13.3
Impact Factor
Theranostics 2017; 7(1):180-195. doi:10.7150/thno.17133 This issue Cite
Research Paper
Exosomes derived from miR-140-5p-overexpressing human synovial mesenchymal stem cells enhance cartilage tissue regeneration and prevent osteoarthritis of the knee in a rat model
1. Department of Orthopaedic Surgery, Shanghai Jiao Tong University Affiliated Sixth People's Hospital, 600 Yishan Road, Shanghai 200233, China;
2. Institute of Microsurgery on Extremities, Shanghai Jiao Tong University Affiliated Sixth People's Hospital, 600 Yishan Road, Shanghai 200233, China.
*Co-first authors: These authors contributed equally to this work.
Received 2016-8-7; Accepted 2016-10-4; Published 2017-1-1
Abstract
OBJECTIVES: Osteoarthritis (OA) is the most common joint disease throughout the world. Exosomes derived from miR-140-5p-overexpressing synovial mesenchymal stem cells (SMSC-140s) may be effective in treating OA. We hypothesized that exosomes derived from SMSC-140 (SMSC-140-Exos) would enhance the proliferation and migration abilities of articular chondrocytes (ACs) without harming extracellular matrix (ECM) secretion.
METHODS: SMSCs were transfected with or without miR-140-5p. Exosomes derived from SMSCs or SMSC-140s (SMSC-Exos or SMSC-140-Exos) were isolated and identified. Proliferation, migration and ECM secretion were measured in vitro and compared between groups. The mechanism involving alternative Wnt signalling and activation of Yes-associated protein (YAP) was investigated using lentivirus, oligonucleotides or chemical drugs. The preventative effect of exosomes in vivo was measured using Safranin-O and Fast green staining and immunohistochemical staining.
RESULTS: Wnt5a and Wnt5b carried by exosomes activated YAP via the alternative Wnt signalling pathway and enhanced proliferation and migration of chondrocytes with the side-effect of significantly decreasing ECM secretion. Highly-expressed miR-140-5p blocked this side-effect via RalA. SMSC-140-Exos enhanced the proliferation and migration of ACs without damaging ECM secretion in vitro, while in vivo, SMSC-140-Exos successfully prevented OA in a rat model.
CONCLUSIONS: These findings highlight the promising potential of SMSC-140-Exos in preventing OA. We first found a potential source of exosomes and studied their merits and shortcomings. Based on our understanding of the molecular mechanism, we overcame the shortcomings by modifying the exosomes. Such exosomes derived from modified cells hold potential as future therapeutic strategies.
Keywords: Osteoarthritis, exosomes, synovial mesenchymal stem cells, Yes-associated protein, extracellular matrix, miR-140-5p.
Introduction
Osteoarthritis (OA) is the most common joint disease throughout the world, affecting approximately 10% of men and 18% of women over 60 years of age [1]. Traditional treatment of OA consists of pain management with arthroplasty for end-stage disease [2]. This method does not reduce the morbidity of the early phase of the disease or address the limits of joint replacement surgery, such as the finite lifespan of the prostheses and the possibility of adverse outcomes [3]. Anatomical factors are strongly associated with OA, such as hip dysplasia [4, 5], femoroacetabular impingement morphology [6, 7], varus (or valgus) knee alignment [8, 9], and leg length inequality [10]. Remarkably, age is the strongest risk factor [11], while knee injury also dramatically increases the risk of OA of the knee by more than four times [12]. It is therefore important to prevent or reverse the progression of early OA before it is too late.
Isolated from tissues such as bone marrow, adipose tissue, muscle, perichondrium, periosteum and recently synovium, mesenchymal stem cells (MSCs) have become a popular choice for repairing damaged tissues [13, 14]. Synovial mesenchymal stem cells (SMSCs) were first isolated from the synovial membrane surrounding joints in 2001[15]. With tissue specificity for cartilage regeneration [16], SMSCs have been regarded as showing the greatest potential for cartilage regeneration research. However, the direct use of stem cells has limitations, such as potential immunological rejection and chromosomal variation [17, 18]. It is therefore important to develop a superior strategy which can fully utilize the advantages of stem cells without the potential risks of direct usage.
Recently, increasing evidence has shown that the activation of resident cells via a paracrine mechanism may play an important role in progenitor cell-mediated tissue regeneration [19, 20]. Exosomes—membrane-bound vesicles from 30 to 150 nm in diameter [21] produced by almost all cell types—may be critical for cell-cell communication [22]. Evidence has shown that exosomes have similar biological function to the cells from which they are derived, and that direct use of these nanoparticles has no obvious adverse effects such as immunogenicity or tumorigenicity [23, 24]. Currently, there are no reports regarding the application of exosomes derived from SMSCs in OA therapy. We hypothesized that exosomes released from SMSCs (SMSC-Exos) could be effective in helping to prevent OA.
In the present study, we used synovial membrane tissue-derived SMSCs as a “factory” to generate exosomes and found that SMSC-Exos effectively promoted chondrocyte proliferation and migration but inhibited their secretion of extracellular matrix (ECM). We planned to establish the reasons and improve the function of SMSCs-Exos.
Whilst studying the reasons for the aforementioned phenomenon, our focus turned to MSC-derived exosome-mediated-Wnt signalling [25]. Exosomes were recently regarded as a potential carrier for secretion and extracellular transportation of Wnt[26, 27]. We measured the gene expression levels of known Wnt proteins, and found that Wnt5a and Wnt5b were highly expressed in SMSCs while other Wnt proteins were rarely expressed. Furthermore, Wnt5a and Wnt5b were enriched in SMSC-Exos.
In the Hippo-YAP signalling pathway, YAP (Yes-associated protein) is activated by de-phosphorylation and translocated into the nucleus to bind TEA domain family transcription factors and thus stimulate the expression of target genes involved in cell proliferation and migration [28-30]. Wnt5a/b promotes YAP activation via the alternative Wnt signalling pathway [31]. Interestingly, Wnt5a/b-YAP signalling antagonises canonical Wnt/β-catenin signalling [31] and decreases expression of a panel of the major β-catenin/TCF target genes, including SOX9[31], which regulates genes related to cartilage ECM [32] and cartilage formation [33, 34].
Genome-wide and gene analysis identified that miR-140-5p plays an essential role in chondrogenic differentiation of MSCs [35]. Recently, miR-140-5p gained a lot of attention for playing dual roles in both cartilage homeostasis and development [36, 37] by targeting RalA to enhance SOX9 and aggrecan (ACAN) [38] and showing potential for preventing OA [39, 40].
We used miRNA microarray to measure the composition of miRNAs in SMSC-Exos and found a low expression level of miR-140-5p.
We overexpressed miR-140-5p in SMSCs and generated an improved version of the exosomes (SMSC-140-Exos) which were able to promote chondrocyte proliferation and migration with less influence on the secretion of extracellular matrix (ECM). In the OA rat model, SMSC-140-Exos successfully prevented OA.
Methods
This study was carried out in compliance with the Declaration of Helsinki. All clinical samples were collected in our department, with written informed consent provided by each patient. Ethical approval was granted by the ethical committee of the Shanghai Sixth People's Hospital. All animal experiments complied with the ARRIVE guidelines. All experimental protocols were approved by the Animal Research Committee of Shanghai Jiao Tong University School of Medicine.
Isolation and characterization of SMSCs
SMSCs were isolated from synovial membranes. To identify the multiple differentiation potential, the cells were induced to differentiate by switching to osteogenic, adipogenic or chondrogenic differentiation medium. Differentiation was confirmed by analysis of cell surface markers using a Guava® easyCyte™ flow cytometer (Merck-Millipore, Darmstadt, Germany). The details are described in Supplementary Methods.
Lentivirus transfection
The lentivectors miR-140-5p, S127A, shYAP #1, and shYAP #2, were obtained from GenePharma (Shanghai, China). Lentivectors RalA, shRalA #1 and shRalA #2 were obtained from Obio Technology (Shanghai, China). Lentiviral particles of Wnt5a (sc-41112) and Wnt5b (sc-95781) shRNAs were purchased from Santa Cruz Biotechnology (Santa Cruz, CA, USA). Lentivirus transfection was performed following the manufacturer's instructions. The shRNA sequences used in this study were as follows:shYAP #1: 5′-CTGGTCAGAGATACTTCTTAA-3′;shYAP #2: 5′-AAGCTTTGAGTTCTGACATCC-3′;shRalA #1: 5′-CCAAGGGTCAGAATTCTTT-3′;shRalA #2: 5′-GGTCAGAATTCTTTGGCTT-3′;
Inhibition of miR-140-5p
To confirm the function of miR-140-5p, use of an inhibitor and antagomir are reliable approaches [41-43]. The miR-140-5p-inhibitor, miR-140-5p-antagomir and negative control were purchased from Ribobio (Guangzhou, China). Oligonucleotide transfection was performed with riboFECT™ CP Reagents (Ribobio).
Isolation and identification of exosomes
Exosomes were isolated from the conditioned medium of SMSCs. Dynamic light scattering (DLS) analysis and transmission electron microscopy (TEM) were used to identify exosomes. RNA and protein were extracted from exosomes using a Total Exosome RNA & Protein Isolation Kit (Invitrogen, Carlsbad, CA, USA) for further analysis. DiO-labelled exosomes were used to observe exosome uptake by chondrocytes. The details are described in Supplementary Methods.
Isolation of RNA from cells
Total RNA was extracted from cells using TRIzol Reagent (Invitrogen) following the manufacturer's instructions.
Real-time quantitative polymerase chain reaction (qPCR)
For miRNA, synthesis of cDNA and qRT-PCR was performed using a human microRNA qRT-PCR detection kit specifically for hsa-miR-140-5p and RNU6B (BioTNT, Shanghai, China). RNU6B was used for normalization of the results. The sequences are provided in Supplementary Methods.
For the mRNA, synthesis of cDNA was performed using TransScript® All-in-One First-Strand cDNA Synthesis SuperMix for qPCR (Transgen Biotech, Beijing, China). Real-time qPCR was performed using TransStart® Top Green qPCR SuperMix (Transgen Biotech), and β-actin was used for normalization of the results. The primer sequences are provided in Supplementary Methods.
Western blotting
The protocol and procedure for western blotting were as described in our previous reports [30, 44, 45]. The anti-p-YAP (Ser127), anti-YAP, anti-RalA and anti-β-actin primary antibodies were obtained from Cell Signalling Technology (Danvers, MA, USA). The anti-Wnt5a, anti-Wnt5b, anti-Aggrecan and anti-SOX9 primary antibodies were obtained from Abcam (Cambridge, MA, USA). The anti-CD63, anti-CD9, anti-CD81 and anti-Alix were obtained from System Biosciences (Palo Alto, CA, USA). Based on previous research [46, 47], collagen type II was analysed using 5% (wt/vol) SDS-PAGE with anti-collagen type II primary antibodies (Chemicon®, Merck-Millipore).
miRNA microarray
Total RNA was analysed by microarray according to the manufacturer's protocol. Briefly, total RNA (100 ng) was labelled with the microRNA Complete Labelling and Hyb Kit (Agilent, Santa Clara, CA, USA), and then hybridized to the Agilent Human miRNA microarray V21.0 (8×60K) for 2549 human microRNAs. After hybridization, the microarray was washed using a Gene Expression Wash Buffer kit (Agilent). Hybridization signals were then scanned with the Agilent Microarray Scanner (Agilent) using Agilent scan control software Version A7.0 (Agilent). The data collection, background subtraction and within array normalization were performed by Agilent Feature Extraction Software (Agilent). Percentile normalization and principal component analysis were performed by Gene spring 12.0 (Agilent).
Immunofluorescence analysis
For immunofluorescence, after reaching more than 80% confluence, chondrocytes were transferred to serum-free culture medium for 24 h and then incubated in complete culture medium with various exosomes for an additional three hours. The cells were then washed in phosphate-buffered saline (PBS) and fixed with 4% paraformaldehyde for 15 min. The fixed cells were permeabilised with 0.1% Triton X-100 for 5 min and washed with PBS three times. Then, the cells were treated with primary antibodies overnight at 4°C in PBS supplemented with 1% bovine serum albumin (BSA). Cells were washed a further three times, then the secondary Alexa Fluor® 488-conjugated antibody (Cell Signalling Technology) was added and incubated for 1 h at room temperature. After washing using PBS, cells were mounted in ProLong® Diamond Antifade Mountant with DAPI (Thermo Fisher Scientific) and randomly-selected fields were imaged using an LSM-880 confocal fluorescence microscope (Carl Zeiss, Jena, Germany).
In vitro response of chondrocytes to SMSC-Exos and SMSC-140-Exos
Cells and cell culture
Human cartilage obtained from the resected lateral femoral condyle of patients, aged from 45 to 55 years old, undergoing total knee arthroplasty (THA), was minced into small pieces and digested using collagenase type II (Gibco) which was diluted in complete chondrocyte culture medium: DMEM/F12 (Hyclone) supplemented with 10% foetal bovine serum (FBS), penicillin/streptomycin (PS) and 250 ng/mL amphotericin B (Gibco). After overnight digestion, the whole mix was filtered through a 70 μm cell strainer (Falcon, BD Biosciences, Franklin Lakes, NJ, USA) then plated into T25 flasks in 10 mL complete chondrocyte culture medium and cultured for 14 days as Passage 0 (P0). Then the cells were digested using trypsin-EDTA and counted. Half of the P0 articular chondrocytes were preserved at -80°C using CELLSAVING (New Cell & Molecular Biotech, Suzhou, China). The rest were cultured and passaged for subsequent study.
Proliferation of chondrocytes
The effect of stimulation with various exosomes or lentivirus transfection on chondrocytes was measured using an EdU-488 Cell Proliferation Kit (Ribobio) with flow cytometry following the manufacturer's instructions. In brief, normal chondrocytes or chondrocytes transfected with lentiviral vectors or empty vector, at an initial density of 2 × 104 cells/well, were seeded into 48-well plates and cultured with various exosomes for 12 h. Next, EdU working solution, consisting of 150 μL of complete chondrocyte culture medium containing 0.15 μL of EdU, was added into each well and incubated for 3 h at 37°C. Cultures were then digested using trypsin-EDTA, washed using PBS, fixed in 4% paraformaldehyde (PFA) for 15 min, neutralized with 2 mg/mL glycine and washed twice in PBS before permeabilising with 0.4% Triton X-100 for 5 mins and finally washing twice with PBS. The labelled cells were resuspended using the Apollo staining solution in the kit by incubating for 10 min, then washed twice in 0.4% Triton X-100 and resuspended in PBS for analysis using the Guava® easyCyte™ flow cytometer.
Migration of chondrocytes
The effect of stimulation of chondrocytes with various exosomes or lentivirus transfection was evaluated using a transwell assay. In brief, after digestion and counting, approximately 5 × 104 cells were seeded into the upper chamber of a 24-well 8-μm-pore-size transwell plate (Corning, Corning, NY, USA). Next, 600 μL of complete chondrocyte culture medium containing various exosomes was added into the lower chamber of the transwell plate before incubation for 12 h at 37°C. The upper chamber was then fixed with 4% PFA for 15 min, stained with 0.5% crystal violet for 10 min and washed with PBS three times. The upper surface of the upper chamber was carefully wiped using a cotton swab to remove cells which had not migrated to the lower surface. Five randomly-selected fields (100 × magnification) per well were photographed using a Leica microscope and assessed by two pathologists in a blinded manner.
Animal studies
Male Sprague-Dawley rats (approximately 12 weeks old) weighing 300-350 g, housed in a specific pathogen-free (SPF) animal laboratory with 12:12 hours light/dark cycle, controlled temperature environment (23-25°C) and steady humidity (55-70%), were used in these studies. Based on former studies [48], the OA model was established by transecting the medial collateral ligament and the medial meniscus completely, by cutting the meniscus at the narrowest point without damaging the tibial surface, and transecting the anterior cruciate ligament. After surgery, all rats received 0.05 mg/kg buprenorphine for pain relief together with 5 mg/kg gentamicin. All Sprague-Dawley rats were randomly divided into four groups: (1) Normal group (without surgery; received articular cavity injection of normal saline every time the OA group was given an injection; 10 knee joints from 5 rats, n = 10) (2) OA group (rats underwent surgery and received articular cavity injection of normal saline on the first day of every week from the 5th to the 8th week after surgery; 10 knee joints from 5 rats, n = 10); (3) OA+SMSC-Exos group (rats received articular cavity injection of SMSC-Exos (100 μL; 1011 exosome particles/mL; 10 knee joints from 5 rats, n = 10) when the OA group received saline injections); (4) OA+SMSC-140-Exos group (Rats received articular cavity injection of SMSC-140-Exos (100 μL; 1011 exosomes particles/mL) when the OA group received saline injection; 10 knee joints from 5 rats, n = 10).
Twelve weeks after surgery, rats were sacrificed by anaesthetic overdose and the knee samples were harvested to evaluate disease progression.
Histology and immunohistochemical analysis
The tibiofemoral joints were removed and the femoral condyles were fixed in neutral-buffered formalin (containing 4% formaldehyde) for 24 h after the knee samples were harvested. The fixed femoral condyles were decalcified in EDTA for 21 days, refreshed every day, before graded ethanol dehydration, dimethylbenzene vitrification, paraffin embedding and tissue sectioning (5 μm).
Safranin-O & fast green-stained sections were used to evaluate matrix proteoglycan and overall joint morphology [49, 50].
Immunohistochemistry was performed to evaluate the function of articular chondrocytes. Anti-type II collagen, anti-type I collagen and anti-aggrecan antibodies were used as primary antibodies (all from Abcam, 1:100).
Pictures at high magnification were randomly captured and evaluated by two different pathologists.
Based on safranin O-fast green staining, OARSI score [51], which is a well-recognized histological scoring system [52], was used by two different blinded pathologists to evaluate the OA progress of every sample in each group.
Statistical analysis
Numerical data are presented as mean ± standard deviation (SD). All statistical analyses in this study were performed using SPSS 16.0 software (IBM Corp., Armonk, NY, USA). The significance of differences between two groups was analysed using Student's t test. A P value < 0.05 was considered significant.
Results
Isolation and identification of SMSCs
SMSCs were isolated from synovial membrane as described in the Methods section. At passage 5 (P5), SMSCs were identified and used in subsequent experiments. After reaching 80-90% confluence, cells adopted a spindle-like shape (Figure 1A).
Classification of human synovial mesenchymal stem cells (SMSCs) and exosomes. (A) SMSCs exhibited a representative spindle-like morphology (scale bar: 50 μm). (B) SMSCs showed multi-potential differentiation capacity for osteogenesis (scale bar: 50 μm), adipogenesis (scale bar: 20 μm) and chondrogenesis (scale bar: 50 μm). (C) Flow cytometric analysis of characteristic cell surface markers of SMSCs. Unfilled curves represent isotype controls and solid grey curves represent measured surface markers (CD34, CD44, CD45, CD73, CD90, CD105, CD133 and CD151). This experiment was repeated independently three times. (D) Particle size distribution of exosomes measured by dynamic light scattering (DLS). This experiment was repeated independently three times and representative results are shown. (E) Morphology of exosomes observed by transmission electron microscopy (TEM). Scale bar: 100 nm. (F) Exosome surface markers (Alix, CD81, CD9, and CD63) and carried proteins (Wnt5a and Wnt5b) measured using western blotting. This experiment was repeated independently three times and representative results are shown. (G) Representative immunofluorescence photomicrograph of DiO (green)-labelled exosomes absorbed by chondrocytes, the nuclei of which were stained by DAPI (blue). Scale bar: 50 μm.
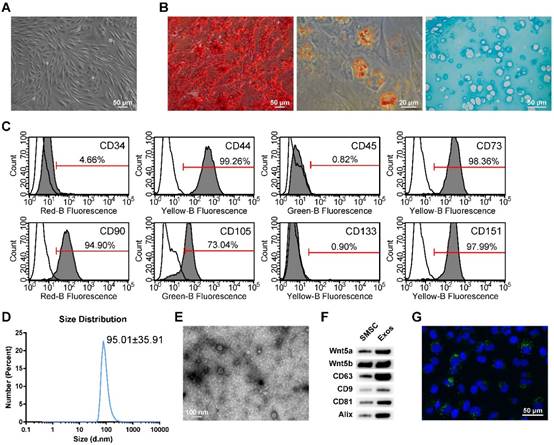
When cultured in osteogenic, adipogenic or chondrogenic medium, the SMSCs could readily be induced to differentiate along the osteogenic, adiopogenic and chondrogenic pathways, respectively. Osteogenic potential was confirmed by staining of calcium mineral deposits with Alizarin Red (Figure 1B, left panel), while adipogenic potential was evaluated by observation of small cytoplasmic lipid droplets stained using Oil Red O (Figure 1B, middle panel). Chondrogenic potential was confirmed by sectioning beads and staining sulphated glycosaminoglycans using Alcian blue (Figure 1B, right panels). The representative morphology of chondrocytes, typically round and plump, was observed.
Following the criteria for identifying stem cells, we analysed the surface markers of SMSCs. Analysis of surface antigen expression using flow cytometry demonstrated that SMSCs were positive for CD44, CD73, CD90, CD105 and CD151, and negative for CD34, CD45, and CD133 (Figure 1C).
Isolation and identification of exosomes derived from SMSCs
Dynamic light scattering (DLS), transmission electron microscopy (TEM) and western blotting were used to characterize the particles secreted from SMSCs. The results showed that most of these vesicles ranged from 30 to 150 nm in size (Figure 1D). These hollow spherical microvesicles (Figure 1E) were further confirmed by western blotting. The expression of the exosome markers CD63, CD9, CD81 and Alix (Figure 1F) were significantly enriched in exosomes. All these data indicate that exosomes were successfully isolated.
Internalization of exosomes by articular chondrocytes
To confirm whether the chondrocytes could take up exosomes derived from SMSCs, SMSCs were labelled using green fluorescent lipophilic dye (Vybrant DiO) before harvesting the exosomes. Chondrocytes were then incubated for 6 h with exosomes derived from labelled cells. After washing with PBS, the DiO-labelled exosomes were observed in the perinuclear region of the chondrocytes (Figure 1G), confirming the internalization by chondrocytes.
SMSC-Exos activated YAP, decreased ECM secretion of articular chondrocytes, and induced proliferation and migration of articular chondrocytes
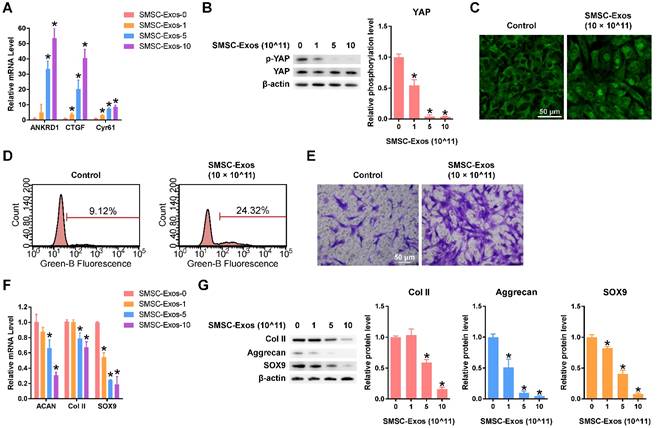
After quantifying the exosomes using an ExoELISA kit, we stimulated chondrocytes using 0, 1, 5, or 10 × 1011 particles/mL of SMSC-Exos for 24 h. Then, the levels of characterized downstream targets of YAP signalling, including ANKRD1, CTGF and Cyr61[53], were analysed using qPCR. The data showed that levels of these YAP downstream genes were enhanced in a dose-dependent manner (Figure 2A). To confirm the activation of YAP, western blotting and fluorescence confocal assays were performed. The results revealed that YAP was activated by SMSC-Exos via de-phosphorylation (Figure 2B). The activated YAP was localized in the nucleus (Figure 2C) and activated the downstream genes to be translated.
Flow cytometry with an EdU kit showed that the proliferative ability of chondrocytes was dramatically enhanced by SMSC-Exos (Figure 2D). We also found that the migratory ability of chondrocytes was obviously strengthened by SMSC-Exos (Figure 2E).
However, the function of chondrocytes including ECM secretion was obviously suppressed by SMSC-Exos (Figure 2F-G), as analysed using qPCR and western blotting.
SMSC-Exos activated YAP, decreased ECM secretion, and induced proliferation and migration of articular chondrocytes via Wnt5a and Wnt5b
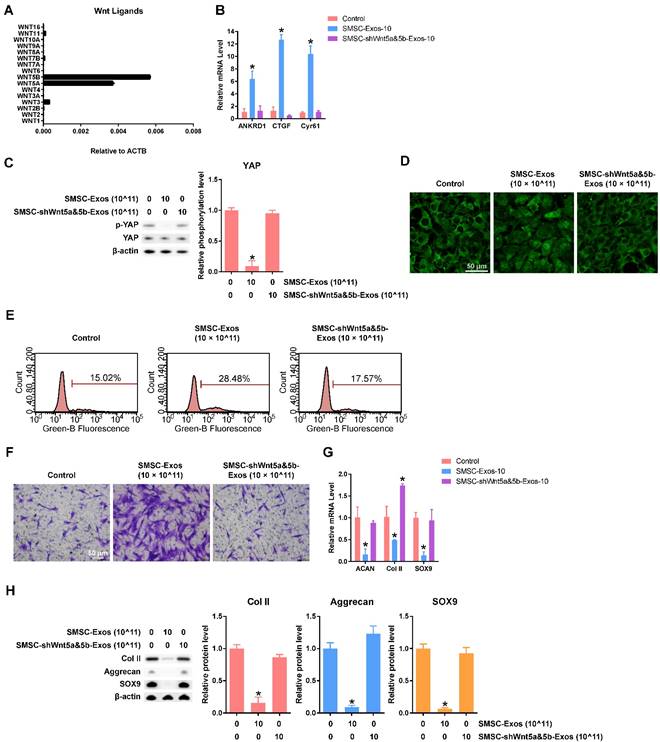
To elucidate the mechanism behind these phenomena in chondrocytes stimulated by SMSC-Exos, we analysed the gene expression levels of Wnt family members in SMSCs and found that Wnt5a and Wnt5b were highly expressed (Figure 3A). We also confirmed that the protein levels of Wnt5a and Wnt5b were highly expressed in both SMSCs and exosomes derived from them using western blotting (Figure 1F).
After suppressing the expression level of both Wnt5a and Wnt 5b using lentiviral shRNA, we found no evidence of YAP activation (Figure 3B-D), nor of enhanced proliferation and migration abilities (Figure 3E-F) or inhibition of chondrocyte functions including ECM secretion (Figure 3G-H).
All these data indicated that Wnt5a and Wnt5b were the major effector molecules of SMSC-Exos.
SMSC-Exos decreased ECM secretion and induced proliferation and migration of articular chondrocytes through activation of YAP
Next, we aimed to clarify whether YAP activation via the alternative Wnt signalling pathway was responsible for the effects caused by Wnt5a and Wnt5b.
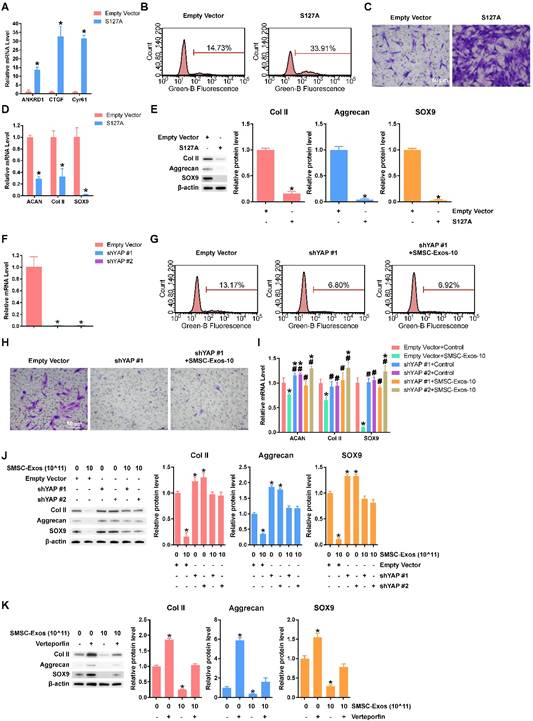
We overexpressed S127A, a mutant of YAP protein that generates a constitutively active YAP [30], in chondrocytes. The efficiency of S127A transfection was analysed using qPCR (Figure 4A). We found that YAP activation had the same effects as SMSC-Exos stimulation on cell proliferation (Figure 4B), cell migration (Figure 4C) and chondrocyte functions including ECM secretion (Figure 4D-E).
To further verify that YAP is the key molecule, two YAP shRNAs (shYAP #1 and shYAP #2) were designed and their inhibitory efficiency was verified using qPCR (Figure 4F). The proliferation and migration abilities were weak in chondrocytes transfected with shYAP and could not be enhanced by SMSC-Exos (Figure 4G-H). Further, the effects on chondrocyte function including ECM secretion were blocked by down-regulation of YAP (Figure 4I-J).
To verify the role of the YAP-TEAD complex in the effects on chondrocyte function including ECM secretion, we used verteporfin, a chemical which can disrupt YAP-TEAD interaction [54], and further confirmed that the suppression of ECM secretion caused by SMSC-Exos was rescued by verteporfin (Figure 4K).
miR-140-5p maintained the function of articular chondrocytes
In addition to proteins, miRNAs are another type of key molecule in exosomes [24]. We used miRNA microarray to measure expression levels of various miRNAs (Figure 5A). The expression levels of highly-expressed miRNAs and miR-140-5p/miR-140-3p which had attracted our interest are shown in Figure 5B. To date there is no clear link between the highly-expressed miRNA in exosomes and chondrocyte function. The miR-140-5p which is closely related to chondrocyte function is rarely expressed in SMSC-Exos.
Responses of articular chondrocytes stimulated by SMSC-Exos. (A) Changes in the expression of the representative downstream genes ANKRD1, CTGF and Cyr61 following YAP activation. SMSC-Exos-0, 1, 5 and 10 indicates that 0, 1, 5, or 10 × 1011 exosome particles/mL were used in the corresponding groups. This experiment was repeated three times. *P < 0.05 compared to SMSC-Exos-0. (B) Representative pictures of de-phosphorylation of YAP, reflecting its activation, and the results of statistical analysis of three replicates. *P < 0.05 compared to SMSC-Exos-0. (C) Subcellular localization of YAP detected using confocal immunofluorescence microscopy. This experiment was repeated independently three times and representative results are shown. Scale bar: 50 μm. (D) EdU assays performed using flow cytometry. The percentage of EdU-positive cells (labelled green) was determined. This experiment was repeated independently three times and representative results are shown. (E) Chondrocyte migration assays. Migrated chondrocytes were stained by crystal violet. This experiment was repeated independently three times and representative results are shown. Scale bar: 50 μm. (F) Gene expression changes of Aggrecan (ACAN), Type II Collagen (Col II) and SOX9 after stimulation with different concentrations of SMSC-Exos. This experiment was repeated three times. *P < 0.05 compared to SMSC-Exos-0. (G) Protein expression levels of Col II, Aggrecan and SOX9 were detected by western blotting and the results of statistical analysis of three replicates are also shown. *P < 0.05 compared to SMSC-Exos-0.
Functions of Wnt5a and Wnt5b in SMSC-Exos-stimulated articular chondrocytes. (A) Normalized gene expression levels of Wnt family members in SMSCs. This experiment used three replicates. (B) Expression of YAP downstream genes in chondrocytes treated with exosomes, at a concentration of 10 × 1011 exosome particles/mL, derived from SMSCs transfected by empty vector (SMSC-Exos-10) and SMSCs transfected with shWnt5a and shWnt5b (SMSC-shWnt5a&5b-Exos). This experiment was repeated three times. (C) YAP activation was detected by western blotting by observing the de-phosphorylation of YAP. The results of statistical analysis of three independent replicates are also shown. *P < 0.05 compared to control. (D) Subcellular localization of YAP was observed using confocal immunofluorescence microscopy. This experiment was repeated independently three times and representative results are shown. Scale bar: 50 μm. (E) EdU assays were performed using flow cytometry. The percentage of EdU-positive cells (labelled green) was determined. This experiment was repeated independently three times and representative results are shown. (F) Migration assays of chondrocytes. Migrated chondrocytes were stained by crystal violet. This experiment was repeated independently three times and representative results are shown. Scale bar: 50 μm. (G) Gene expression changes of Aggrecan, Col II and SOX9 after stimulation with the indicated exosomes, at a concentration of 10 × 1011 exosome particles/mL. This experiment was repeated three times. *P < 0.05 compared to control. (H) Protein expression levels of Col II, Aggrecan and SOX9 were detected using western blotting. The results of statistical analysis of three independent replicates are also shown. *P < 0.05 compared to control.
YAP plays an important role in mediating the effects of SMSC-Exos stimulation. (A) Representative gene expression levels of YAP downstream genes after chondrocytes were transfected with empty vector or S127A. This experiment was repeated three times. *P < 0.05 compared to empty vector. (B) EdU assays were performed, using chondrocytes transfected with empty vector or S127A. The percentage of EdU-positive cells (labelled green) was determined. This experiment was repeated three times. (C) Migration of chondrocytes after transfection with empty vector or S127A. Migrated chondrocytes were stained with crystal violet. This experiment was repeated three times. Scale bar: 50 μm. (D) Gene expression levels of Aggrecan, Col II and SOX9 after chondrocytes were transfected with empty vector or S127A. This experiment was repeated three times. *P < 0.05 compared to empty vector. (E) Protein levels of Aggrecan, Col II and SOX9 were analysed using western blotting. The results of statistical analysis of three independent replicates are also shown. *P < 0.05 compared to Empty Vector. (F) YAP gene expression levels after chondrocytes were transfected by shYAP #1 or shYAP #2. This experiment was repeated three times. *P < 0.05 compared to empty vector. (G) EdU assays were performed on chondrocytes transfected with empty vector or shYAP #1 with or without SMSC-Exos treatment, at a concentration of 10 × 1011 exosome particles/mL. This experiment was repeated three times. (H) Chondrocyte migration assays. Migrated chondrocytes were stained by crystal violet. This experiment was repeated three times. Scale bar: 50 μm. (I) Chondrocytes, transfected with empty vector, shYAP #1 or shYAP #2, were treated with or without SMSC-Exos at a concentration of 10 × 1011 exosome particles/mL. This experiment was repeated three times. *P < 0.05 compared to chondrocytes transfected with empty vector without exosome treatment (Empty Vector+Control). #P < 0.05 compared to chondrocytes transfected with empty vector with SMSC-Exos treatment, at a concentration of 10 × 1011 exosome particles/mL (Empty Vector+SMSC-Exos-10). (J-K) Protein expression levels of Col II, Aggrecan and SOX9 were analysed using western blotting. The results of statistical analysis of three independent replicates are also shown. *P < 0.05 compared to control.
miR-140-5p maintains the function of articular chondrocytes (A) Normalized gene expression levels in SMSC-Exos detected by miRNA microarray. Three independent samples were used. (B) Gene expression levels of highly-expressed genes in SMSC-Exos and gene expression levels of miR-140-5p and miR-140-3p. (C) Efficiency of shRalA #1, shRalA #2 and RalA lentiviral vectors were detected by qPCR. This experiment was repeated three times. *P < 0.05 compared to chondrocytes transfected with empty vector. (D) Aggrecan or SOX9 gene expression levels of chondrocytes transfected with miR-140-5p, RalA or shRalA #1. This experiment was repeated three times. *P < 0.05 compared to chondrocytes transfected with empty vector. (E) Protein expression levels of Col II, Aggrecan, SOX9 and RalA were analysed using western blotting. The results of statistical analysis of three independent replicates are also shown. *P < 0.05 compared to Empty Vector. #P < 0.05 compared between each other. (F) Expression level changes of miR-140-5p in SMSC-Exos, SMSC-140-Exos, or in articular chondrocytes which were treated with SMSC-Exos (AC-SMSC-Exos) or SMSC-140-Exos (AC-SMSC-140-Exos). This experiment was repeated three times.
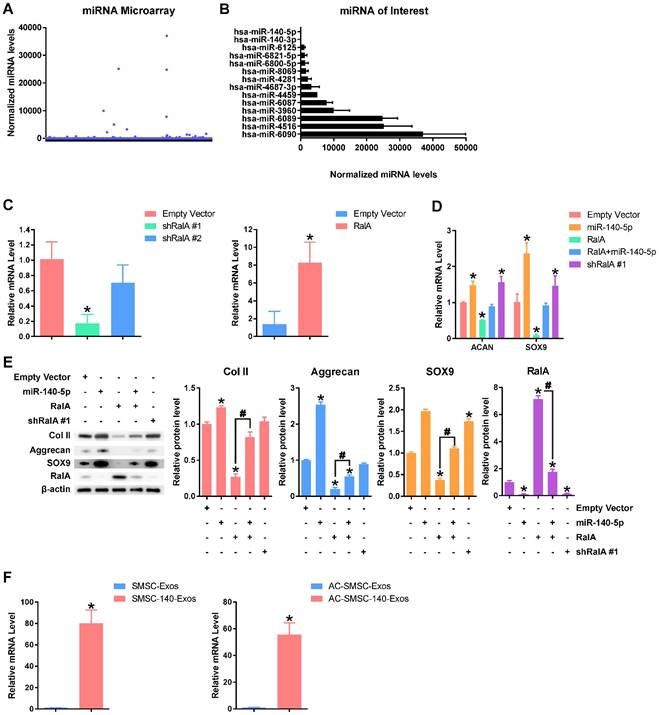
We modified SMSC-Exos by miR-140-5p overexpression after verifying the function of miR-140-5p. Existing research [38] indicated that miR-140-5p may target RalA to translationally enhance SOX9 and Aggrecan. We designed two shRNAs for RalA (shRalA #1 and shRalA #2) and tested the inhibitory efficiency using qPCR (Figure 5C). The chondrocytes transfected with shRalA #1 were then used for subsequent experiments. The efficiency of RalA overexpression was also tested by qPCR (Figure 5C).
The experimental results (Figure 5D-E) show that miR-140-5p suppressed RalA and upregulated SOX9, ACAN and collagen type II and that this effect could be blocked by RalA overexpression. Further, shRalA #1 transfection had similar effects to miR-140-5p overexpression.
Exosomes derived from miR-140-5p-overexpressing SMSCs (SMSC-140-Exos) induced proliferation and migration of AC without decreasing ECM secretion
We overexpressed miR-140-5p in SMSCs and obtained the exosomes derived from miR-140-5p-overexpressing SMSCs (SMSC-140-Exos). The expression level of miR-140-5p was obviously upregulated in SMSC-140-Exos compared to SMSC-Exos. In addition, the expression level of miR-140-5p of chondrocytes treated with SMSC-140-Exos was higher compared to chondrocytes treated with SMSC-Exos (Figure 5F) when analysed by qPCR.
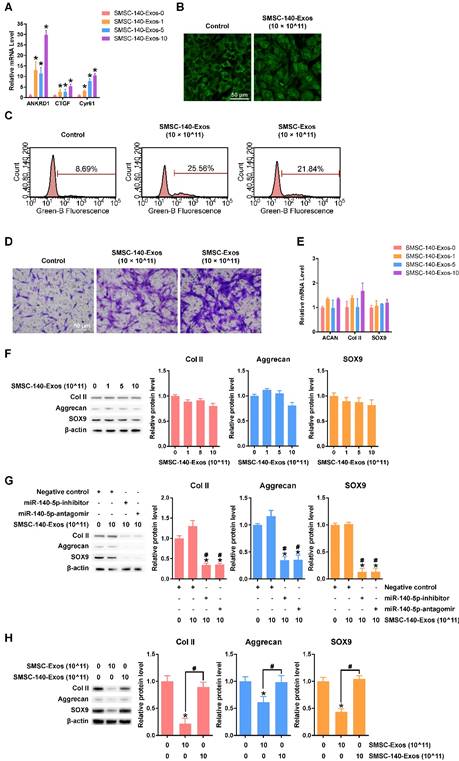
SMSC-140-Exos had similar effects on YAP activation, proliferation and migration of chondrocytes (Figure 6A-D) without obvious suppression of ECM secretion (Figure 6E-F and H) compared to SMSC-Exos.
To verify the function of miR-140-5p in SMSC-140-Exos, we used two methods, miR-140-5p-inhibitor and miR-140-5p-antagomir, to block the miR-140-5p which was transferred into chondrocytes. The results indicated that the functions of chondrocytes, including ECM secretion, were maintained by miR-140-5p (Figure 6 G).
SMSC-140-Exos prevented OA
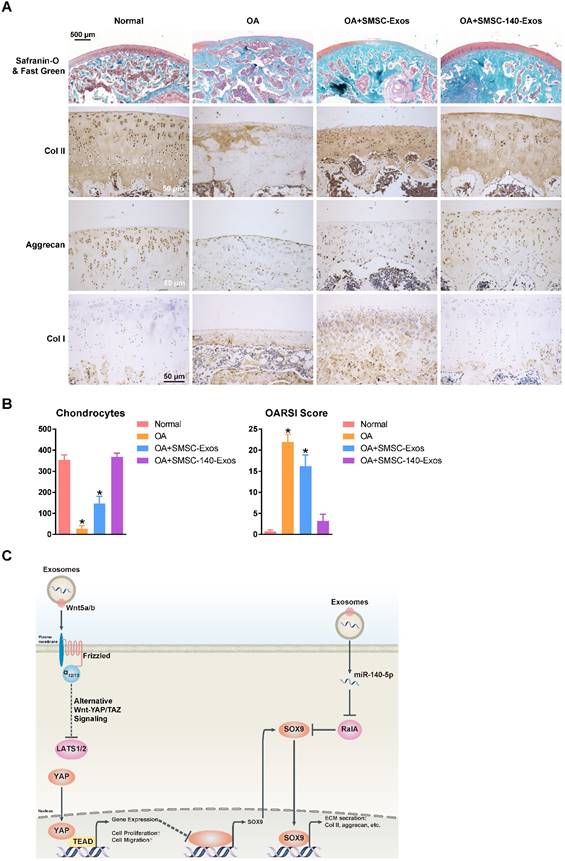
We verified the potential of exosomes for OA prevention in an OA rat model (Figure 7A). We also found that no obvious adverse events occurred in any experimental group.
In the OA group, severe joint wear and cartilage matrix loss were observed. The expression of type II collagen and aggrecan in cartilage had decreased, while expression of type I collagen was observed on the cartilage surface.
In the OA+SMSC-Exos group, joint wear and cartilage matrix loss were also observed. However, these were much less severe compared to the OA group. The cartilage matrix consisting of type II collagen was thin and cartilage cells were arranged in dense clusters of chondrocytes rather than the normal arrangement of neat rows. Expression of aggrecan was low while type I collagen was obviously expressed in the cartilage matrix.
In the OA+SMSC-140-Exos group, joint wear was still present but the severity was very mild. The cartilage matrix consisting of type II collagen was slightly thinner than in the normal group but significantly better than in the OA group or the OA+SMSC-Exos group. There was no obvious decrease of aggrecan expression, and no type I collagen expression was observed in the cartilage matrix.
These data indicated that SMSC-140-Exos slowed the progression of early OA and prevented the severe damage to knee articular cartilage in the OA model caused by instability of the knee joint.
The chondrocyte counts and OARSI scores are shown in Figure 7B.
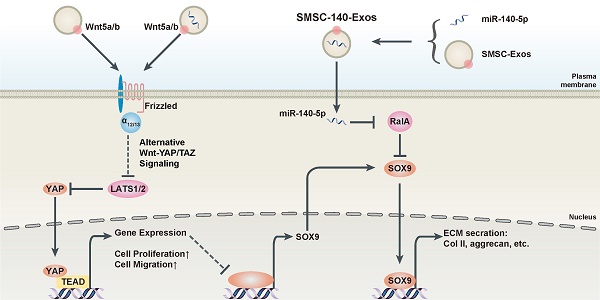
Summary of the therapeutic mechanism
In Figure 7C, we summarize the therapeutic mechanism of SMSC-140-Exos. Wnt5a and Wnt5b activated YAP via the alternative Wnt-YAP/TAZ signalling pathway. Activation of YAP led to the enhancement of chondrocyte proliferation and migration with the side effect of decreasing SOX9 expression and ECM secretion. With the help of miR-140-5p highly expressed in SMSC-140-Exos, SOX9 was rescued via RalA inhibition and ECM secretion was successfully restored.
Discussion
The limited potential of articular cartilage for self-regeneration has been attributed to the fact that it is difficult for cells to access the injured site due to the inability of adjacent articular chondrocytes to migrate and produce matrix [55]. In our study, SMSC-Exos promoted chondrocyte proliferation and migration, which are inhibited in OA [56]. The drawback of SMSC-Exos is their inhibition of SOX9 and ECM generation, which was overcome by overexpressing miR-140-5p in SMSCs and using the SMSC-140-Exos.
SMSCs can easily be isolated from synovial membrane, which is easy to obtain. SMSCs were observed to proliferate vigorously and age slowly in our study. We found that SMSCs can retain their multi-potential differentiation ability even after reaching passage 10. We therefore believe that SMSCs will be a popular new option among the family of mesenchymal stem cells for tissue regeneration. Their specialties of “tissue specific” regeneration [16] give SMSCs unlimited potential in cartilage regeneration.
Responses of articular chondrocytes to stimulation by SMSC-140-Exos. (A) Representative downstream gene expression changes following YAP activation. SMSC-140-Exos-0, 1, 5 and 10 indicates that 0, 1, 5, or 10 × 1011 exosome particles/mL were used in the corresponding groups. This experiment was repeated three times. *P < 0.05 compared to SMSC-140-Exos-0. (B) Subcellular localization of YAP detected using confocal immunofluorescence microscopy. This experiment was repeated three times. Scale bar: 50 μm. (C) EdU assays performed using flow cytometry. The percentage of EdU-positive cells (labelled green) was determined. This experiment was repeated three times. (D) Chondrocyte migration assays. Migrated chondrocytes were stained by crystal violet. This experiment was repeated three times. Scale bar: 50 μm. (E) Gene expression levels of Aggrecan, Col II and SOX9 after stimulation with SMSC-140-Exos at different concentrations. This experiment was repeated three times. *P < 0.05 compared to SMSC-140-Exos-0. (F) Protein expression levels of Col II, Aggrecan and SOX9 were analysed using western blotting. The results of statistical analysis of three independent replicates are also shown. *P < 0.05 compared to SMSC-140-Exos-0. (G) Protein expression levels of Col II, Aggrecan and SOX9 in the articular chondrocytes transfected with negative control, miR-140-5p-inhibitor or miR-140-5p-antagomir, and treated with or without SMSC-140-Exos at a concentration of 10 × 1011 exosome particles/mL, analysed using western blotting. The results of statistical analysis of three independent replicates are also shown. *P < 0.05 compared to negative control only. #P < 0.05 compared to negative control treated with 10 × 1011 SMSC-140-Exos. (H) Protein expression levels of Col II, Aggrecan and SOX9 were analysed using western blotting, and the results of statistical analysis of three independent replicates are shown. *P < 0.05 compared to control. #P < 0.05 compared between each other.
SMSC-140-Exos prevent OA (A) Sections of femoral condyle (n = 10 for each group) were stained using Safranin-O & fast green (Scale bar: 500 μm). Photomicrographs of femoral condyle sections (n = 10 for each group) stained using anti-type II collagen, anti-aggrecan or anti-type I collagen as primary antibodies (Scale bar: 50 μm). (B) Statistical results of chondrocytes counted in randomly-selected high magnification fields and the result of statistical analysis of OARSI score in each group. *P < 0.05 compared to Normal. (C) Diagram illustrating the proposed mechanism of action of SMSC-140-Exos in OA.
Recent studies have shown that paracrine mechanisms including exosomes are responsible for stem cell- or progenitor cell-mediated tissue regeneration [19, 20]. We found that exosomes derived from SMSCs (SMSC-Exos) clearly promoted chondrocyte proliferation and migration. However, they had one crucial shortcoming: the inhibition of ECM protein synthesis including aggrecan and collagen II. To establish the reason for this, we measured the mRNA expression levels of Wnt family members in SMSCs and found that Wnt5a and Wnt5b were highly expressed in SMSCs. Furthermore, we found that Wnt5a and Wnt5b promoted chondrocyte proliferation and migration by activating YAP through alternative Wnt signalling pathways. The activation of YAP clearly suppressed the expression of SOX9 and led to the suppression of ECM formation.
Wnt5a and Wnt5b play important roles in early chondrogenesis[57] and are able to inhibit chondrocyte hypertrophy via nuclear factor κB (NFκB) and JNK, respectively[58, 59]. These effects combined with enhancement of proliferation and migration may be the reason why SMSC-Exos have some protective effect on OA.
The precise function of YAP in chondrocyte differentiation is unclear. Previous studies have shown that while early chondrocyte proliferation is promoted, maturation is inhibited [60]. This phenomenon is similar to the SMSC-Exos-induced activation of YAP in our study. The inhibition of chondrocyte maturation is the drawback of the use of SMSC-Exos, and consequently the therapeutic strategy needed to be optimized.
In our study, the inhibition of SOX9 and its downstream genes, including aggrecan and type II collagen, was key to the YAP-induced inhibition of maturation. The optimized therapeutic strategy is based on SOX9.
For further analysis of the composition of SMSC-Exos, we performed a miRNA microarray assay. However, the relationship between highly-expressed miRNA in SMSC-Exos and ECM formation was not clear. The function of these microRNAs needs further research. The essential microRNA in cartilage homeostasis and development, miR-140-5p, and its sister, miR-140-3p, were rarely expressed in SMSC-Exos. In view of its importance, we came up with the idea of increasing the miR-140-5p levels in SMSC-Exos.
Nucleic acids can be positively incorporated into exosomes by relying on elevated intracellular RNA concentration, which can be obtained by overexpressing nucleic acids using lentiviral-based or lipid-based systems [61]. We overexpressed miR-140-5p in SMSCs and found that miR-140-5p was enriched in its derived exosomes (SMSC-140-Exos). The miR-140-5p-overexpressed SMSCs were found to be a superior cell line for preventing OA.
SMSC-140-Exos retained the advantages of SMSC-Exos without the disadvantages by rescuing SOX9 through RalA. The ability of chondrocytes to proliferate and migrate when stimulated by SMSC-140-Exos was clearly promoted without significant suppression of ECM protein synthesis.
The highly-expressed miRNAs in exosomes are shown in Figure 5B. Key miRNAs are miR-6090, which may play a role in embryo development [62]; miR-4516, which is involved in autophagy[63]; miR-3960, which is related to Runx2[64], and miR-4459, which is involved in altering the stemness of embryonic stem cells[65]. However, the relationship between these microRNAs and chondrocytes remains unknown and needs to be further researched.
We chose an OA rat model based on knee joint instability induced by surgery. This kind of model can perfectly simulate knee joint instability caused by knee injury, including anterior cruciate ligament (ACL) injury. It can also reasonably simulate joint wear caused by anatomical factors.
In vivo, we used SMSC-140-Exos in an OA rat model. The progression of early stage OA was delayed and the knee joint cartilage damage, which was caused by OA, was prevented by SMSC-140-Exos while the effect of SMSC-Exos was limited.
We conclude that SMSCs and their exosomes with or without some kind of modification still show tremendous potential for future exploration and use in clinical practice.
Supplementary Material
Supplementary methods.
Abbreviations
OA: Osteoarthritis
SMSC-140s: miR-140-5p-overexpressing synovial mesenchymal stem cells
SMSC-140-Exos: Exosomes derived from SMSC-140s
ACs: Articular chondrocytes
ECM: Extracellular matrix
YAP: Yes-associated protein
MSCs: Mesenchymal stem cells
SMSCs: Synovial mesenchymal stem cells
ACAN: Aggrecan
DLS: Dynamic light scattering
TEM: Transmission electron microscopy
qPCR: Real-time quantitative polymerase chain reaction
BSA: Bovine serum albumin
THA: Total knee arthroplasty
P0: Passage 0
PFA: Paraformaldehyde
PBS: Phosphate Buffered Saline
SD: Standard deviation
P5: Passage 5
shRNA: Small hairpin RNA
NFκB: Nuclear factor κB
ACL: Anterior cruciate ligament
Acknowledgements
This study was supported by the National Natural Science Foundation of China (Grant Number. 81301589, 81572239 and 81472066). We thank OEbiotech Company for technical support with the microarray studies.
Author Contributions
S.C.T. and S.C.G. carried out the main part of the studies and drafted the manuscript. Y.L.Z and W.J.Y. participated in the in vivo experiments. T.Y. participated in the design of the study. S.C.T. performed the statistical analysis. S.C.G. and C.Q.Z. conceived the study, participated in its design and coordination, and helped to draft the manuscript. All authors read and approved the final manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Woolf AD, Pfleger B. Burden of major musculoskeletal conditions. Bulletin of the World Health Organization. 2003;81:646-56
2. Skou ST, Roos EM, Laursen MB, Rathleff MS, Arendt-Nielsen L, Simonsen O. et al. A Randomized, Controlled Trial of Total Knee Replacement. The New England journal of medicine. 2015;373:1597-606
3. Glyn-Jones S, Palmer AJ, Agricola R, Price AJ, Vincent TL, Weinans H. et al. Osteoarthritis. Lancet (London, England). 2015;386:376-87
4. Agricola R, Heijboer MP, Roze RH, Reijman M, Bierma-Zeinstra SM, Verhaar JA. et al. Pincer deformity does not lead to osteoarthritis of the hip whereas acetabular dysplasia does: acetabular coverage and development of osteoarthritis in a nationwide prospective cohort study (CHECK). Osteoarthritis and cartilage / OARS, Osteoarthritis Research Society. 2013;21:1514-21
5. Dezateux C, Rosendahl K. Developmental dysplasia of the hip. Lancet (London, England). 2007;369:1541-52
6. Agricola R, Heijboer MP, Bierma-Zeinstra SM, Verhaar JA, Weinans H, Waarsing JH. Cam impingement causes osteoarthritis of the hip: a nationwide prospective cohort study (CHECK). Annals of the rheumatic diseases. 2013;72:918-23
7. Zadpoor AA. Etiology of Femoroacetabular Impingement in Athletes: A Review of Recent Findings. Sports medicine (Auckland, NZ). 2015;45:1097-106
8. Sharma L, Chmiel JS, Almagor O, Felson D, Guermazi A, Roemer F. et al. The role of varus and valgus alignment in the initial development of knee cartilage damage by MRI: the MOST study. Annals of the rheumatic diseases. 2013;72:235-40
9. Felson DT, Niu J, Gross KD, Englund M, Sharma L, Cooke TD. et al. Valgus malalignment is a risk factor for lateral knee osteoarthritis incidence and progression: findings from the Multicenter Osteoarthritis Study and the Osteoarthritis Initiative. Arthritis and rheumatism. 2013;65:355-62
10. Harvey WF, Yang M, Cooke TD, Segal NA, Lane N, Lewis CE. et al. Association of leg-length inequality with knee osteoarthritis: a cohort study. Annals of internal medicine. 2010;152:287-95
11. Lawrence RC, Felson DT, Helmick CG, Arnold LM, Choi H, Deyo RA. et al. Estimates of the prevalence of arthritis and other rheumatic conditions in the United States. Part II. Arthritis and rheumatism. 2008;58:26-35
12. Muthuri SG, McWilliams DF, Doherty M, Zhang W. History of knee injuries and knee osteoarthritis: a meta-analysis of observational studies. Osteoarthritis and cartilage / OARS, Osteoarthritis Research Society. 2011;19:1286-93
13. Jiang Y, Jahagirdar BN, Reinhardt RL, Schwartz RE, Keene CD, Ortiz-Gonzalez XR. et al. Pluripotency of mesenchymal stem cells derived from adult marrow. Nature. 2002;418:41-9
14. Kurth TB, Dell'accio F, Crouch V, Augello A, Sharpe PT, De Bari C. Functional mesenchymal stem cell niches in adult mouse knee joint synovium in vivo. Arthritis and rheumatism. 2011;63:1289-300
15. De Bari C, Dell'Accio F, Tylzanowski P, Luyten FP. Multipotent mesenchymal stem cells from adult human synovial membrane. Arthritis and rheumatism. 2001;44:1928-42
16. Jones BA, Pei M. Synovium-derived stem cells: a tissue-specific stem cell for cartilage engineering and regeneration. Tissue engineering Part B, Reviews. 2012;18:301-11
17. Amariglio N, Hirshberg A, Scheithauer BW, Cohen Y, Loewenthal R, Trakhtenbrot L. et al. Donor-derived brain tumor following neural stem cell transplantation in an ataxia telangiectasia patient. PLoS medicine. 2009;6:e1000029
18. Herberts CA, Kwa MS, Hermsen HP. Risk factors in the development of stem cell therapy. Journal of translational medicine. 2011;9:29
19. Kim JY, Song SH, Kim KL, Ko JJ, Im JE, Yie SW. et al. Human cord blood-derived endothelial progenitor cells and their conditioned media exhibit therapeutic equivalence for diabetic wound healing. Cell transplantation. 2010;19:1635-44
20. Zhang M, Malik AB, Rehman J. Endothelial progenitor cells and vascular repair. Current opinion in hematology. 2014;21:224-8
21. Zhou J, Wang S, Sun K, Chng WJ. The emerging roles of exosomes in leukemogeneis. Oncotarget. 2016
22. Thery C, Ostrowski M, Segura E. Membrane vesicles as conveyors of immune responses. Nature reviews Immunology. 2009;9:581-93
23. Burger D, Vinas JL, Akbari S, Dehak H, Knoll W, Gutsol A. et al. Human endothelial colony-forming cells protect against acute kidney injury: role of exosomes. The American journal of pathology. 2015;185:2309-23
24. Xin H, Li Y, Chopp M. Exosomes/miRNAs as mediating cell-based therapy of stroke. Frontiers in cellular neuroscience. 2014;8:377
25. Zhang B, Wang M, Gong A, Zhang X, Wu X, Zhu Y. et al. HucMSC-Exosome Mediated-Wnt4 Signaling Is Required for Cutaneous Wound Healing. Stem cells (Dayton, Ohio). 2015;33:2158-68
26. Zhang L, Wrana JL. The emerging role of exosomes in Wnt secretion and transport. Curr Opin Genet Dev. 2014;27:14-9
27. Gross JC, Chaudhary V, Bartscherer K, Boutros M. Active Wnt proteins are secreted on exosomes. Nat Cell Biol. 2012;14:1036-45
28. Meng Z, Moroishi T, Guan KL. Mechanisms of Hippo pathway regulation. Genes Dev. 2016;30:1-17
29. Yu FX, Zhao B, Guan KL. Hippo Pathway in Organ Size Control, Tissue Homeostasis, and Cancer. Cell. 2015;163:811-28
30. Tao SC, Gao YS, Zhu HY, Yin JH, Chen YX, Zhang YL. et al. Decreased extracellular pH inhibits osteogenesis through proton-sensing GPR4-mediated suppression of yes-associated protein. Sci Rep. 2016;6:26835
31. Park HW, Kim YC, Yu B, Moroishi T, Mo JS, Plouffe SW. et al. Alternative Wnt Signaling Activates YAP/TAZ. Cell. 2015;162:780-94
32. Bell DM, Leung KK, Wheatley SC, Ng LJ, Zhou S, Ling KW. et al. SOX9 directly regulates the type-II collagen gene. Nat Genet. 1997;16:174-8
33. Oldershaw RA, Baxter MA, Lowe ET, Bates N, Grady LM, Soncin F. et al. Directed differentiation of human embryonic stem cells toward chondrocytes. Nat Biotechnol. 2010;28:1187-94
34. Liu CF, Lefebvre V. The transcription factors SOX9 and SOX5/SOX6 cooperate genome-wide through super-enhancers to drive chondrogenesis. Nucleic Acids Res. 2015;43:8183-203
35. Barter MJ, Tselepi M, Gomez R, Woods S, Hui W, Smith GR. et al. Genome-Wide MicroRNA and Gene Analysis of Mesenchymal Stem Cell Chondrogenesis Identifies an Essential Role and Multiple Targets for miR-140-5p. Stem cells (Dayton, Ohio). 2015;33:3266-80
36. Miyaki S, Sato T, Inoue A, Otsuki S, Ito Y, Yokoyama S. et al. MicroRNA-140 plays dual roles in both cartilage development and homeostasis. Genes & development. 2010;24:1173-85
37. Buechli ME, Lamarre J, Koch TG. MicroRNA-140 expression during chondrogenic differentiation of equine cord blood-derived mesenchymal stromal cells. Stem Cells Dev. 2013;22:1288-96
38. Karlsen TA, Jakobsen RB, Mikkelsen TS, Brinchmann JE. microRNA-140 targets RALA and regulates chondrogenic differentiation of human mesenchymal stem cells by translational enhancement of SOX9 and ACAN. Stem cells and development. 2014;23:290-304
39. Liang Y, Duan L, Xiong J, Zhu W, Liu Q, Wang D. et al. E2 regulates MMP-13 via targeting miR-140 in IL-1beta-induced extracellular matrix degradation in human chondrocytes. Arthritis Res Ther. 2016;18:105
40. Li X, Zhen Z, Tang G, Zheng C, Yang G. MiR-29a and MiR-140 Protect Chondrocytes against the Anti-Proliferation and Cell Matrix Signaling Changes by IL-1beta. Mol Cells. 2016;39:103-10
41. Ding J, Huang S, Wu S, Zhao Y, Liang L, Yan M. et al. Gain of miR-151 on chromosome 8q24.3 facilitates tumour cell migration and spreading through downregulating RhoGDIA. Nat Cell Biol. 2010;12:390-9
42. Wu J, Wu G, Lv L, Ren YF, Zhang XJ, Xue YF. et al. MicroRNA-34a inhibits migration and invasion of colon cancer cells via targeting to Fra-1. Carcinogenesis. 2012;33:519-28
43. Du J, Wang J, Tan G, Cai Z, Zhang L, Tang B. et al. Aberrant elevated microRNA-146a in dendritic cells (DC) induced by human pancreatic cancer cell line BxPC-3-conditioned medium inhibits DC maturation and activation. Med Oncol. 2012;29:2814-23
44. Zhu H, Cheng X, Niu X, Zhang Y, Guan J, Liu X. et al. Proton-sensing GPCR-YAP Signalling Promotes Cell Proliferation and Survival. Int J Biol Sci. 2015;11:1181-9
45. Zhu H, Guo S, Zhang Y, Yin J, Yin W, Tao S. et al. Proton-sensing GPCR-YAP Signalling Promotes Cancer-associated Fibroblast Activation of Mesenchymal Stem Cells. Int J Biol Sci. 2016;12:389-96
46. Aszodi A, Chan D, Hunziker E, Bateman JF, Fassler R. Collagen II is essential for the removal of the notochord and the formation of intervertebral discs. J Cell Biol. 1998;143:1399-412
47. Lee WK, Yu SM, Cheong SW, Sonn JK, Kim SJ. Ectopic expression of cyclooxygenase-2-induced dedifferentiation in articular chondrocytes. Exp Mol Med. 2008;40:721-7
48. Prasadam I, Zhou Y, Shi W, Crawford R, Xiao Y. Role of dentin matrix protein 1 in cartilage redifferentiation and osteoarthritis. Rheumatology (Oxford). 2014;53:2280-7
49. Sagar DR, Ashraf S, Xu L, Burston JJ, Menhinick MR, Poulter CL. et al. Osteoprotegerin reduces the development of pain behaviour and joint pathology in a model of osteoarthritis. Annals of the rheumatic diseases. 2014;73:1558-65
50. Janusz MJ, Hookfin EB, Heitmeyer SA, Woessner JF, Freemont AJ, Hoyland JA. et al. Moderation of iodoacetate-induced experimental osteoarthritis in rats by matrix metalloproteinase inhibitors. Osteoarthritis and cartilage / OARS, Osteoarthritis Research Society. 2001;9:751-60
51. Pritzker KP, Gay S, Jimenez SA, Ostergaard K, Pelletier JP, Revell PA. et al. Osteoarthritis cartilage histopathology: grading and staging. Osteoarthritis and cartilage / OARS, Osteoarthritis Research Society. 2006;14:13-29
52. Zhen G, Wen C, Jia X, Li Y, Crane JL, Mears SC. et al. Inhibition of TGF-beta signaling in mesenchymal stem cells of subchondral bone attenuates osteoarthritis. Nat Med. 2013;19:704-12
53. Yimlamai D, Christodoulou C, Galli GG, Yanger K, Pepe-Mooney B, Gurung B. et al. Hippo pathway activity influences liver cell fate. Cell. 2014;157:1324-38
54. Liu-Chittenden Y, Huang B, Shim JS, Chen Q, Lee SJ, Anders RA. et al. Genetic and pharmacological disruption of the TEAD-YAP complex suppresses the oncogenic activity of YAP. Genes & development. 2012;26:1300-5
55. Im GI. Regeneration of articular cartilage using adipose stem cells. J Biomed Mater Res A. 2016;104:1830-44
56. Hu W, Zhang W, Li F, Guo F, Chen A. miR-139 is up-regulated in osteoarthritis and inhibits chondrocyte proliferation and migration possibly via suppressing EIF4G2 and IGF1R. Biochemical and biophysical research communications. 2016;474:296-302
57. Church V, Nohno T, Linker C, Marcelle C, Francis-West P. Wnt regulation of chondrocyte differentiation. J Cell Sci. 2002;115:4809-18
58. Bradley EW, Drissi MH. Wnt5b regulates mesenchymal cell aggregation and chondrocyte differentiation through the planar cell polarity pathway. J Cell Physiol. 2011;226:1683-93
59. Bradley EW, Drissi MH. WNT5A regulates chondrocyte differentiation through differential use of the CaN/NFAT and IKK/NF-kappaB pathways. Mol Endocrinol. 2010;24:1581-93
60. Deng Y, Wu A, Li P, Li G, Qin L, Song H. et al. Yap1 Regulates Multiple Steps of Chondrocyte Differentiation during Skeletal Development and Bone Repair. Cell Rep. 2016;14:2224-37
61. Ferguson SW, Nguyen J. Exosomes as therapeutics: The implications of molecular composition and exosomal heterogeneity. J Control Release. 2016;228:179-90
62. Ma G, Luo Y, Zhu H, Luo Y, Korhonen PK, Young ND. et al. MicroRNAs of Toxocara canis and their predicted functional roles. Parasit Vectors. 2016;9:229
63. Li X, Lv Y, Hao J, Sun H, Gao N, Zhang C. et al. Role of microRNA-4516 involved autophagy associated with exposure to fine particulate matter. Oncotarget. 2016
64. Xia ZY, Hu Y, Xie PL, Tang SY, Luo XH, Liao EY. et al. Runx2/miR-3960/miR-2861 Positive Feedback Loop Is Responsible for Osteogenic Transdifferentiation of Vascular Smooth Muscle Cells. Biomed Res Int. 2015;2015:624037
65. Lu W, Han L, Su L, Zhao J, Zhang Y, Zhang S. et al. A 3'UTR-associated RNA, FLJ11812 maintains stemness of human embryonic stem cells by targeting miR-4459. Stem cells and development. 2015;24:1133-40
Author contact
![]() Corresponding authors: Chang-Qing Zhang, Department of Orthopaedic Surgery, Shanghai Jiao Tong University Affiliated Sixth People's Hospital, 600 Yishan Road, Shanghai 200233, China. E-mail: zhangcqedu.cn. Shang-Chun Guo, Institute of Microsurgery on Extremities, Shanghai Jiao Tong University Affiliated Sixth People's Hospital, 600 Yishan Road, Shanghai 200233, China. E-mail: achunicom.
Corresponding authors: Chang-Qing Zhang, Department of Orthopaedic Surgery, Shanghai Jiao Tong University Affiliated Sixth People's Hospital, 600 Yishan Road, Shanghai 200233, China. E-mail: zhangcqedu.cn. Shang-Chun Guo, Institute of Microsurgery on Extremities, Shanghai Jiao Tong University Affiliated Sixth People's Hospital, 600 Yishan Road, Shanghai 200233, China. E-mail: achunicom.
 Global reach, higher impact
Global reach, higher impact