Impact Factor
Theranostics 2016; 6(5):638-649. doi:10.7150/thno.14596 This issue Cite
Research Paper
Spectrum and Classification of ATP7B Variants in a Large Cohort of Chinese Patients with Wilson's Disease Guides Genetic Diagnosis
1. Department of Neurology and Research Center of Neurology in Second Affiliated Hospital, and the Collaborative Innovation Center for Brain Science, Zhejiang University School of Medicine, Hangzhou, China;
2. Department of Neurology and Institute of Neurology, First Affiliated Hospital, Fujian Medical University, Fuzhou, China;
3. State Key Laboratory of Genetic Engineering, Institutes of Biomedical Sciences, School of Life Science, Fudan University, Shanghai, China;
4. Department of Neurology and Institute of Neurology, Huashan Hospital, Shanghai Medical College, Fudan University, Shanghai, China;
5. Center for Genomic Translation and Health Intervention, School of Public Health, Fudan University, Shanghai, China.
* These authors contributed equally to this work.
Abstract
Background: Wilson's disease (WD) is an autosomal recessive disorder of copper metabolism caused by ATP7B pathogenic mutations. The symptoms of WD can be effectively prevented if the affected individuals are identified and intervened early. However, clinical utility of this molecular analysis is challenging due to hundreds of variants with various clinical effects in the gene. Here, we aim to describe the spectrum of ATP7B variants and assess their clinical effects in the Han Chinese population.
Methods: The ATP7B gene was directly sequenced in 632 unrelated WD patients and 503 unrelated healthy individuals. The effects of identified variants were classified according to the American College of Medical Genetics and Genomics (ACMG) standards and guidelines. Different frequency of variants observed in both cases and controls were tested using Chi-square or Fisher's exact tests.
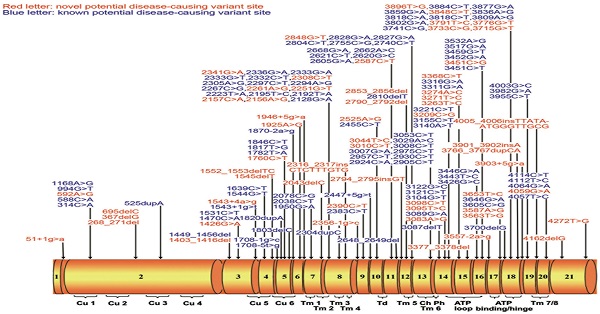
Results: We detected 161 non-synonymous variants in these 632 WD patients, 58 of which were novel. Among these variants, 78, 64, 8, 4, and 7 were classified as 'pathogenic variants', 'likely pathogenic variants', 'variants with uncertain significance', 'likely benign variants', and 'benign variants', respectively. Ninety percent (569/632) of these WD patients can be genetically diagnosed with two or more 'pathogenic' or 'likely pathogenic' variants. The 14 most common disease-causing variants were found at least once in 94% (537/569) of genetically diagnosed patients.
Conclusions: These data expand the spectrum of ATP7B variants and facilitate effective screening for ATP7B variants for early diagnosis of WD and development of individualized treatment regimens.
Keywords: Wilson's disease, ATP7B, variants classification.
 Global reach, higher impact
Global reach, higher impact