13.3
Impact Factor
Theranostics 2015; 5(12):1328-1342. doi:10.7150/thno.11432 This issue Cite
Research Paper
Salmonella typhimurium Suppresses Tumor Growth via the Pro-Inflammatory Cytokine Interleukin-1β
1. Laboratory of In Vivo Molecular Imaging, Department of Nuclear Medicine, Chonnam National University Medical School and Hwasun Hospital, Gwangju, Republic of Korea
2. Department of Microbiology, Chonnam National University Medical School, Gwangju, Republic of Korea
3. School of Life Sciences, Gwangju Institute of Science and Technology (GIST), 261 Cheomdan-gwagiro, Buk-gu, Gwangju 500-712, Republic of Korea
4. Academy of Immunology and Microbiology (AIM), Institute of Basic Science (IBS), Pohang, Republic of Korea
5. Fraunhofer IZI, Leipzig, Germany
6. Division of Integrative Biosciences and Biotechnology (IBB), Pohang University of Science and Technology, Pohang, 790-784, Republic of Korea
*These authors contributed equally to this work.
Received 2014-12-25; Accepted 2015-8-23; Published 2015-10-6
Abstract
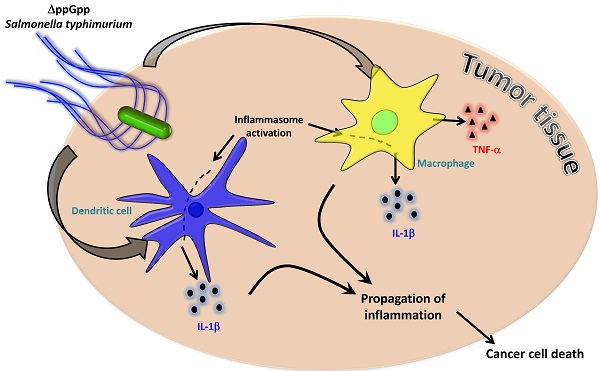
Although strains of attenuated Salmonella typhimurium and wild-type Escherichia coli show similar tumor-targeting capacities, only S. typhimurium significantly suppresses tumor growth in mice. The aim of the present study was to examine bacteria-mediated immune responses by conducting comparative analyses of the cytokine profiles and immune cell populations within tumor tissues colonized by E. coli or attenuated Salmonellae. CT26 tumor-bearing mice were treated with two different bacterial strains: S. typhimurium defective in ppGpp synthesis (ΔppGpp Salmonellae) or wild-type E. coli MG1655. Cytokine profiles and immune cell populations in tumor tissue colonized by these two bacterial strains were examined at two time points based on the pattern of tumor growth after ΔppGpp Salmonellae treatment: 1) when tumor growth was suppressed ('suppression stage') and 2) when they began to re-grow ('re-growing stage'). The levels of IL-1β and TNF-α were markedly increased in tumors colonized by ΔppGpp Salmonellae. This increase was associated with tumor regression; the levels of both IL-1β and TNF-α returned to normal level when the tumors started to re-grow. To identify the immune cells primarily responsible for Salmonellae-mediated tumor suppression, we examined the major cell types that produce IL-1β and TNF-α. We found that macrophages and dendritic cells were the main producers of TNF-α and IL-1β. Inhibiting IL-1β production in Salmonellae-treated mice restored tumor growth, whereas tumor growth was suppressed for longer by local administration of recombinant IL-1β or TNF-α in conjunction with Salmonella therapy. These findings suggested that IL-1β and TNF-α play important roles in Salmonella-mediated cancer therapy. A better understanding of host immune responses in Salmonella therapy may increase the success of a given drug, particularly when various strategies are combined with bacteriotherapy.
Keywords: Bacteria-mediated cancer therapy, Salmonella typhimurium, E. coli, IL-1β, TNF-α, dendritic cells
Introduction
Bacteria-mediated cancer therapy (BCT) was first introduced in the mid-19th century, when bacteria were used to treat solid tumors [1]. This approach has several advantages over other conventional therapies: bacteria have the ability to target tumors specifically; they actively proliferate in a variety of malignant tumors; they are easy to manipulate at the genetic level; and they are inexpensive to produce [2]. A number of obligatory and facultative bacteria, such as Clostridium [3-6], Salmonella typhimurium [7-11], Bifidobacterium [12], Escherichia coli [13, 14], and Listeria [15] preferentially localize, proliferate, and produce anti-cancer molecules in solid tumors. Although BCT is a promising strategy for cancer treatment, the actual mechanisms that suppress tumor growth are unclear.
We previously showed that two bacterial strains, E. coli K-12 strain (MG1655) and attenuated S. typhimurium defective in the synthesis of ppGpp (ΔppGpp S. typhimurium) have tumor-targeting ability and developed diverse BCT strategies based on these strains [9, 13, 14, 16-20]. Of these two strains, only ΔppGpp Salmonellae significantly suppressed tumor growth [9, 14]. Thus, the following questions arise: why do these two bacterial strains have different tumor-suppressing abilities? Might the difference be related to the triggering of different host immune responses? If so, what kind of immune response is triggered by Salmonella?
One possible answer is that different bacterial strains induce different cytokine profiles in the tumor microenvironment. Bacteria (or bacterial components) induce immune responses by increasing the production of pro-inflammatory cytokines and by activating different immune cell populations [21]. For example, gram-negative bacteria such as Salmonellae and E. coli express lipopolysaccharide (LPS), which is the ligand for TLR4 (LPS triggers inflammatory reactions and the secretion of pro-inflammatory cytokines [22, 23]).
The role of TNF-α in BCT has been studied [22, 24, 25]. TNF-α induced tumoricidal effects associated with gram-negative tumor-colonizing bacteria; thus this does not explain the contrary results achieved with the two different gram-negative strains. To date, although several studies reported underlying mechanism of tumor regression upon bacterial injection, it has not been suggested clearly for involvement of other immune modulators, except TNF-α [26].
Although ΔppGpp Salmonellae suppressed tumor growth, the effect was not permanent. Tumor growth was significantly reduced for several days (1-10 days), but the tumor began to re-grow after this time. Therefore, we examined bacteria-mediated immune responses by comparing the cytokine and immune cell profiles in tumor tissues colonized by E. coli or ΔppGpp Salmonellae during two separate time periods: the period of tumor suppression ('suppression stage') and the period of tumor re-growth ('re-growing stage'). The aim was to identify the specific factors responsible for the anti-tumor immunity evoked by ΔppGpp Salmonellae.
Materials and Methods
Tumor cell line and animal model
The murine CT26 colon adenocarcinoma cell line was obtained from the American Type Culture Collection (CRL-2638). The cells were grown at 37°C/5% CO2 in complete DMEM (cDMEM, endotoxin-free DMEM high glucose medium [Hyclone, Thermal scientific] supplemented with 10% FBS [Gibco-Invitrogen] and 1% penicillin and streptomycin [Gibco-Invitrogen]).
Male BALB/c mice (4-5 weeks old) were purchased from Orient BIO (Charles River Laboratories, Korea) and maintained under specific pathogen-free conditions. All animal experiments conformed to the Chonnam National University Animal Research Committee protocols. CT26 cells (1 × 106) were suspended in PBS and implanted subcutaneously into the right thigh of each mouse. Tumor volume (mm3) was measured with a caliper. The optimal size for analysis was 180 mm3. Tumor volume was measured every 3 days until the end of experiment. Bacteria-mediated immune responses were examined by comparing the cytokine and immune cell profiles in tumor tissues during two separate time periods: the suppression stage (2 days post-inoculation, 2dpi) and the re-growing stage (at 15 dpi or when the tumor volume reached 1200 mm3). When the tumor volume exceeded 2000mm3, the animals were euthanized and excluded from the experiment.
Bacterial infection
Attenuated S. typhimurium (defective in the synthesis of ppGpp (RelA::cat, SpoT::kan)) expressing the bacterial luciferase gene lux (StΔppGpp-lux; SHJ2168), and wild-type E. coli MG1655 were used for the study [9, 13]. Mice received an intravenous injection of SLΔppGpp/lux (4.5 × 107 cells) or E. coli MG1655 (5 × 107 cells) in PBS, respectively. Culture, harvesting, and preparation of bacteria was described previously [19].
Optical bioluminescence imaging
Bioluminescence imaging was performed as previously described using an IVIS 100 system (Caliper) [19].
Preparation of single cell suspensions
Tumors were excised at both the suppression and re-growing stages (based on the ΔppGpp Salmonellae-treated group). Single cell suspensions were created according to the 'Dissociation of cells from primary tissue' protocol provided by Invitrogen Laboratories. Briefly, the tumor was washed twice with PBS and minced into 2-3 mm3 pieces. The pieces were then incubated with 0.25% trypsin (Gibco-Invitrogen) for 2 hr at 4°C, followed by a further incubation for 20 min at 37°C. After inactivating the trypsin, the tissue was ground gently, followed by filtration through 40 µm cell strainer (Falcon) to collect digested tissues. Dead cells were removed by Percoll gradient centrifugation (isopynic centrifugation) as previously described [27]. Collected cells were washed with PBS and re-suspended in cDMEM. To obtain lymphocytes from the tumor-draining lymph nodes, a single cell suspension was first prepared, and total lymphocytes were suspended in RPMI (Gibco) supplemented with 10% fetal bovine serum (Hyclone), 3 mM L-glutamine (Sigma), 10 mM HEPES (Sigma), 100 U/ml penicillin (Sigma), 100 U/ml streptomycin (Sigma), and 0.05 mM 2-beta-mercaptoethanol (Sigma).
mRNA isolation, cDNA synthesis, and quantitative RT-PCR (qRT-PCR)
Cells isolated from the tumor were homogenized in TRIzol Reagent (Molecular Research Center), and mRNA was obtained following the recommended protocol. To generate cDNA, 1 μg of total mRNA was firstly mixed with oligo (dT) primers (Promega, USA) and incubated at 65°C for 5 min. The primer/template mix was chilled on ice and reaction mix containing Improm-II Reverse Transcriptase (Promega) was added. The samples were then incubated at 30°C for 10 min, followed by 42°C for 60 min, and 70°C for 15 min in PCR Thermal Cycler Dice machine (Takara, Japan). cDNA was amplified using the RT-PCR primer sets listed in the Table 1. PCR reactions were performed in Rotor-Gene 3000 (Corbett Robotics, Australia). SYBR Premix Ex Taq (Takara) was used to detect amplification under the following conditions: 10 min at 95°C, followed by 40 three-temperature cycles (15 s at 95°C, 30 s at 60°C, and 15 s at 72°C). All reactions were run in triplicate. Results were analyzed with Rotor-Gene Software. HPRT was used as housekeeping gene to assess target gene expression.
Western blot analysis
To measure the amount of pro-IL-1β and caspase-1, isolated tumor cells were lysed in protein extraction solution (Intron Biotechnology), subjected to SDS-PAGE, and subsequently transferred to a nitrocellulose membrane (Bio-Rad). The membranes were probed with polyclonal goat anti-mouse IL-1β (AF-401-NA, R&D Systems; 1:2000 dilution) or polyclonal rabbit anti-mouse caspase-1 (sc-514, Santa Cruz Biotechnology; 1:1000 dilution) antibodies, followed by horseradish peroxidase-conjugated donkey anti-goat IgG (sc-2020, Santa Cruz Biotechnology; 1:2000 dilution) or polyclonal goat anti-rabbit IgG/HRP (P0448, Dako; 1:2000 dilution), respectively. Bioluminescence was catalyzed using a Western Blotting Luminol Reagent (Santa Cruz), and bands were detected in a luminescent image analyser LAS-3000 (Fujifilm).
Primer sequences for cytokines and housekeeping genes.
| Name | Sequence (5' → 3') | |
|---|---|---|
| IL-1β | IL-1β Forward | GCA ACT GTT CCT GAA CTC AAC T |
| IL-1β Reverse | ATC TTT TGG GGT CCG TCA ACT | |
| TNF-α | TNF-α Forward | CAT CTT CTC AAA ATT CGA GTG ACA A |
| TNF-α Reverse | TGG GAG TAGACA AGG TAC AAC CC | |
| IL-6 | IL-6 Forward | GAG GAT ACC ACT CCC AAC AGA CC |
| IL-6 Reverse | AAG TGC ATC ATC ATC GTT GTT CA | |
| IFN-γ | IFN-γ Forward | TCA AGT GGC ATA GAT GTG GAAGAA |
| IFN-γ Reverse | TGG CTC TGC AGG ATT TTC ATG | |
| TGF-β | TGF-β Forward | GAA GGC AGA GTT CAG GGT CTT |
| TGF-β Reverse | GGT TCC TGT CTT TGT GGT GAA | |
| HPRT | HPRT Forward | TTA TGG ACA GGA CTG AAA GAC |
| HPRT Reverse | GCT TTA ATG TAA TCC AGC AGG T |
Measurement of cytokines in tumor tissues and serums
Tumor tissues were homogenized in protein extraction solution (Intron Biotechnology), and the supernatant was collected by centrifugation. Blood was collected by cardiac puncture at 0, 0.5, 3, 12h and 2dpi and then serum was harvested by removing blood clot after centrifugation. Cytokine levels were measured using individual ELISA kits (eBioscience) according to the manufacturer's instructions. The substrate color reaction was measured at 450 nm in an ELISA reader (SpectraMax, Molecular Devices).
Flow cytometry analysis
For surface staining, cells were washed with PBS and then incubated with fluorescence-labeled antibodies against target cell surface molecules for 20 min in the dark. For intracellular staining, cells were re-stimulated with phorbol 12-myristate 13-acetate (PMA) and ionomycin (Calbiochem) in GolgiStop (1 mg/ml; BD Biosciences) at 37℃ for 5 hours. After cells were stained for surface markers (see below) for 20 min, they were fixed in IC Fixation buffer (eBioscience) for 30 min. Cells were then washed and re-suspended in permeabilization buffer (eBioscience) containing intracellular detection antibodies (see below) for 20 min. To detect Foxp3 and Helios, cells stained for surface markers were suspended in 1 ml of fixation/permeabilization buffer for 1 hour, re-suspended in permeabilization buffer, and then stained with each of the antibodies. Cells were then washed and re-suspended in PBS, and at least 10,000 events were analyzed using an LSR-Fortessa flow cytometer (BD Bioscience), according to the manufacturer's instructions. Data were evaluated using FlowJo software (TreeStar). Antibodies to the following were used for surface staining: CD3 (17A2), CD4 (RM4-5), CD8 (53-6.7), B220 (RA3-6B2), CD11b (M1/70), CD11c (N418), CD25 (PC61), CD45 (30-F11), F4/80 (BM8), MHCII (M5/114.15.2), CD68 (FA-11), Gr1 (RB6-8C5), Ly6C (HK1.4), Ly6G (1A8), CD103 (2E7), CTLA4 (UC10-4B9), and ICOS (C398.4A) (all from BioLegend). The following antibodies were used for intracellular staining: IL-1β (NJTEN3), TNF-α (MP6-XT22), Foxp3 (FJK-16s), and Helios (22F6) (all from eBioscience).
Apoptosis assay
Cells were isolated from tumor tissue and separated using magnetic beads (Miltenyi Biotech). As there is no common marker for CT26 tumor cells, antibodies against the following representative immune cell markers were used: CD4 (RM4-5), CD8 (53-6.7), CD19 (6D5), B220 (RA3-6B2), CD11b (M1/70), CD11c (N418), CD24 (M1/69), CD25 (PC61), CD49b (DX5) (all from BioLegend), and γδTCR (eBioCL3) (eBioscience). Cells that expressed at least one of these molecules were considered to be immune cells, whereas those that did not were considered to be tumor cells. This procedure was used to examine apoptotic immune cells and tumor cells isolated from tumor tissues. Cells were washed with PBS and re-suspended in binding buffer (BD Pharmingen) prior to incubation with Annexin V and 7-AAD (BD Pharmingen). The populations of viable cells (Annexin V-7AAD-), early apoptotic cells (Annexin V+7AAD-), and late apoptotic or necrotic cells (Annexin V+7AAD+) were analyzed using an LSR-Fortessa flow cytometer (BD Biosciences), according to the manufacturer's instructions. Data were evaluated using FlowJo software (TreeStar).
Treatment with an anti-IL-1β antibody or recombinant IL-1β
To deplete IL-1β, mice were injected with 5 μg of an IL-1β-specific antibody (AF401-NA, R&D Systems) 1 day before infection by bacteria and then twice a week for 2 weeks thereafter. The control group received hamster immunoglobulin (EQUITECH Bio) according to the same schedule. The Salmonellae and IL-1β combination therapy groups received an intratumoral injection of recombinant IL-1β (0.5 μg; 401-ML/CF, R&D Systems) suspended in PBS every 2 days starting at 5 dpi and continuing until 11 dpi. Recombinant IL-1β was injected using a Microliter syringe (Hamilton Company) fitted with a PrecisionGlide Needle (BDM011455-1, BD Bioscience).
Treatment with an anti-TNF-α antibody or recombinant TNF-α
Procedures similar to those described for the anti-IL-1β antibody or recombinant IL-1β were used. To deplete TNF-α, mice were injected with 50 μg of a TNF-α-specific antibody (AF410-NA; R&D Systems) 1 day before infection by bacteria and then twice per week for 2 weeks thereafter. The Salmonellae and TNF-α combination therapy groups received an intratumoral injection of recombinant TNF-α (410-MT/CF; 0.25 μg in PBS; R&D Systems) every 2 days starting at 5 dpi and continuing until 11 dpi. Recombinant TNF-α injections were performed using a Microliter syringe (Hamilton Company) fitted with a PrecisionGlide Needle (BDM011455-1; BD Biosciences).
Immunohistochemistry and immunofluorescence staining
Tumor tissues were excised at 2 dpi, fixed in 4% paraformaldehyde at 4°C for 4 hr, immersed in 30% sucrose/PBS overnight at 4°C, washed with PBS, and then embedded in OCT compound (Leica microsystem) in a plastic container.
OCT-mounted tissues were cryo-sectioned (5 μm) using a Microm HM 525 cryotome (Thermo Scientific). Sections were incubated with streptavidin-biotin labeled antibodies (diluted 1:100-1:500) against CD45 (MCA1031GA, ABD Serotec), CD68 (MCA1957GA, ABD Serotec), Ly-6G/Ly-6C (sc-71674, Santa Cruz), and CD11c (550283, BD Pharmingen). Briefly, sections were rehydrated in cold acetone for 15 min and washed with PBS. Endogenous peroxidase activity and non-specific binding were blocked with 0.3% H2O2 and 5% BSA in TBS containing 0.1% Tween 20, respectively. After blocking, the sections were incubated with the primary antibody overnight at 4°C, followed by the appropriate secondary antibody for 2 hr at room temperature. Nuclei were stained with hematoxylin. Finally, the sections were mounted in Immu-Mount reagent (238402, Thermo Scientific).
For immunofluorescence staining, tissues sections were permeabilized and blocked with TBS containing 0.1% Tween 20, 0.3% Triton X-100, and 5% BSA. The sections were then incubated with rat anti-neutrophil antibodies (sc-71674, Santa Cruz), rat anti-CD68 (MCA1957GA, ABD Serotec), hamster anti-CD11c (550283, BD Pharmingen), rabbit anti-IL-1β (sc-7884, Santa Cruz), or goat anti-TNF-α (sc-1350, Santa Cruz) overnight at 4°C. Alexa Fluor 488 donkey anti-rat (A21208), Alexa Fluor 555 donkey anti-rabbit (A31572), Alexa Fluor 568 goat anti-hamster (A21112), Alexa Fluor 488 goat anti-rabbit (A11008), Alexa Fluor 555 donkey anti-goat (A21432), and Alexa Fluor 488 donkey anti-goat (A11055) antibodies were used as secondary antibodies (all from Invitrogen). All antibodies were diluted to 1:200 in TBS containing 0.1% Tween 20, 1% BSA, and 0.1% Triton X-100. Samples were mounted with Pro-long® Gold antifade reagent (P36930, Invitrogen) and analyzed under a Fluoview-1000 (FV-1000) laser scanning confocal microscope (Olympus).
Statistical analysis
All statistical analyses were performed using Excel and SPSS 15.0 software. The results are expressed as the mean ± SD. Data were analyzed using an independent t-test. P < 0.05 was considered significant.
Results
Salmonellae inhibit tumor growth
First, we examined the tumor-targeting activity of MG1655 and ΔppGpp Salmonellae in BALB/c mice bearing subcutaneous CT26 colon carcinomas. Both MG1655 and ΔppGpp Salmonellae showed similar tumor-targeting and proliferative capabilities when injected intravenously (i.v.) into tumor-bearing mice [9, 13] (Fig. 1A and S1A).
Injection of ΔppGpp Salmonellae led to significant suppression of tumor growth for around 10 dpi; however, the tumors tended to re-grow (Fig. 1B, C). By contrast, tumors infected with MG1655 grew at the same rate as non-treated tumors. Based on the results observed for ΔppGpp Salmonellae, we divided the tumor growth curve into two phases: the period of tumor suppression ('suppression stage', 1-10 dpi) and the period of tumor re-growth ('re-growing stage', ≥15 dpi). As previously reported [28], tumors colonized by Salmonellae showed massive hemorrhage and necrosis during the suppression stage (Fig. 1C).
Systemic injection of ΔppGpp Salmonellae (SLΔppGpp) into tumor-bearing mice induces significant growth suppression compared with E. coli (MG1655) or PBS injection. Mice (n = 5/group) were subcutaneously injected (into the right thigh) with CT26 cells (1 × 106). When the tumor volume reached 180 mm3, mice received an intravenous (i.v.) injection of PBS, MG1655 (5 × 107 CFU), or SLΔppGpp (4.5 × 107 CFU). (A) Distribution of bacteria visualized by in vivo bioluminescence imaging after injection of bacteria expressing bacterial luciferase (lux). (B) Tumor volume was then measured every 3 days until the end of experiment. The dotted line represents maximum tumor volume; mice were sacrificed when the tumor volume reached 2000mm3. Growth stages were divided into two phases based on the pattern of tumor growth after SLΔppGpp treatment; the suppression stage is the period during which the tumor stopped growing and/or shrunk in size, whereas the re-growing stage is the period during which the tumor grew again in SLΔppGpp-treated mice. (C) Images of the tumor graft in CT26-bearing mice. The shape of the tumor was noted before (0 dpi) and after treatment with PBS or bacteria (2 and 5 dpi). (D) Two days after treatment, cells were isolated from the tumor and the degree of apoptosis was measured using annexin V and 7AAD. Isolated cells were then separated into tumor cells and infiltrating immune cells to identify the cells in which apoptosis was induced. Thus, infiltrated immune cells were isolated by magnetic bead method using representative immune cell markers including CD4, CD8, CD19, B220, CD11b, CD11c, CD24, CD25, CD49b, and γδTCR. Apoptotic populations were investigated in two types of cells; immune cells and tumor cells isolated from tumor tissue. Data represent the mean ± SD of three independent experiments. *P < 0.05, **P < 0.005, and ***P < 0.001 vs. the control at Day 9 and 12 and vs. MG1655 at Day 15.
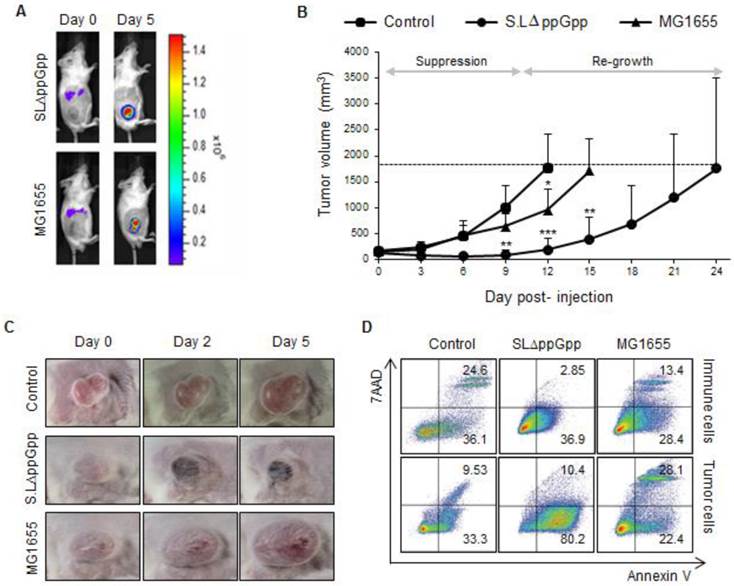
Next, we measured the amount of cell death. Cells isolated from the tumor mass during the suppression stage and re-growing stage were stained with markers of early and late apoptosis (annexin V and 7-aminoactinomycin (7-AAD), respectively). Isolated cells were then separated into tumor cells and infiltrating immune cells using magnetic beads and antibodies against representative immune cell markers (CD4, CD8, CD19, B220, CD11b, CD11c, CD24, CD25, CD49b, and γδTCR). Cells expressing at least one of these molecules were considered to be immune cells, whereas cells that did not were considered to be tumor cells. Apoptosis of immune cells and tumor cells was also examined (Fig. 1D). Compared with control and MG1655-treated tumors, a significant population of tumor cells was apoptotic within the ΔppGpp Salmonellae-treated tumors. The number of apoptotic tumor cells significantly decreased to low level during the re-growing stage in control, MG1655-, and ΔppGpp Salmonellae-treated tumors (Fig. S2A). On the other hand, the number of apoptotic immune cells in control, MG1655-, and ΔppGpp Salmonellae-treated tumors was not significantly different. These results suggest that the tumor-suppressive activity of ΔppGpp Salmonellae is responsible for the different levels of apoptosis induction observed at the suppression and re-growth stages.
Increased infiltration of immune cells into tumors colonized by ΔppGpp Salmonellae
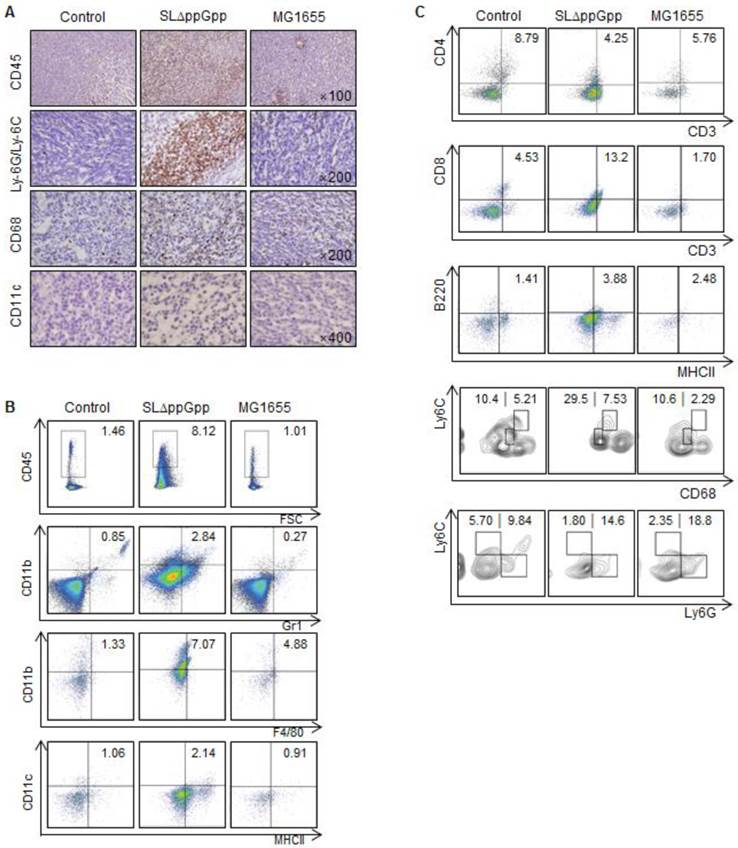
We next examined immune cell infiltration into tumors colonized by bacteria. We found that ΔppGpp Salmonellae treatment led to increased infiltration of tumor tissues by leukocytes (CD45+), including macrophages (CD11b+F4/80+ or CD68+), dendritic cells (DCs, CD11c+MHCII+), CD8+ T cells (CD3+CD8+), B cells (B220/CD45R+MHCII+), and neutrophils (CD11b+Gr1+ or Ly-6G/Ly-6C+) (Fig. 2A-C). Consistent with previous studies [24, 29-31], we found that infiltrated neutrophils accumulated at the border between the proliferative and necrotic areas of tumors colonized by ΔppGpp Salmonellae; however, only clusters of neutrophils were observed in tumors colonized by MG1655 (Fig. S3A). We also found that the numbers of neutrophils, macrophages, and DCs in the tumor-draining lymph nodes (tLNs) or spleens were higher in mice injected with ΔppGpp Salmonellae than in mice injected with MG1655 or in control mice, respectively (Fig. S3B, C).
We next used multi-color flow cytometry to further investigate the characteristics of macrophages and myeloid-derived suppressor cells (MDSCs) isolated from tumor tissue (Fig. S2A, C). To differentiate between M1 and M2 macrophages, we examined CD68 and Ly6C expression in CD11b+ F4/80+ cells [32]. The M1-like population (CD68intLy6Cint) in tumors colonized by ΔppGpp Salmonellae increased, whereas the M2-like population (CD68hiLy6Chi) was similar between groups (Fig. 2C). To examine the MDSC population, we gated on the CD11b+ cell population and then examined the expression of Ly6C and Ly6G [33]. Each MDSC subset within the monocytic MDSC (M-MDSC) and polymorphonuclear MDSC (PMN-MDSC) populations were examined according to their expression of Ly6C and Ly6G (M-MDSC: CD11b+Ly6ChiLy6G-; PMN-MDSC: CD11b+Ly6Clow/intLy6G+). The overall population of MDSCs within the tumor was not significantly different after infection by ΔppGpp Salmonellae (Fig. 2C).
At the re-growing stage, the proportion of infiltrating immune cells within tumors colonized by ΔppGpp Salmonella was comparable with that in tumors colonized by MG1655 or control tumors (Fig. S2B, C). In addition, the proportion of M1/M2 macrophages and MDSC was similar among all groups.
We also tested whether treatment with ΔppGpp Salmonellae induced changes in the regulatory T cell (Treg) population. Although the function of Tregs during tumor progression is still unclear, increased Treg numbers within tumors are thought to indicate a poor prognosis [34]. Thus, we examined whether the number and characteristics of Tregs are affected by ΔppGpp Salmonellae. We examined Treg cell characteristics by analyzing the expression of known Treg markers, including CD25, CTLA4, CD103, and ICOS, along with Helios (a thymic Treg marker) [35, 36]. Tumors colonized by ΔppGpp Salmonellae showed reduced expression of these markers at the suppression stage (Fig. S4A); however, the expression levels increased at the re-growing stage, reaching levels similar to those in control tumors or tumors colonized by MG1655 (Fig. S4B). These results suggest that ΔppGpp Salmonellae induce tumor suppression not only by increasing tumor-infiltrating immune cells such as M1 macrophages, neutrophils, DCs, CD8 T cells and B cells, but also by reducing both the Treg population and Treg effector function, mainly in tumor-bearing regions.
The production of IL-1β and TNF-α is elevated in tumors colonized by ΔppGpp Salmonellae
Because different immune cell populations accumulated in tumors colonized by Salmonellae, we speculated that effector molecules produced by these infiltrating cells might be involved in the observed tumor-suppressive effects. Thus, we next explored immunological changes in the tumor, focusing on cytokines that regulate the survival, proliferation, and differentiation of both immune cells and tumor cells. Tumor-bearing mice were treated with ΔppGpp Salmonellae, MG1655, or PBS, and cells were isolated at the suppression stage and re-growing stage. First, we examined the cytokine profiles using RT-PCR. We found that there was a marked increase in the amount of IL-1β and TNF-α transcripts in ΔppGpp Salmonellae-infected tumors; however, the levels of transcripts for other cytokines related to tumor development (IL-6, IFN-γ, and TGF-β) were similar to those in control tumors (Fig. 3A). There was no significant difference between the expression of these cytokines in MG1655-infected tumors and control tumors.
Colonization by SLΔppGpp leads to increased tumor infiltration by immune cells. (A) Tumor infiltration by immune cells was examined by immunohistochemistry at 2 dpi. Tumor tissue was stained for CD45, Ly-6G/Ly-6C, CD68, and CD11c, which are surface markers for hematopoietic cells, neutrophils, macrophages, and dendritic cells, respectively. Positive cells are brown. Sections were counterstained with hematoxylin. (B) Tumor-infiltrating cells were analyzed by flow cytometry. The indicated percentages represent hematopoietic cells (CD45+), neutrophils (CD11b+Gr1+), macrophages (CD11b+F4/80+), and dendritic cells (CD11c+MHCII+), respectively. (C) The indicated numbers represent the proportion of CD4+ T cells (CD3+CD4+), CD8+ T cells (CD3+CD8+), and B cells (B220+MHCII+) within CT26 tumors. The population of M1/M2 macrophages was examined by measuring CD68 and Ly6C expression in CD11b+F4/80+ cells (M1: CD68intLy6Cint, M2: CD68hiLy6Chi). MDSCs within CD11b+ cell population was also categorized M-MDSC (monocytic MDSC, CD11b+Ly6ChiLy6G-) and PMN-MDSC (polymorphonuclear MDSC, CD11b+Ly6ClowLy6G+). Data are representative of two individual experiments, each showing similar results.
Cytokine profile in SLΔppGpp- or MG1655-colonized tumors during the suppression stage. (A) Cells were isolated from tumors at 2dpi, and the expression of different cytokines was analyzed by qRT-PCR. HPRT was used as housekeeping gene to assess target gene expression. (B) Tumors were excised and homogenized in protein extraction solution, and the level of cytokines in the supernatant was measured by ELISA. (C) Single cell suspensions derived from tumor tissues were lysed, and total protein was obtained. The expression of caspase-1, pro-IL-1β, IL-1β, IL-18 and IPAF was examined by western blotting. β-actin was used as the loading control. (D) Serum TNF-α levels were measured in an ELISA at the indicated times. Data represent the mean ± SD. *P < 0.05, **P < 0.005, and ***P < 0.001.
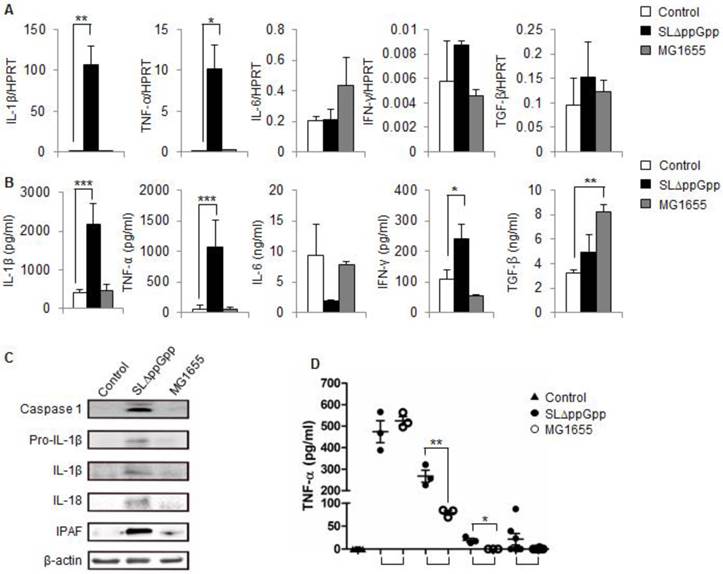
We next measured the expression of cytokines at the protein level. Again, IL-1β and TNF-α were highly expressed in ΔppGpp Salmonellae-infected tumors compared with non- or M1655-infected tumors. The expression of IL-6 and TGF-β in the ΔppGpp Salmonellae group was comparable with that in the control and MG1655 groups. The level of IFN-γ in the ΔppGpp Salmonellae group was about twice that in the control group; however, it was not as significant as IL-1β and TNF-α (Fig. 3B).
Previous studies highlighted the role of TNF-α in bacteriotherapy [22, 24, 25]; however, none reported changes in IL-1β expression. Thus, we examined whether the IL-1β processing pathway is activated in colonized tumors by comparing the levels of pro-IL-1β and caspase-1/IL-1 converting enzyme (ICE, which promotes the maturation of pro-IL-1β [37]) in cells from the ΔppGpp Salmonellae, MG1655, and PBS groups by western blotting. Strong signals were observed for both pro-IL-1β and caspase-1 in ΔppGpp Salmonellae-treated tumors, but not in MG1655-treated tumors (Fig. 3C). Consistent with pro-IL-1β production, we found that IL-1β was cleaved into its 17 kD mature form in ΔppGpp Salmonellae-treated tumors. In addition to IL-1β, we also detected mature IL-18 in ΔppGpp Salmonellae-treated tumors but not in MG1655- or PBS-treated tumors (Fig. 3C). The main pathway responsible for processing IL-1β is the inflammasome, a multi-protein complex that forms in the cytosol and activates caspase-1, leading to the subsequent cleavage and secretion of active IL-1β. Interestingly, we found that expression of the core molecule involved in inflammasome signaling, IPAF (ICE-protease-activating factor), was significantly higher in ΔppGpp Salmonellae-treated tumors than in MG1655- or PBS-treated tumors (Fig. 3C).
We also examined whether treatment with ΔppGpp Salmonellae affected serum levels of IL-1β or other cytokines. There was no significant difference in the serum levels of IL-1β (data not shown) in mice treated with ΔppGpp Salmonellae or MG1655 at any of the time points tested; however, the serum levels of TNF-α were significantly higher in mice treated with ΔppGpp Salmonellae than in mice treated with MG1655 at 3 h and 12 h post-bacterial injection (Fig. 3D).
Next, we examined whether the expression of these cytokines changed during the re-growing stage. We found that the expression of IL-1β and TNF-α transcripts was significantly down-regulated in ΔppGpp Salmonellae-treated tumors at re-growing stage, almost approaching the levels of the control group (Fig. S5A). Western blotting showed reduced expression of IL-1β, pro-IL-1β, and caspase-1 at this stage (Fig. S5B). Taken together, these results strongly suggest that ΔppGpp Salmonellae suppresses tumor growth by increasing the expression of IL-1β and TNF-α.
Dendritic cells and macrophages are responsible for increased IL-1β and TFN-α levels in tumors colonized by ΔppGpp Salmonellae
Monocyte lineage cells (macrophages in particular) may be the main sources of pro-inflammatory IL-1β and TNF-α [22, 38]. We next tried to identify the cells within colonized tumors responsible for producing these two cytokines. First, we double stained macrophages (CD68), neutrophils (Ly-6G/Ly-6C) or DCs (CD11c) for IL-1β or TNF-α and examined them under an immunofluorescence (IF) microscope. Second, we performed flow cytometry analysis using isolated cells from tLNs and spleens to identify and quantify IL-1β/TNF-α-producing cells. We could not collect data from the tumor region in the ΔppGpp Salmonellae-treated group because cells were either dying or already dead. Instead, we analyzed IL-1β and TNF-α expression in the tLNs and spleen. The IL-1β- and TNF-α-producing cells within the total cell, macrophage (CD11b+F4/80+), neutrophil (CD11b+Gr1+), and DC (CD11c+MHCII+) populations were analyzed after re-stimulation with PMA and ionomycin.
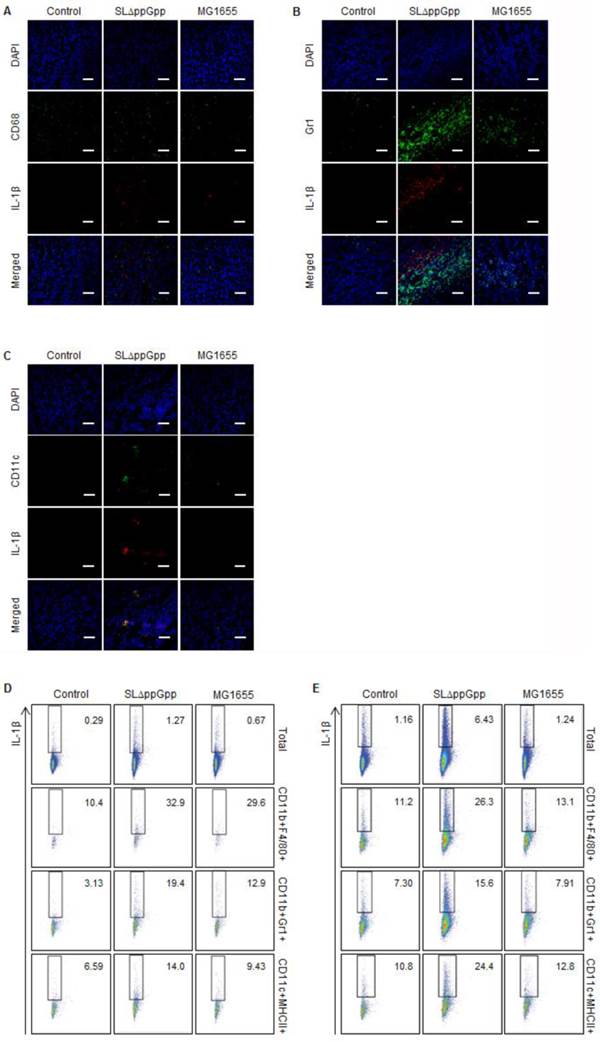
The results showed that macrophages and DCs might produce IL-1β. Some macrophages co-localized with the IL-1β signal (Fig. 4A), while neutrophils located beside IL-1β (Fig. 4B). Intriguingly, the pattern of the fluorescence signals suggested precise co-localization of IL-1β and DCs, indicating that most DCs within the tumor produce IL-1β (Fig. 4C). Flow cytometry analysis supported the IF microscopic findings showing that treatment with ΔppGpp Salmonellae led to a significant increase in IL-1β expression by DCs and macrophages in tLNs and the spleen (Fig. 4D, E). Thus, although DCs may not be the only producer of IL-1β, they appear to be the main producer in response to Salmonellae colonization.
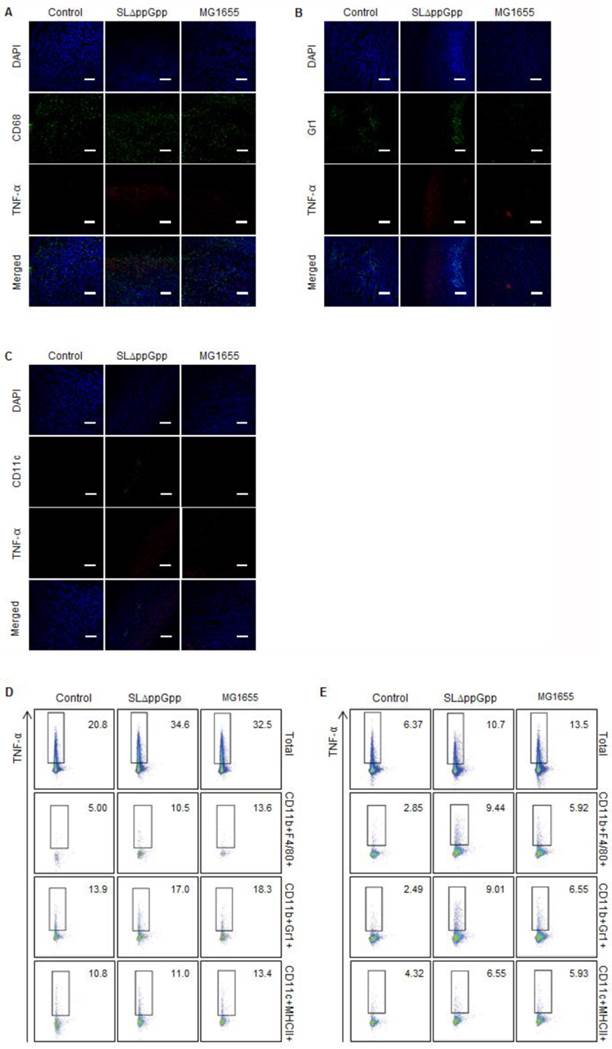
Next, we identified the immune cells that secrete TNF-α in bacteria-infected tumors. As shown previously, IF double-staining revealed that most CD68+ macrophages co-localized with the TNF-α signals in ΔppGpp Salmonella-infected tumors; however, some macrophages co-localized with TNF-α signals in MG1655-infected tumors (Fig. 5A). This clearly suggests that macrophages are the main producers of TNF-α in response to bacterial colonization. There was no correlation between the localization of neutrophils or DCs and TNF-α signals (Fig. 5B, C). Quantitative analysis by intracellular flow cytometry also revealed a significant increase in TNF-α expression by macrophages (Fig. 5D, E). Taken together, these results suggest that TNF-α and IL-1β in ΔppGpp Salmonellae-colonized tumors are mainly produced by CD68+ macrophages and CD11c+ DCs, respectively.
Inhibiting IL-1β production reduces the therapeutic efficacy of ΔppGpp Salmonellae, but co-treatment with recombinant IL-1β or TNF-α increases the therapeutic efficacy
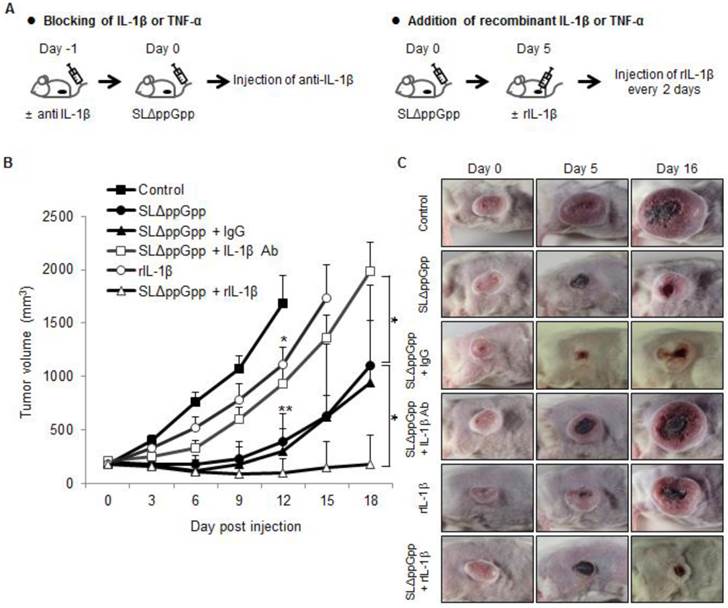
The anti-cancer activity of TNF-α is considered a promising target for BCT [25, 39, 40]. Here, we examined the role of IL-1β in Salmonella-mediated cancer therapy by blocking its activity with an anti-IL-1β antibody (Fig. 6A). An anti-IL-1β antibody abrogated the anti-tumor activity of ΔppGpp Salmonellae in tumor-bearing mice (Fig. 6B, C). Co-injection of an isotype control antibody (hamster IgG) did not abrogate the anti-tumor effect.
We next asked whether injecting recombinant IL-1β would increase the anti-tumor effects of ΔppGpp Salmonellae. Intratumoral injection of recombinant IL-1β from 5 dpi (followed by re-injection every 2 days) prolonged the tumor-suppressive activity of ΔppGpp Salmonellae (tumor growth remained suppressed until the end of the experiment). However, injection of recombinant IL-1β alone had only a mild suppressive effect compared with PBS (Fig. 6B, C). Treatment with an anti-IL-1β antibody or recombinant IL-1β did not lead to significant changes in the number of bacteria when compared with ΔppGpp Salmonellae treatment alone (Fig. S1C). Thus, neither recombinant IL-1β nor an anti-IL-1β antibody regulates the persistence of ΔppGpp Salmonellae in tumor tissue.
Considering that TNF-α has the potential to both stimulate and suppress tumor growth, we also examined its effect on tumor growth. Co-treatment with recombinant TNF-α prolonged the tumor-suppressive effects of ΔppGpp Salmonellae (Fig. S6A, B) in a manner similar to that observed after treatment with recombinant IL-1β (Fig. 6B, C). These results strongly suggest that both IL-1β and TNF-α play an important role in Salmonella-mediated cancer therapy.
Analysis of IL-1β producing cells. Tumor tissue was excised and fixed at 2 dpi, cryo-sectioned, and immunostained for IL-1β plus CD68 (macrophages), Ly-6G/Ly-6C (neutrophils), or CD11c (dendritic cells). Representative images from three independent experiments are shown (A-C). The green signal represents CD68 (A), Ly-6G/Ly-6C (B), or CD11c (C), whereas the red signal represents IL-1β. Sections were stained with DAPI (blue signal) as a control. Scale bar = 50 µm. (D, E) Cells were re-stimulated with PMA and ionomycin in the presence of GolgiStop, and the IL-1β+ population was examined by intracellular staining. Multi-color analysis was used to examine the proportion of IL-1β+ cells within the total cell, macrophage (CD11b+F4/80+), neutrophil (CD11b+Gr1+), and dendritic cell (CD11c+MHCII+) populations. The indicated values represent the IL-1β+ population in the tumor-draining lymph nodes (D) and spleen (E). Data are representative of two individual experiments, each with similar results.
Analysis of TNF-α producing cells. TNF-α expression in tumor tissue at 2 dpi was examined as described in the legend to Figure 4, except cells were stained for TNF-α (red) rather than IL-1β. Representative images from three independent experiments are shown (A-C). DAPI was used as a control. Scale bar = 100 µm. (D, E) Cells were re-stimulated with PMA and ionomycin in the presence of GolgiStop and the TNF-α+ cell population examined by intracellular staining. Multi-color analysis was used to examine the proportion of TNF-α+ cells within the total cell, macrophage (CD11b+F4/80+), neutrophil (CD11b+Gr1+), and dendritic cell (CD11c+MHCII+) populations. The indicated values represent the TNF-α+ population in the tumor-draining lymph nodes (D) and spleen (E). Data are representative of two individual experiments, each with similar results.
Role of IL-1β in SLΔppGpp-mediated cancer therapy. (A) Protocols for co-treatment with an IL-1β blocking antibody (left) or recombinant IL-1β (right), plus PBS or SLΔppGpp. (B) CT26 cells were transplanted into mice. Mice then received an intravenous (i.v.) injection of PBS (black rectangles) or SLΔppGpp (black circles). IL-1β depletion: mice received an i.v. injection of anti-IL-1β-specific antibody (IL-1β Ab) 1 day before SLΔppGpp treatment (Day 1). The antibody was then injected twice a week for 2 weeks (open rectangles). Control mice received isotype (IgG) according to the same schedule (black triangles). Treatment with recombinant IL-1β (rIL-1β): mice received an intratumoral (i.t.) injection of rIL-1β on 5 dpi (open triangles), followed by another i.t injection every other day up until 11 dpi. Another group of tumor-bearing mice received four i.t injections of rIL-1β alone (open circles). Data represent the mean ± SD. Results from at least three individual experiments are shown. *P < 0.05, **P < 0.005, and ***P < 0.001 vs. control at Day 12 and vs. SLΔppGpp at Day 18. (C) Photographs of representative animals in each group were taken before (0 dpi) and after treatment (5 and 16 dpi).
Discussion
The present study showed that tumor colonization by ΔppGpp Salmonellae led to significant increases in the amount of IL-1β and TNF-α within the tumor mass, particularly during the tumor suppression stage. Tumor colonization by ΔppGpp Salmonellae led to increased infiltration of the tumor by host immune cells, which in turn led to increased expression of IL-1β and TNF-α by DCs and macrophages. Taken together these results suggest that increased expression of IL-1β and TNF-α mediates a protective anti-cancer immune response via ΔppGpp Salmonellae.
ΔppGpp Salmonellae treatment led to a significant increase in tumor infiltration by leukocytes such as macrophages, DCs, neutrophils, CD8+ T cells, and B cells. Colonization and proliferation of ΔppGpp Salmonellae significantly increased the proportion of M1-like macrophages and reduced Treg numbers in tumor tissue during the suppression period. These changes were reversible; the numbers of M1-like macrophages and Treg cells were similar to those in MG1655-treated or control tumor during the re-growing stage. This implies that ΔppGpp Salmonellae suppress tumor growth by making intratumoral myeloid cells less suppressive and by enhancing the anti-tumor immune response. Indeed, a previous study showed that the anti-tumor effects of Salmonella depend on a functional MyD88-TLR pathway, and that Salmonella treatment significantly reduces the intratumoral myeloid cell population [41].
TNF-α is involved in apoptosis-induced tumor cell death [42-44]; thus its role in BCT has been extensively investigated [24, 25, 45]. The rapid increases of circulating TNF-α induced by bacterial infection may make tumor vessels hyper-permeable, resulting in rapid influx of blood into the tumor due to vascular disruption [24]. A previous study shows that administration of TNF-α with chemotherapy enhanced tumor-selective drug uptake during perfusion and revealed high response rates in soft tissue sarcoma patients [46]. Here, we found that co-administration of recombinant TNF-α prolonged the tumor-suppressive effects of ΔppGpp Salmonellae.
Here, we showed that tumor-infiltrating cells produced large amounts of IL-1β in response to i.v. injection of ΔppGpp Salmonellae, leading to suppressed tumor growth; however, IL-1β levels returned to baseline when the tumor started to re-grow. A recent study showed that certain types of tumor cell produce high levels of IL-1β, which recruits tumor-associated neutrophils (TANs) to the site (TANs play a crucial role in anti-tumor activity) [47]. In addition, we found that immune cells, in particular DCs, are the main source of IL-1β in colonized tumors. Several mechanisms are responsible for processing IL-1β, including the caspase-1-dependent pathway (monocytes [48], macrophages [48], and dendritic cells [49]) and the caspase-1-independent pathway (monocytes [50] and neutrophils [51]) [52]. Here, we examined the expression of molecules associated with the inflammasome. We found that expression of the core molecules involved in the inflammasome signaling pathway, such as IPAF, caspase-1, IL-18, and IL-1β were up-regulated in ΔppGpp Salmonellae-treated tumors. Several nod-like receptor (NLR)-family proteins, including IPAF and NLRP3, are thought to regulate caspase-1 activation [49]. Franchi et al. showed that activation of caspase-1 in response to Salmonellae invasion was triggered by cytosolic exposure to bacterial flagellin; caspase-1 activation was mediated via IPAF signaling and was independent of TLR signaling [53].
Although chronic exposure to IL-1β is thought to play a role in angiogenesis and invasion in some cancers [54-57], the cytokine was also successfully used to treat cancer [22, 49, 58-61]. For example, a previous study suggests that DC-derived IL-1β induces tumor cell death by priming CD8+ T lymphocytes or promoting the polarization of CD4+ T cells into Th1 cells, which is crucial for a therapeutic response [49]. Because chemotherapeutic agents are cytotoxic, tumor cells release 'danger signals', which then activate the inflammasome complex (NLRP3-ASC-Caspase-1) in DCs, thereby triggering the maturation of caspase-1 [62]. Hence, IL-1β is a potential treatment for cancer.
Notably, the data presented herein show that blocking IL-1β activity abrogated the anti-tumor effects of ΔppGpp Salmonellae. Thus, IL-1β has potential anti-cancer activity in the context of BCT. Treatment with ΔppGpp Salmonellae alone led to a transient tumor suppression (1 week to 10 days), and the tumors subsequently regrew. This may be because (i) there were too few bacteria in the tumor at the later stages, or (ii) immuno-evasive/tolerance mechanisms were activated to subvert the bacteria-mediated immune responses. Further studies are needed to confirm these hypotheses. Here, we showed that the period of tumor suppression was increased by combined treatment with recombinant IL-1β and Salmonellae, thereby confirming that IL-1β plays a crucial role in tumor suppression.
MG1655 (LPS-competent E. coli) failed to elicit an effective anti-tumor response, presumably because the responses it did elicit were irrelevant in this context. MG1655-treated tumors showed production of pro-IL-1β and pro-IL-18, but no significant expression of caspase-1, IL-1β, or IL-18. Thus, MG1655 may activate the TLR4 signaling pathway but not the inflammasome pathways required for IL-1β production. More detailed studies are underway to identify the mechanisms by which different bacterial strains activate the inflammasome pathway.
We also detected increased numbers of macrophages and DCs in the spleen and draining lymph nodes as well as in the colonized tumors. The number of CD8+ T cells was significantly increased in ΔppGpp Salmonellae-colonized tumors, even during the early suppression stage. It is speculated that macrophages and DCs recruited to the draining lymph nodes may activate T cells. Considering the short period of tumor suppression, it is hard to connect inhibited tumor growth with the adaptive response triggered by CD8+ T cells infiltrating tumors colonized by ΔppGpp Salmonellae. Further studies are needed to elucidate the role of the adaptive immune response in this context.
ΔppGpp Salmonellae is deficient in synthesis of ppGpp, the signal molecule absolutely required for expression of SPI1 genes as well as the rest of SPI genes [63] and thus is non-invasive. The reduction of tumor size was highly correlated with Salmonella persistence in tumor in suppression stage. In contrast, when the tumor re-grew again at re-growing stage, the number of tumor-colonizing bacteria decreased. The mechanism of immune response by ΔppGpp Salmonellae might be intriguing when compared with that of invasive tumor-colonizing Salmonellae such as A1R [64-67].
Conclusion
Our results strongly suggest that TNF-α (secreted by macrophages) and IL-1β (secreted by various innate immune cells, particularly DCs) are responsible for the anti-cancer effects of S. typhimurium. Unlike wild-type E. coli MG1655, attenuated ΔppGpp Salmonellae triggered effective IL-1β/TNF-α-mediated immune responses in the tumor, leading to transient tumor regression. The main pathway responsible for IL-1β processing was the inflammasome, which in turn activated caspase-1, followed by the cleavage and secretion of active IL-1β and IL-18. The current study is the first to show that IL-1β plays a pivotal role in Salmonellae-mediated cancer therapy and provides a new paradigm of combined immunotherapy involving the administration of attenuated S. typhimurium plus a strategy to increase IL-1β activity.
Supplementary Material
Figures S1-S6.
Abbreviations
BCT: bacteria-mediated cancer therapy; DC: dendritic cell; dpi: days post-inoculation; ICE: IL-1 converting enzyme or caspase-1; IPAF: ICE-protease-activating factor; i.v.: intravenous; i.t.: intratumoral; LPS: lipopolysaccharide; MDSC: myeloid-derived suppressor cell; M-MDSC: monocytic MDSC; PMN-MDSC: polymorphonuclear MDSC; M1: M1-polarized macrophage; M2: M2-polarized macrophage; NLR: nod-like receptor; qRT-PCR: quantitative RT-PCR; TANs: tumor-associated neutrophils; tLN: tumor-draining lymph node; Treg: regulatory T cell.
Acknowledgements
This work was supported by the Pioneer Research Center Program through the National Research Foundation of Korea funded by the Ministry of Science, ICT & Future Planning (2015M3C1A3056410), the Leading Foreign Research Institute Recruitment Program (2011-0030034) and from a research program by IBS-R005-G1-2015-a00. Y.H. was supported by the Pioneer Research Center Program “Bacteriobot” (2012-0001031).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Coley WB. Treatment of inoperable malignant tumors with the toxines of erysipelas and the bacillus prodigiosus. Am J Med Sci. 1894;108:50-66
2. Forbes NS. Engineering the perfect (bacterial) cancer therapy. Nat Rev Cancer. 2010;10:785-94
3. Barbé S, Van Mellaert L, Anné J. The use of clostridial spores for cancer treatment. J Appl Microbiol. 2006;101:571-8
4. Lemmon M, Van Zijl P, Fox M, Mauchline M, Giaccia A, Minton N. et al. Anaerobic bacteria as a gene delivery system that is controlled by the tumor microenvironment. Gene Ther. 1997;4:791-6
5. Liu S, Minton N, Giaccia A, Brown J. Anticancer efficacy of systemically delivered anaerobic bacteria as gene therapy vectors targeting tumor hypoxia/necrosis. Gene Ther. 2002;9:291-6
6. Nuyts S, Van Mellaert L, Theys J, Landuyt W, Lambin P, Anné J. Clostridium spores for tumor-specific drug delivery. Anticancer Drugs. 2002;13:115-25
7. Avogadri F, Martinoli C, Petrovska L, Chiodoni C, Transidico P, Bronte V. et al. Cancer immunotherapy based on killing of Salmonella-infected tumor cells. Cancer Res. 2005;65:3920-7
8. Loeffler M, Le'Negrate G, Krajewska M, Reed JC. Attenuated Salmonella engineered to produce human cytokine LIGHT inhibit tumor growth. Proc Natl Acad Sci U S A. 2007;104:12879-83
9. Nguyen VH, Kim H-S, Ha J-M, Hong Y, Choy HE, Min J-J. Genetically engineered Salmonella typhimurium as an imageable therapeutic probe for cancer. Cancer Res. 2010;70:18-23
10. Pawelek JM, Low KB, Bermudes D. Tumor-targeted Salmonella as a novel anticancer vector. Cancer Res. 1997;57:4537-44
11. Zhao M, Yang M, Li X-M, Jiang P, Baranov E, Li S. et al. Tumor-targeting bacterial therapy with amino acid auxotrophs of GFP-expressing Salmonella typhimurium. Proc Natl Acad Sci U S A. 2005;102:755-60
12. Yazawa K, Fujimori M, Nakamura T, Sasaki T, Amano J, Kano Y. et al. Bifidobacterium longum as a delivery system for gene therapy of chemically induced rat mammary tumors. Breast Cancer Res Treat. 2001;66:165-70
13. Min J-J, Kim H-J, Park JH, Moon S, Jeong JH, Hong Y-J. et al. Noninvasive real-time imaging of tumors and metastases using tumor-targeting light-emitting Escherichia coli. Mol Imaging Biol. 2008;10:54-61
14. Jiang S-N, Phan TX, Nam T-K, Nguyen VH, Kim H-S, Bom H-S. et al. Inhibition of tumor growth and metastasis by a combination of Escherichia coli-mediated cytolytic therapy and radiotherapy. Mol Ther. 2010;18:635-42
15. Yoshimura K, Jain A, Allen HE, Laird LS, Chia CY, Ravi S. et al. Selective targeting of antitumor immune responses with engineered live-attenuated Listeria monocytogenes. Cancer Res. 2006;66:1096-104
16. Jeong J-H, Kim K, Lim D, Jeong K, Hong Y, Nguyen VH. et al. Anti-tumoral effect of the mitochondrial target domain of Noxa delivered by an engineered Salmonella typhimurium. PloS One. 2014;9:e80050
17. Jiang S-N, Park S-H, Lee HJ, Zheng JH, Kim H-S, Bom H-S. et al. Engineering of bacteria for the visualization of targeted delivery of a cytolytic anticancer agent. Mol Ther. 2013
18. Kim K, Jeong JH, Lim D, Hong Y, Yun M, Min J-J. et al. A novel balanced-lethal host-vector system based on glmS. PloS One. 2013;8:e60511
19. Min J-J, Nguyen VH, Kim H-J, Hong Y, Choy HE. Quantitative bioluminescence imaging of tumor-targeting bacteria in living animals. Nat Protoc. 2008;3:629-36
20. Park SJ, Park S-H, Cho S, Kim D-M, Lee Y, Ko SY. et al. New paradigm for tumor theranostic methodology using bacteria-based microrobot. Sci Rep. 2013:3
21. Degré M. Cytokines and bacterial infections. Biotherapy. 1996;8:219-28
22. Reisser D, Pance A, Jeannin JF. Mechanisms of the antitumoral effect of lipid A. Bioessays. 2002;24:284-9
23. Weibel S, Stritzker J, Eck M, Goebel W, Szalay AA. Colonization of experimental murine breast tumours by Escherichia coli K-12 significantly alters the tumour microenvironment. Cell Microbiol. 2008;10:1235-48
24. Leschner S, Westphal K, Dietrich N, Viegas N, Jablonska J, Lyszkiewicz M. et al. Tumor invasion of Salmonella enterica serovar Typhimurium is accompanied by strong hemorrhage promoted by TNF-alpha. PLoS One. 2009;4:e6692
25. Theys J, Nuyts S, Landuyt W, Van Mellaert L, Dillen C, Böhringer M. et al. Stable Escherichia coli-Clostridium acetobutylicum shuttle vector for secretion of murine tumor necrosis factor alpha. Appl Environ Microbiol. 1999;65:4295-300
26. Hong E-H, Chang S-Y, Lee B-R, Pyun AR, Kim J-W, Kweon M-N. et al. Intratumoral injection of attenuated Salmonella vaccine can induce tumor microenvironmental shift from immune suppressive to immunogenic. Vaccine. 2013;31:1377-84
27. Bosslet K, Ruffmann R, Altevogt P, Schirrmacher V. A rapid method for the isolation of metastasizing tumour cells from internal organs with the help of isopycnic density-gradient centrifugation in Percoll. Br J Cancer. 1981;44:356-62
28. Liu F, Zhang L, Hoffman RM, Zhao M. Vessel destruction by tumor-targeting Salmonella typhimurium A1-R is enhanced by high tumor vascularity. Cell Cycle. 2010;9:4518-24
29. Coburn B, Grassl GA, Finlay B. Salmonella, the host and disease: a brief review. Immunol Cell Biol. 2006;85:112-8
30. Vendrell A, Gravisaco MJ, Pasetti MF, Croci M, Colombo L, Rodríguez C. et al. A novel Salmonella Typhi-based immunotherapy promotes tumor killing via an antitumor Th1-type cellular immune response and neutrophil activation in a mouse model of breast cancer. Vaccine. 2011;29:728-36
31. Wick MJ. Living in the danger zone: innate immunity to Salmonella. Curr Opin Microbiol. 2004;7:51-7
32. Hope C, Ollar SJ, Heninger E, Hebron E, Jensen JL, Kim J. et al. TPL2 kinase regulates the inflammatory milieu of the myeloma niche. Blood. 2014;123:3305-15
33. Youn J-I, Kumar V, Collazo M, Nefedova Y, Condamine T, Cheng P. et al. Epigenetic silencing of retinoblastoma gene regulates pathologic differentiation of myeloid cells in cancer. Nat Immunol. 2013;14:211-20
34. Wolf D, Wolf AM, Rumpold H, Fiegl H, Zeimet AG, Muller-Holzner E. et al. The expression of the regulatory T cell-specific forkhead box transcription factor FoxP3 is associated with poor prognosis in ovarian cancer. Clin Cancer Res. 2005;11:8326-31
35. Martin-Orozco N, Li Y, Wang Y, Liu S, Hwu P, Liu Y-J. et al. Melanoma cells express ICOS ligand to promote the activation and expansion of T-regulatory cells. Cancer Res. 2010;70:9581-90
36. Anz D, Mueller W, Golic M, Kunz WG, Rapp M, Koelzer VH. et al. CD103 is a hallmark of tumor-infiltrating regulatory T cells. Int J Cancer. 2011;129:2417-26
37. Yu HB, Finlay BB. The caspase-1 inflammasome: a pilot of innate immune responses. Cell Host Microbe. 2008;4:198-208
38. Männel D, Moore R, Mergenhagen S. Macrophages as a source of tumoricidal activity (tumor-necrotizing factor). Infect Immun. 1980;30:523-30
39. Ganai S, Arenas R, Forbes N. Tumour-targeted delivery of TRAIL using Salmonella typhimurium enhances breast cancer survival in mice. Br J Cancer. 2009;101:1683-91
40. Nuyts S, Van Mellaert L, Theys J, Landuyt W, Bosmans E, Anné J. et al. Radio-responsive recA promoter significantly increases TNFalpha production in recombinant clostridia after 2 Gy irradiation. Gene Ther. 2001;8:1197-201
41. Kaimala S, Mohamed Y, Nader N, Issac J, Elkord E, Chouaib S. et al. Salmonella-mediated tumor regression involves targeting of tumor myeloid suppressor cells causing a shift to M1-like phenotype and reduction in suppressive capacity. Cancer Immunol Immunother. 2014;63:587-99
42. Blankenstein T, Qin Z, Uberla K, Müller W, Rosen H, Volk H-D. et al. Tumor suppression after tumor cell-targeted tumor necrosis factor alpha gene transfer. J Exp Med. 1991;173:1047-52
43. Leong K, Karsan A. Signaling pathways mediated by tumor necrosis factor alpha. Histol Histopathol. 2000;15:1303-25
44. Sanlioglu AD, Aydin C, Bozcuk H, Terzioglu E, Sanlioglu S. Fundamental principals of tumor necrosis factor-alpha gene therapy approach and implications for patients with lung carcinoma. Lung Cancer. 2004;44:199-211
45. Asher A, Mulé JJ, Reichert CM, Shiloni E, Rosenberg SA. Studies on the anti-tumor efficacy of systemically administered recombinant tumor necrosis factor against several murine tumors in vivo. J Immunol. 1987;138:963-74
46. Verhoef C, de Wilt JH, Grünhagen DJ, van Geel AN, ten Hagen TL, Eggermont AM. Isolated limb perfusion with melphalan and TNF-alpha in the treatment of extremity sarcoma. Curr Treat Options Oncol. 2007;8:417-27
47. Chen LC, Wang LJ, Tsang NM, Ojcius DM, Chen CC, OuYang CN. et al. Tumour inflammasome-derived IL-1β recruits neutrophils and improves local recurrence-free survival in EBV-induced nasopharyngeal carcinoma. EMBO Mol Med. 2012;4:1276-93
48. Netea MG, Nold-Petry CA, Nold MF, Joosten LAB, Opitz B, van der Meer JHM. et al. Differential requirement for the activation of the inflammasome for processing and release of IL-1beta in monocytes and macrophages. Blood. 2009;113:2324-35
49. Ghiringhelli F, Apetoh L, Tesniere A, Aymeric L, Ma Y, Ortiz C. et al. Activation of the NLRP3 inflammasome in dendritic cells induces IL-1beta-dependent adaptive immunity against tumors. Nat Med. 2009;15:1170-8
50. Hazuda DJ, Strickler J, Kueppers F, Simon PL, Young PR. Processing of precursor interleukin 1 beta and inflammatory disease. J Biol Chem. 1990;265:6318-22
51. Guma M, Ronacher L, Liu-Bryan R, Takai S, Karin M, Corr M. Caspase 1-independent activation of interleukin-1beta in neutrophil-predominant inflammation. Arthritis Rheum. 2009;60:3642-50
52. van de Veerdonk FL, Netea MG, Dinarello CA, Joosten LA. Inflammasome activation and IL-1β and IL-18 processing during infection. Trends Immunol. 2011;32:110-6
53. Franchi L, Amer A, Body-Malapel M, Kanneganti T-D, Ozoren N, Jagirdar R. et al. Cytosolic flagellin requires Ipaf for activation of caspase-1 and interleukin 1beta in salmonella-infected macrophages. Nat Immunol. 2006;7:576-82
54. Krelin Y, Voronov E, Dotan S, Elkabets M, Reich E, Fogel M. et al. Interleukin-1beta-driven inflammation promotes the development and invasiveness of chemical carcinogen-induced tumors. Cancer Res. 2007;67:1062-71
55. Talmadge JE, Donkor M, Scholar E. Inflammatory cell infiltration of tumors: Jekyll or Hyde. Cancer Metastasis Rev. 2007;26:373-400
56. Voronov E, Shouval DS, Krelin Y, Cagnano E, Benharroch D, Iwakura Y. et al. IL-1 is required for tumor invasiveness and angiogenesis. Proc Natl Acad Sci U S A. 2003;100:2645-50
57. Yoo SY, Lee SY, Yoo NC. Cytokine expression and cancer detection. Med Sci Monit. 2009;15:RA49-RA56
58. Carmi Y, Rinott G, Dotan S, Elkabets M, Rider P, Voronov E. et al. Microenvironment-derived IL-1 and IL-17 interact in the control of lung metastasis. J Immunol. 2011;186:3462-71
59. Carrier MJ, Chatfield SN, Dougan G, Nowicka U, O'Callaghan D, Beesley JE. et al. Expression of human IL-1 beta in Salmonella typhimurium. A model system for the delivery of recombinant therapeutic proteins in vivo. J Immunol. 1992;148:1176-81
60. Friedlander RM, Gagliardini V, Rotello RJ, Yuan J. Functional role of interleukin 1 beta (IL-1 beta) in IL-1 beta-converting enzyme-mediated apoptosis. J Exp Med. 1996;184:717-24
61. Zitvogel L, Apetoh L, Ghiringhelli F, Kroemer G. Immunological aspects of cancer chemotherapy. Nat Rev Immunol. 2008;8:59-73
62. Aymeric L, Apetoh L, Ghiringhelli F, Tesniere A, Martins I, Kroemer G. et al. Tumor cell death and ATP release prime dendritic cells and efficient anticancer immunity. Cancer Res. 2010;70:855-8
63. Song M, Kim H-J, Kim EY, Shin M, Lee HC, Hong Y. et al. ppGpp-dependent stationary phase induction of genes on Salmonella pathogenicity island 1. J Biol Chem. 2004;279:34183-90
64. Zhao M, Yang M, Ma H, Li X, Tan X, Li S. et al. Targeted therapy with a Salmonella typhimurium leucine-arginine auxotroph cures orthotopic human breast tumors in nude mice. Cancer Res. 2006;66:7647-52
65. Zhao M, Geller J, Ma H, Yang M, Penman S, Hoffman RM. Monotherapy with a tumor-targeting mutant of Salmonella typhimurium cures orthotopic metastatic mouse models of human prostate cancer. Proc Natl Acad Sci U S A. 2007;104:10170-4
66. Yam C, Zhao M, Hayashi K, Ma H, Kishimoto H, McElroy M. et al. Monotherapy with a tumor-targeting mutant of S. typhimurium inhibits liver metastasis in a mouse model of pancreatic cancer. J Surg Res. 2010;164:248-55
67. Yano S, Zhang Y, Zhao M, Hiroshima Y, Miwa S, Uehara F. et al. Tumor-targeting Salmonella typhimurium A1-R decoys quiescent cancer cells to cycle as visualized by FUCCI imaging and become sensitive to chemotherapy. Cell Cycle. 2014;13:3958-63
Author contact
![]() Corresponding author: Jung-Joon Min, Laboratory of In Vivo Molecular Imaging, Department of Nuclear Medicine, Chonnam National University Hwasun Hospital, 160 Ilsimri, Hwasun, Jeonnam, Republic of Korea. Tel.: (+82) 10-622-06406; Fax: (+82) 61-379-8455; E-mail: jjminac.kr or Sin-Hyeog Im, Academy of Immunology and Microbiology (AIM), Institute for Basic Science (IBS), Pohang, Republic of Korea, and Division of Integrative Biosciences and Biotechnology (IBB), Pohang University of Science and Technology, Pohang, 790-784, Republic of Korea. Tel.: 82-54-279-2356; Fax: 82-54-279-8768; E-mail: iimshac.kr
Corresponding author: Jung-Joon Min, Laboratory of In Vivo Molecular Imaging, Department of Nuclear Medicine, Chonnam National University Hwasun Hospital, 160 Ilsimri, Hwasun, Jeonnam, Republic of Korea. Tel.: (+82) 10-622-06406; Fax: (+82) 61-379-8455; E-mail: jjminac.kr or Sin-Hyeog Im, Academy of Immunology and Microbiology (AIM), Institute for Basic Science (IBS), Pohang, Republic of Korea, and Division of Integrative Biosciences and Biotechnology (IBB), Pohang University of Science and Technology, Pohang, 790-784, Republic of Korea. Tel.: 82-54-279-2356; Fax: 82-54-279-8768; E-mail: iimshac.kr
 Global reach, higher impact
Global reach, higher impact