:409-18
41. Liu H, Radisky DC, Yang D, Xu R, Radisky ES, Bissell MJ. et al. MYC suppresses cancer metastasis by direct transcriptional silencing of αv and β3 integrin subunits. Nature Cell Biology. 2012;14:567-74
42. Dang Chi V. MYC on the Path to Cancer. Cell. 2012;149:22-35
43. Arsic N, Bendris N, Peter M, Begon-Pescia C, Rebouissou C, Gadéa G. et al. A novel function for Cyclin A2: Control of cell invasion via RhoA signaling. The Journal of Cell Biology. 2012;196:147-62
Author contact
![]() Corresponding author: Tel.: +86 20 87343182; fax: +86 20 87343170. Email address: kangtborg.cn (T. Kang).
Corresponding author: Tel.: +86 20 87343182; fax: +86 20 87343170. Email address: kangtborg.cn (T. Kang).
 Global reach, higher impact
Global reach, higher impact