13.3
Impact Factor
Theranostics 2017; 7(13):3338-3353. doi:10.7150/thno.20512 This issue Cite
Research Paper
MicroRNA-647 Targets SRF-MYH9 Axis to Suppress Invasion and Metastasis of Gastric Cancer
1. Department of General Surgery, Nanfang Hospital, Southern Medical University, Guangdong Provincial Engineering Technology Research Center of Minimally Invasive Surgery, Guangzhou, 510515 China;
2. Guangdong Key Laboratory of Laboratory Animal, Guangdong Laboratory Animal Monitoring Institute, Guangzhou 510663, China;
3. Leder Human Biology and Translational Medicine, Biology and Biomedical Sciences, Division of Medical Sciences, Harvard Medical School, Boston, MA 02115.
* These authors contributed equally to this work
† Present address: Department of General Surgery, The First Affiliated Hospital of Xiamen University, Xiamen, 361003 China.
‡ Present address: Department of General Surgery, Panyu Central Hospital, Guangzhou, 511400 China
Received 2017-4-10; Accepted 2017-5-29; Published 2017-8-2
Abstract
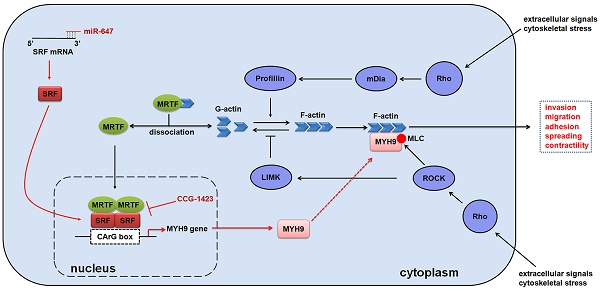
MicroRNAs (miRNAs) play important roles in regulating tumour development and progression. Here we show that miR-647 is repressed in gastric cancer (GC), and associated with GC metastasis. Moreover, we identify that miR-647 can suppress GC cell migration and invasion in vitro. Mechanistically, we confirm miR-647 directly binds to the 3' untranslated regions of SRF mRNA, and SRF binds to the CArG box located at the MYH9 promoter. CCG-1423, an inhibitor of RhoA/SRF-mediated gene transcription, inhibits the expression of MYH9, especially in SRF downregulated cells. Overexpression of miR-647 inhibits MGC 80-3 cells' metastasis in orthotropic GC models, but increasing SRF expression in these cells reverses this change. Importantly, we found the synergistic inhibition effect of CCG-1423 and agomir-647, an engineered miRNA mimic, on cancer metastasis in orthotropic GC models. Our study demonstrates that miR-647 functions as a tumor metastasis suppressor in GC by targeting SRF/MYH9 axis.
Keywords: Gastric cancer, Metastasis, MicroRNA-647, MYH9, SRF.
Introduction
Gastric cancer (GC) is the third leading cause of cancer-related death worldwide and causes more than 723,000 deaths every year [1]. Despite recent therapeutic advances, the mortality of GC remains high due to its silent nature, late clinical presentation and high levels of biological and genetic heterogeneity [2]. GC is a heterogeneous disease that evolves in various genetic and epigenetic alterations [2-9], including transcriptional changes of noncoding RNAs.
MicroRNAs (miRNAs), a class of noncoding RNAs, regulates protein expression through incomplete base pairing with the 3' untranslated regions (3'UTR) of target messenger RNAs (mRNAs) [10]. Accumulating evidences suggest that miRNAs play an important role in GC development and progression. Though certain miRNAs, such as miR-21 [11], miR-148a [12], miR-218 [13] and miR-29c [14], have been confirmed to act as oncogenes or tumor suppressor genes in GC, there are many other miRNAs suggested to be dysregulated by microRNA arrays, of which the detailed molecular mechanisms in GC are still not well elucidated. MicroRNA-647 (miR-647) has been reported to be a predictive biomarker for prostate cancer recurrence and a prognostic factor for Taxol-sensitive ovarian cancer patients [15, 16]. It was also identified as one of the 7 population-differentiated-miRNAs that are currently implicated as cancer biomarkers or diagnostics [17]. Although miR-647 has been reported to be dysregulated in GC [18, 19], the molecular mechanism of miR-647 in cancer metastasis was not fully uncovered.
Here we report that miR-647 expression is repressed in GC. In vitro and in vivo restorations of miR-647 via exogenous transfection are sufficient to inhibit both GC cell migration and invasion. SRF, a transcription factor known to promote tumor metastasis, and MYH9, a well-known cytoskeleton, are direct and indirect downstream targets of miR-647, respectively. MiR-647/SRF/MYH9 axis is also strongly correlated with GC metastasis in patients. Importantly, we demonstrate that agomir-647, an engineered miRNA mimic, and CCG-1423, an inhibitor of the Rho/MKL1/SRF signaling [20], synergistically inhibit cancer metastasis in orthotropic GC models. Taken together, our results provide an explanation of the metastatic mechanism of GC, and might be novel therapeutic targets or prognostic markers for GC.
Materials and Methods
Patients and Specimens
Two independent cohorts involving 199 GC patients were enrolled in this study. In cohort 1, fresh GC samples and adjacent noncancerous tissues were collected from 109 pathologically verified GC patients who underwent gastrectomy between November 2012 and November 2013 in Nanfang Hospital, Southern Medical University (Guangzhou, China). None of the patients received chemotherapy or radiotherapy before gastrectomy. The GC diagnosis was made by endoscopic biopsy combined with histopathological information after surgery. Diagnosis was confirmed by two pathologists in Nanfang hospital according to American Joint Committee on Cancer (AJCC) TNM Staging Classification for Carcinoma of the Stomach (7th ed., 2010) [21]. Most of cohort 1 patients were followed up for 3 years. This study was approved by the Ethics Committee of Nanfang hospital (Guangzhou, China). For cohort 2, a gastric adenocarcinoma tissue microarray (TMA, HStm-Ade180Sur-05; Shanghai Outdo Biotech) was obtained from the National Engineering Center For Biochip at Shanghai. The TMA was constructed with 90 paired formalin-fixed, paraffin-embedded gastric adenocarcinoma tissues and their corresponding adjacent normal tissues. Cohort 2 patients were followed up for 7.8 years. The corresponding clinicopathological information was listed in Table S1.
Cell Lines and Cell Culture
Human GC cell lines (MGC 80-3, MKN45, SNU-5, AGS and KATOIII) were purchased from the Cell Resource Center, Shanghai Institute of Biochemistry and Cell Biology at the Chinese Academy of Sciences. Cells were cultured in RPMI-1640 medium (MGC 80-3 and MKN45), F12 (AGS) or Dulbecco's Modified Eagle's Media (KATOIII and SNU-5) containing 10% fetal bovine serum (FBS) at 37°C in an atmosphere of 5% CO2.
In Situ Hybridization (ISH)
All the paraffin sections were examined with locked nucleic acid (LNA) based ISH using DIG-labeled miRCURY miRNA probes (Exiqon, Vedbaek, Denmark). The protocol for detection of miRNAs has been previously published [22].
Statistical Analysis
Statistical analysis was performed using GraphPad Prism software 5.0 (GraphPad Software, Inc., San Diego, CA, USA) and SPSS software (Version 21.0; Abbott Laboratories, Chicago, IL). Data were shown as mean ± standard error of mean unless otherwise noted. The Student t test was used to detect significance of data from qPCR experiments, migration and invasion assays, luciferase reporter assay, ChIP assay and in vivo metastatic nodules assay. Moreover, it was used to detect the relationship between miR-647 expression and clinicopathologic characteristics. Pathway enrichment analysis of miR-647 target genes was based on Fisher's exact test using a 2×2 contingency table. Mann-Whitney U test was used to analyze SRF protein expression between normal and GC tissues detected by IHC. The associations between miR-647 expression detected by ISH and proteins expression detected by IHC were analyzed by the χ2 test. Survival curves were calculated using Kaplan-Meier and log-rank tests. The effects of variables on survival were determined by univariate and multivariate Cox proportional hazards modeling. Correlation between SRF and MYH9 proteins in the TMA was analyzed by Spearman rank correlation, while correlation between SRF and MYH9 mRNA data extracted from TCGA was analyzed by Pearson correlation. A p value less than .05 was considered statistically significant.
All supplementary methods are available in supplementary material.
Results
MiR-647 is downregulated in GC tissues and associated with GC metastasis
To identify the expression of miR-647 in GC, we first detected its expression in 109 human GC tissues by qPCR. The result showed that miR-647 was frequently downregulated in GC tissues (T) compared to normal gastric mucosa (N) (p <0.0001, Figure 1A). Moreover, 73.4% (80 of 109) of the GC had at least 2-fold reduced expression of miR-647 compared with their corresponding nontumorous tissues (Figure 1B), which was supported by data from GSE36968 (Figure S1A). To determine whether miR-647 expression levels were related to GC progression, we analyzed the association between miR-647 and clinicopathologic status in 109 GC patients (Table S1-2). As shown in Figure 1C and Table S2, statistical analysis represented a strong correlation between miR-647 expression and TNM stage (p=0.0020), lymph-node metastasis (p=0.0046) and local invasion (p=0.0062). Collectively, these findings strongly suggested that miR-647 was downregulated in GC and negatively associated with GC metastasis.
To detect the function of miR-647 in GC cell invasion and migration, we first determined the expression of miR-647 in 5 normal human gastric tissues from 109 gastric cancer patients, 5 human gastric cancer cell lines (MGC 80-3, MKN45, SNU-5, AGS and KATOIII) and 3 human colorectal cancer cell lines (SW480, SW620 and Caco2) available in our lab by qPCR. Since miR-647 had been identified as one of the 7 miRNAs that were implicated as cancer biomarkers or diagnostics for multiple cancers, including colorectal cancer [17], the human colorectal cancer cell lines were used as positive controls. The results showed miR-647 expression was relatively lower in GC cell lines compared with normal tissues (Figure S2A). Since miR-647 expression was lowest in MGC 80-3 among 5 GC cell lines, followed by MKN45 (Figure 1D), and AGS had relatively higher miR-647 expression level and lentivirus transfection efficiency, we chose MGC 80-3, MKN45 and AGS to establish miR-647 stably overexpressing cell lines by lentivirus infection. Successful overexpression or knockdown of miR-647 was confirmed by qPCR (Figure 1E-F). Intriguingly, overexpression of miR-647 significantly suppressed the migratory and invasive abilities of MGC 80-3 and AGS cells (Figure 1G-H and S2D). Simultaneously, knockdown of miR-647 expression in these LV-miR-647 GC cells by transiently transfecting miR-647 inhibitors could restore the migration and invasion of GC cells (Figure 1G-H and S2D).
MiR-647, associated with Rho Signaling Pathways, negatively regulates SRF and MYH9 expression in GC cells
To identify how miR-647 was involved in GC metastasis, we firstly used 4 common miRNA target prediction engines (miRanda, TargetScan, miRDB, and miRWalk) to search for putative protein-coding gene targets of miR-647 (http://zmf.umm.uni-heidelberg.de/apps/zmf/mirwalk/micrornapredictedtarget.html), and 7724 potential targets were found (Figure 2A). Since miR-647 significantly inhibited GC migration and invasion, we selected 26 genes, listed out in Figure 2A and Table S3, associated with GC metastasis in PubMed. Additionally, we performed pathway enrichment analysis of miR-647 target genes predicted in the miRBase Targets database (http://microrna.sanger.ac.uk/targets) using miRGator online software (http://genome.ewha.ac.kr miRGator) (Table S4) [23, 24]. Specifically, the analysis showed that miR-647 was associated with Rho cell motility signaling pathway (Figure 2B, p=0.0134, BioCarta) and Role of MAL in Rho-mediated activation of SRF (Figure 2B, p=0.0348, BioCarta), which were critical pathways synergistically promoting cancer cell migration and invasion [25]. Because Rho-mediated pathways are closely associated with cancer metastasis and frequently activated in GC [7, 26], we selected out 5 candidate genes (SRF, ROCK1, ZAK, ELMO2, and MYH9) from Figure 2A (red marked) for further research, which were reported to be up-regulated in GC or associated with Rho-ROCK and Rho-MRTF/SRF pathways [12, 27-30].
Since MGC 80-3 and MKN45 expressed miR-647 at relatively low level, AGS and SNU-5 expressed miR-647 at relatively high level (Figure 1D), we transfected MGC 80-3 and MKN45 with miR-647 mimics, AGS and SNU-5 with miR-647 inhibitors (Figure S2E). Then, we carried out qPCR to detect these genes in transfected cells. The results revealed that the expression levels of SRF mRNA were most downregulated in MGC 80-3 and MKN45 cells treated with miR-647 mimics, followed by MYH9 and ZAK mRNAs (Figure 2C and S2F). Consistently, these three mRNAs were upregulated in AGS and SNU-5 cells treated with miR-647 inhibitors (Figure 2C and S2G). Because the fold changes of SRF and MYH9 mRNAs in miR-647-dysregulated GC cells are the highest, we focused on these two genes and carried out western blot and qPCR to confirm their relationship with miR-647. As showed in Figure 2D-E and S2B-C, the expression levels of SRF and MYH9 were both positively associated with miR-647 expression between GC cell lines and normal tissues. Moreover, we detected SRF and MYH9 proteins in GC cell lines stably overexpressing miR-647 (MGC 80-3-LV-miR-647, MKN45-LV-miR-647 and KATOIII-LV-miR-647) by western blot. The results showed SRF and MYH9 levels were significantly downregulated in the case of miR-647 overexpression (Figure 2F, S2H and S2J). In addition, when we transfected MGC 80-3-LV-miR-647, MKN45-LV-miR-647 and SNU-5 with miR-647 inhibitors, the expression of SRF and MYH9 proteins significantly restored (Figure 2G, S2I and S2J). Taken together, these results supported that miR-647 was negatively associated with SRF and MYH9 expression levels.
MiR-647 inhibits the migration and invasion of GC cell lines by directly targeting to SRF mRNA, and indirectly regulating MYH9 expression
We performed luciferase reporter assays to determine whether miR-647 directly interacts with the 3'UTR of SRF and MYH9 mRNAs. Using a computer-based sequence analysis (TargetScan and miRanda), we acquired potential miR-647-targeting sequences of SRF and MYH9. As shown in Figure 3A, three potential binding sites of miR-647 were located in 3'UTR of SRF, while one potential binding site in 3'UTR of MYH9. Then, we subcloned two 235-bp fragments of the SRF 3'UTR (Figure 3A; position 19-253 and 1152-1386, marked as WT1 and WT2) that included all the three predicted miR-647 recognition sites (Figure 3A; position 122-128, 150-155 and 1260-1271), and their corresponding mutant counterparts (Figure 3A; Mut1-3) into a luciferase reporter plasmid (pGV306). For testing whether SRF was a direct target of miR-647, MGC 80-3 and AGS cells were cotransfected with miR-647 mimics (or a negative control) and luciferase vectors containing wild type SRF-3'UTR (WT1 and WT2) (or luciferase vectors containing mutant-type SRF-3'UTR (Mut1-3)). Similarly, for determining whether miR-647 directly interacted with the MYH9 mRNA, we subcloned a 203-bp fragment of the MYH9 3'UTR (Figure 3A; position 732-934, marked as WT3) that included the predicted miR-647 recognition site (Figure 3A; position 832-837), and inserted it or its corresponding mutant counterpart (Mut4) into pGV306. Then, MGC 80-3 and AGS cells were cotransfected with miR-647 mimics (or a negative control) and luciferase vector containing wild type MYH9-3'UTR (WT3) (or luciferase vector containing mutant-type MYH9-3'UTR (Mut4)). As shown in Figure 3B, in cells transfected with WT SRF-3'UTR (WT1 or WT2) vectors and the miR-647 mimics, a significant decrease in luciferase activity was observed compared with cells cotransfected SRF-3'UTR vectors and negative controls. However, the luciferase activity of mutant-type SRF-3'UTR vectors (Mut1 and Mut3), excepting for Mut2, did not change following cotransfection with the miR-647 mimics, indicating miR-647 could directly interact with two sequences (GCAGCCA and UGAGUGCAGCCA) within SRF 3'UTR (Figure 3B). In GC cells (MGC 80-3 and AGS) transfected with WT MYH9-3'UTR (WT3) vector and the miR-647 mimics, no significant change in luciferase activity was observed compared with WT3 vector and negative controls (Figure 3C). And the luciferase activity of mutant-type MYH9-3'UTR vectors (Mut4) also did not change following cotransfection with the miR-647 mimics (Figure 3C). Collectively, these findings suggested miR-647 could directly target to the 3'UTR of SRF, but not the 3'UTR of MYH9.
To understand why MYH9 expression was indirectly associated with miR-647, we analyzed the 1-kb region directly upstream of MYH9 and predicted potential transcriptional factors (TFs) by using bioinformatics analysis websites (Proscan and JASPAR) (Figure 3D). We generated the full-length MYH9 promoter construct (luciferase plasmid construct) pGL3-MYH9-1300 (wild-type) and five truncation constructs (pGL3-MYH9-1000, pGL3-MYH9-903, pGL3-MYH9-753, pGL3-MYH9-528 and pGL3-MYH9-328) according to the locating sites of predicted TFs (Figure 3D). Intriguingly, dual-luciferase reporter assay showed that position -589 to -439, which potentially contained SRF binding site (CCATATATGG), was found to significantly increase the relative luciferase activity (Figure 3D). SRF was a ubiquitously expressed, pleiotropic transcription factor that activated genes by targeting to CArG box, which was a 10-bp CC (A+T-rich)6GG DNA sequence [31]. To further confirm this binding site, we constructed pGL3-MYH9-903-mut, which contained CArG box-mutant sequence (AACGACCAGAG). Luciferase reporter assay showed a significant decrease in reporter activity of MGC 80-3 cells cotransfected with pGV311-SRF and pGL3-MYH9-903-mut (Figure 3E), compared with MGC 80-3 cells cotransfected with pGV311-SRF and pGL3-MYH9-903. ChIP-qPCR analysis was further performed on MGC 80-3 and AGS cells. RNA polymerase II (RNAPII), which binds to the GAPDH promoter, was used as a positive control. The fold enrichment over the IgG control was represented. MYH9 promoter sequences were specifically enriched by anti-SRF and anti-MRTFA antibodies (a coactivator of SRF; Myocardin-related transcription factor A), but not by negative control antibody IgG in both MGC 80-3 and AGS cells (Figure 3F and Figure S3A). These data suggested that MYH9 was a direct target of transcriptional factor SRF in GC cells (Figure 3G). To detect whether SRF-MYH9 axis contributed to miR-647-associated GC metastasis, we transfected MGC 80-3/AGS-LV-miR-647 with SRF or MYH9 plasmids, followed by puromycin or G418 selections. Western blot analysis showed that upregulation of SRF in MGC 80-3/AGS-LV-miR-647 restored SRF and MYH9 expression, while overexpression of MYH9 in these cells only contributed to MYH9 upregulation (Figure 3H). Cell 3D migration and invasion assays suggested that the migratory and invasive abilities of MGC 80-3/AGS-LV-miR-647 restored after being transfected with either SRF or MYH9 plasmids (Figure 3I and S3B). Taken together, miR-647 could inhibit the migration and invasion of GC cell lines by targeting SRF/MYH9 signaling pathway.
MiR-647 is associated with poor prognosis and negatively related with SRF and MYH9 expression in GC patients
To determine whether there was an association between the expression levels of miR-647 and its target SRF in human GC tissues, we firstly analyzed the data from GEO database. Analysis of GSE36968 showed that miR-647 expression tended to be negatively associated with SRF mRNA (rs=-0.2982; p=0.2293; Figure S1B) and MYH9 mRNA (rs=-0.5273; p=0.0245; Figure S1B), although the correlation between miR-647 and SRF mRNA was not significant. To further verify the correlations, we detected the SRF and MYH9 expression of GC tissues and their matched normal mucosa in 52 GC patients from cohort 1 by qPCR. The results showed that both SRF and MYH9 mRNAs were significantly upregulated in GC tissues (Figure S1C), which was supported by GSE63288 (Figure S1D), and their expressions were negatively associated with miR-647 expression detected by qPCR and IHC (Figure S1E-F). Since our studies in vitro suggested miR-647 indirectly regulated MYH9 expression, the negative correlation between miR-647 and MYH9 mRNA could be attributed to dysregulation of SRF protein by miR-647. To confirm the relation between miR-647 expression and SRF protein levels, we further detected the SRF protein expression in 109 GC patients and their corresponding nontumorous tissues by IHC. The results showed that SRF protein was significantly upregulated in GC (Figure 4A and 4B), and high expression of SRF protein was more likely to be seen in GC with low levels of miR-647 (data from qPCR) (Figure 4C), suggesting that the upregulation of SRF protein mainly resulted from repression of miR-647 in GC. Furthermore, we detected the expression of miR-647 by ISH (Figure 4D). It was also found that miR-647 levels were negatively correlated with the levels of SRF and MYH9 protein (Figure 4E and 4F). Low miR-647 expression was significantly associated with higher histological grade (p=0.028), deeper local invasion (p=0.001), lymph-node metastasis (p=0.000) and advanced clinical stage (p=0.024) in Table S5. Kaplan-Meier analysis revealed that low-level expression of miR-647 was associated with poor overall survival in GC patients (p=0.024; Figure 4G). A multivariate Cox regression analysis indicated that low-level expression of miR-647 was an independent prognostic factor for predicting poor survival of GC (Table 1).
Activated SRF/MYH9 axis promotes GC metastasis and is associated with poor prognosis in GC
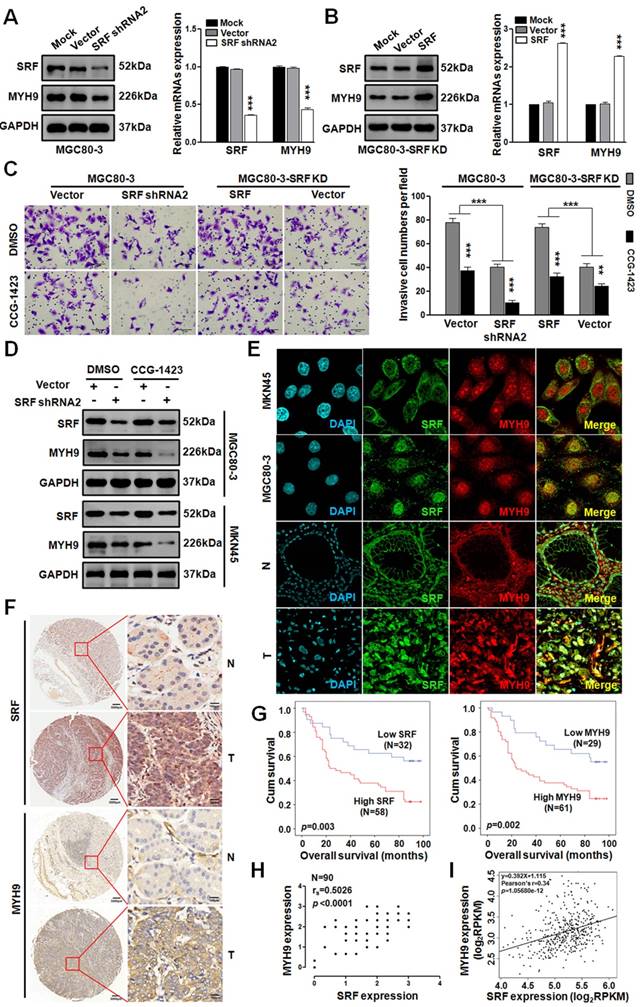
Since miR-647 could inhibit GC cell metastasis by SRF/MYH9 axis in vitro, we further detected the activity of this axis in GC. Firstly, we transfected MGC 80-3 and MKN45 cells with 3 SRF siRNAs and found SRF siRNA2 as the most effective sequence for knocking down SRF expression in GC cell lines by western blot (Figure S4). Then, we constructed a plasmid containing this sequence, marked as shRNA2, and transfected MGC 80-3 cells with it to obtain SRF knocked-down GC cells (MGC 80-3-SRF KD). Western blot and qPCR analysis showed that MYH9 expression in MGC 80-3-SRF KD significantly downregulated (Figure 5A). In addition, we further transfected MGC 80-3-SRF KD with a SRF plasmid, and found that both SRF and MYH9 expression were restored (Figure 5B), which further supported MYH9 was a downstream gene of SRF.
To test whether SRF/MYH9 axis regulated by Rho signaling pathway, we treated GC cells with CCG-1423, an inhibitor of RhoA/SRF-mediated gene transcription [32]. Firstly, we detected its IC50 in MGC 80-3 and AGS cells. The results showed that the IC50 of CCG-1423 in MGC 80-3 was 5.22 uM and in AGS was 5.08 μM (Figure S5A). In addition, before the function assays, GC cells (MGC 80-3 and AGS) were exposed to CCG-1423 with a linear concentration gradient, and 7.5uM CCG-1423 was found to be a better concentration due to its less toxic compared with 9uM in MGC 80-3 (Figure S5B). Intriguingly, we found that CCG-1423 effectively inhibited GC cells' invasion and MYH9 expression (Figure 5C-D and S5C), which suggested that SRF/MYH9 axis in GC was mediated by RhoA signaling pathway. In addition, the dysregulation of SRF expression by directly targeting to SRF gene could change GC cells' invasion. We also detected SRF and MYH9 expressions in GC cell lines (MKN45 and MGC 80-3) and GC tissues by confocal immunofluorescence microscopy. The results showed both proteins broadly located in cytoplasm and nucleus of GC cells (Figure 5E).
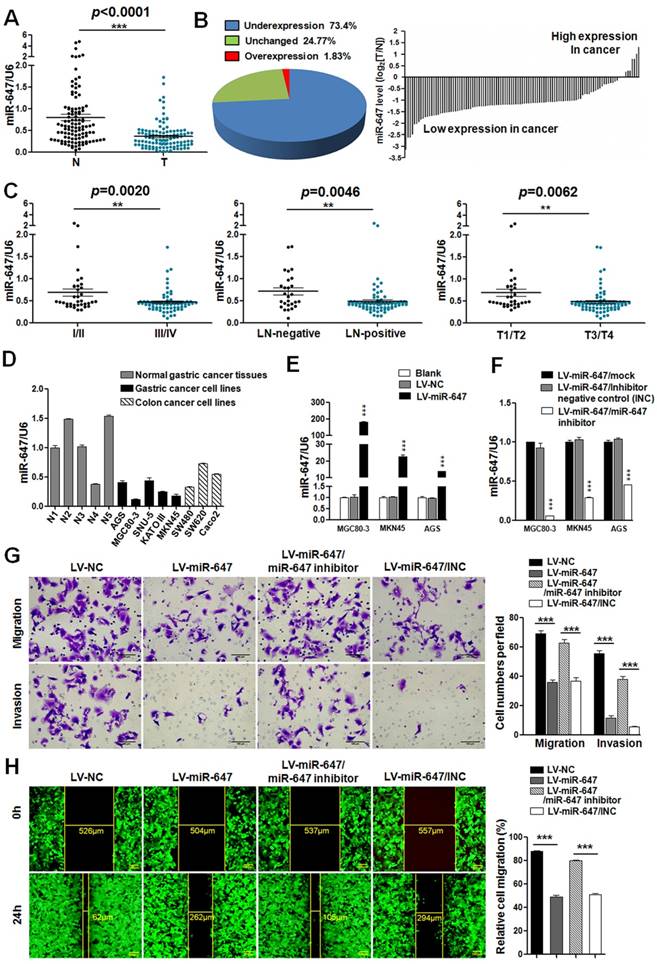
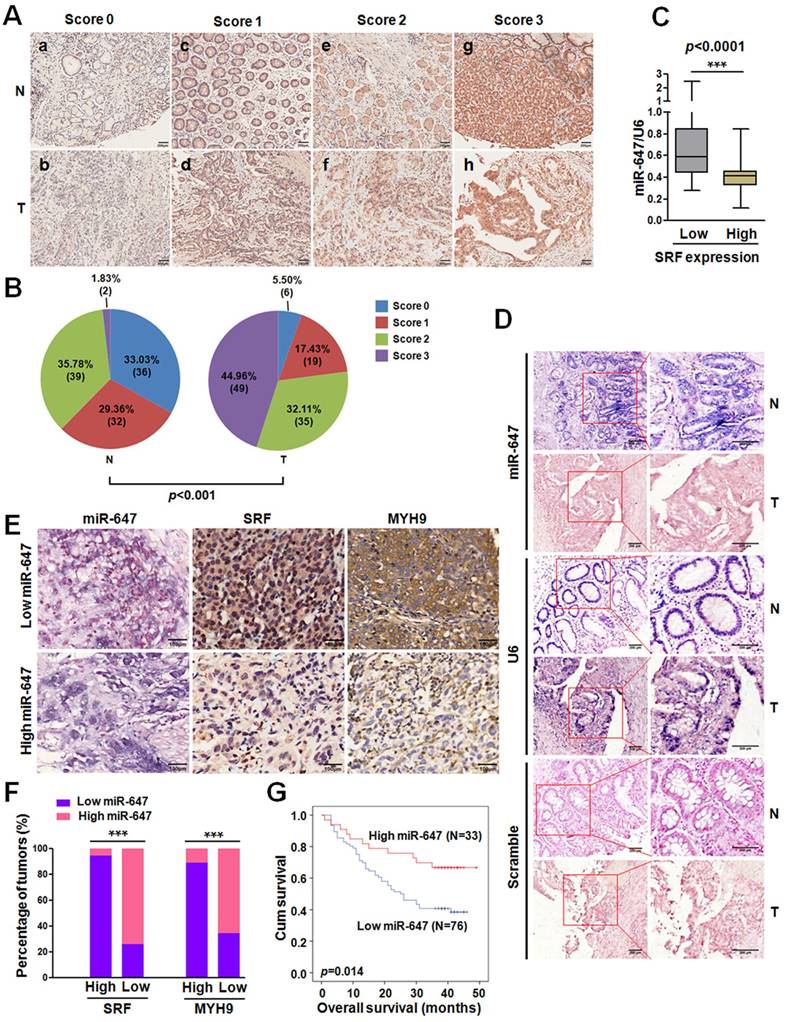
MiR-647 is downregulated in gastric cancer tissues and associated with the migration and invasion of gastric cancer cells. (A) Expression of miR-647 was determined by qPCR in 109 human gastric cancer tissues, which was normalized against an endogenous U6 RNA control. ***p<0.001. (B) Relative miR-647 levels of GC and normal tissues measured by qPCR were shown using pie chart and waterfall plot. The fold change of relative miR-647 expression (T/N) >2 or <1/2 was defined as significant. For waterfall plot, the fold change of relative miR-647 expression (log2[T/N]) >1 or <-1 was defined as significant. (C) The relationship between miR-647 expression and clinical stages, lymph node-metastasis or local invasion. *p<0.05; **p<0.01. (D) Expression of miR-647 in 5 normal human gastric mucosa tissues, 5 GC cell lines and 3 colorectal cancer cell lines was analyzed by qPCR. (E and F) Expression of miR-647 in transfected MGC 80-3, MKN45 and AGS cell lines were analyzed by qPCR. ***p<0.001. (G and H) The 3D migration and invasion changes of different MGC 80-3 cells were tested using transwell chamber migration assay and invasion assay. Cell 2D migration was tested using monolayer wound healing assay. ***p<0.001.
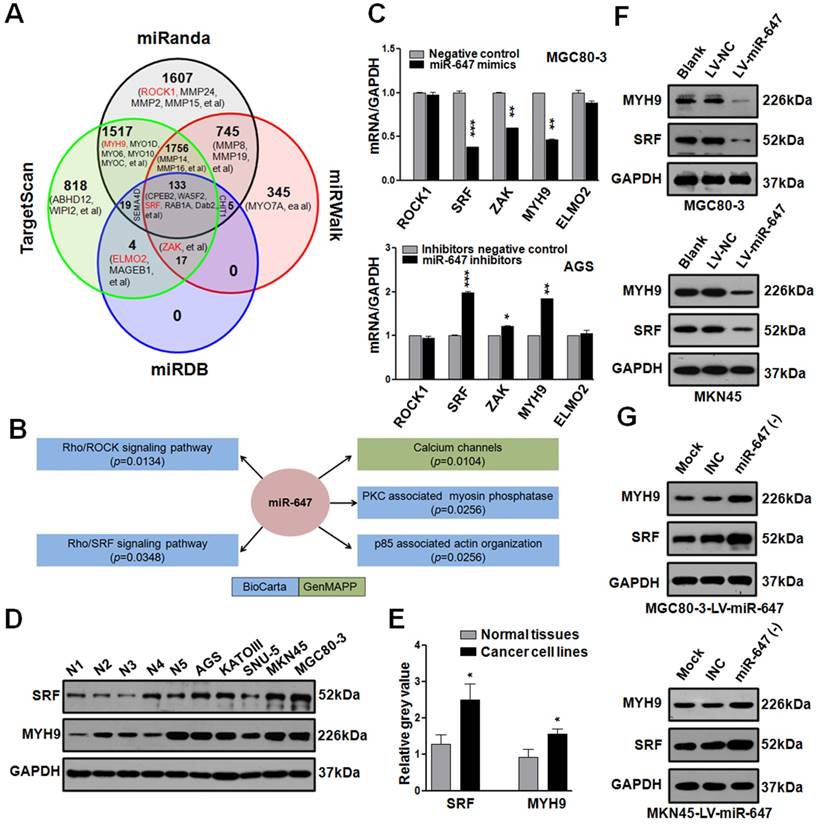
MiR-647 involves in Rho-associated signal pathway and negatively regulates SRF and MYH9 in gastric cancer cell lines. (A) Bioinformatics prediction of potential miR-647 targets by 4 common miRNA databases. Part of the genes associated with cell motility were showed and Rho-associated genes were marked as red. (B) Pathway enrichment analysis of miR-647 target genes predicted in the miRBase Targets database (http://microrna.sanger.ac.uk/targets). Analysis was performed using version 4 of the database and miRGator. Cell motility-associated signal pathways were listed. (C) Expression of Rho-associated genes were analyzed in MGC 80-3 and AGS by qPCR. U6 served as an internal control. ***p<0.001. (D-E) Expression of SRF and MYH9 proteins in 5 normal gastric mucosa tissues and 5 gastric cancer cell lines was analyzed by western blot and followed grayscale value analysis. GAPDH served as an internal control. (F) Western blot analysis was used to detect the expression level of SRF and MYH9 in MGC 80-3 and MKN45 cells after infection with miR-647 expressing or control lentivirus. GAPDH served as an internal control. (G) Western blot analysis was used to detect the expression changes of SRF and MYH9 in MGC 80-3-LV-miR-647 and MKN45-LV-miR-647 cells after infection with miR-647 inhibitors or its negative control (INC). GAPDH served as an internal control.
To further investigate the clinicopathological and prognostic significance of SRF and MYH9 in GC patients, we analyzed two GC cohorts by IHC, which included paired samples of normal gastric mucosa and GC tissues from 109 patients in our hospital (cohort 1) and National Engineering Center for Biochip in Shanghai (TMA; Figure 5F; cohort 2). High SRF expression was positively correlated with higher histological grade (p=0.031), deeper local invasion (p=0.016), lymph-node metastasis (p=0.000), distant metastasis (p=0.017) and advanced clinical stage (p=0.009) in cohort 1 (p<0.05; Table S6), and positively correlated with lymph-node metastasis (p=0.032) in cohort 2 (p<0.05; Table S7). Meanwhile, high MYH9 expression was positively related with lymph-node metastasis (p=0.000) and distant metastasis (p=0.042) in cohort 1 (p<0.05; Table S6), and positively related with lymph-node metastasis (p=0.040), advanced clinical stage (p=0.043) and tumor location at antrum or whole stomach (p=0.038) in cohort 2 (p<0.05; Table S7). Kaplan-Meier analysis showed that high levels of SRF and MYH9 expression were correlated with poor overall survival in both cohorts (Figure 5G and S6A-B). And miR-647, SRF and MYH9 expression levels were correlated with disease-free survival in cohort 1 (Figure S6C-E), respectively. Furthermore, multivariate Cox regression analysis revealed SRF and MYH9 expression to be an independent prognostic factor for poor survival (Table 1). Moreover, the expression of MYH9 protein was positively correlated with the expression of SRF protein (Figure 5H and S6F-G), which was further supported by analysis of these two mRNAs' expression data extracted from TCGA and GEO database (Figure 5I and S1Ga-c), and data detected by qPCR in 52 GC patients involved in cohort 1 (Figure S1Gd).
Ectopic expression of miR-647/SRF regulates tumor growth and metastasis by orthotropic implantation of GC in vivo
To further explore the role of miR-647 on tumor metastasis in vivo, MGC 80-3 cells stably expressed LV-NC/vector, LV-miR-647/vector and LV-miR-647/SRF were transplanted subcutaneously into nude mice, and the growth of resultant primary tumors then was monitored. After growth to an average size of 1 cm3, tumors were orthotopically sutured to the stomach of nude mice (Figure S7) [33]. Mice were sacrificed at the humane end point at two months after operation. The results showed metastatic lesions in the intestine, liver and abdominal wall (Figure 6A). Among these three groups, the tumor volumes of LV-miR-647/vector group were significantly smaller than the control group (LV-NC/vector group), and the tumor volumes of LV-miR-647/SRF group were relatively bigger than the LV-miR-647/vector group (Figure 6B). Additionally, in the group of mice bearing tumors of MGC 80-3-LV-NC/vector cells (control group), 100% (7 of 7), 85.7% (6 of 7) and 71.4% (5 of 7) of mice had intestinal, hepatic and abdominal wall metastases, respectively (Figure 6C). However, in the group of mice bearing tumors of MGC 80-3-LV-miR-647/vector cells, 75% (6 of 8), 25% (2 of 8) and 25% (2 of 8) of mice had intestinal, hepatic and abdominal wall metastases, respectively (Figure 6C). As expected, in the group of mice bearing tumors of MGC 80-3-LV-miR-647/SRF cells, 100% (8 of 8), 87.5% (7 of 8) and 75% (6 of 8) of mice had intestinal, hepatic and abdominal wall metastases, respectively (Figure 6C).
Lastly, the miR-647 expression in primary tumors of three groups was detected by qPCR, and the SRF and MYH9 expression in primary tumors and their metastatic nodules (4 intestinal metastases, 2 hepatic metastases and 2 abdominal wall metastases per group) were detected by IHC (Figure S8A-B and S9A-B). The qPCR data showed that LV-miR-647/vector group and LV-miR-647/SRF group had significantly higher levels of miR-647 expression, and there was no obviously different between these changes in vivo and the data from cell lines used for orthotropic implantation (Figure S8A). Meanwhile, the IHC data showed SRF and MYH9 expression was negatively correlated with miR-647 expression (Figure S8B). Although the SRF expression levels in randomly chosen metastatic nodules had no significantly difference between LV-miR-647/vector group and LV-miR-647/SRF group, there was expected tendency in two groups. Besides, all primary tumors and metastatic lesions were confirmed by HE staining (Figure S8G). These results further suggested that miR-647 inhibited tumor metastasis in vivo by downregulating SRF expression.
Agomir-647 and CCG-1423 synergistically inhibited GC metastasis in vivo
Since CCG-1423 (a SRF inhibitor) [32, 34] could downregulate MYH9 expression in our study, we carried out the CCG-1423 treatment study in orthotopic-transplant nude-mouse models of GC [33]. Two weeks after orthotopic-transplant surgery, the mice were treated with CCG-1423 (0.15 mg/kg/d, intraperitoneally) for 2 weeks [34]. Then, the same procedure was repeated at six weeks after surgery. The mice were sacrificed at 2 months after surgery. As shown in Figure 6D-F, the tumor volumes of CCG-1423-treated group were smaller than the control group, and the metastatic nodules were also significantly less in CCG-1423-treated group. The qPCR data showed that the CCG-1423-treated group and its control group had no significantly different levels of miR-647 expression, and this expression was not obviously different with the expression of cell lines used for orthotropic implantation (Figure S8C). Meanwhile, the IHC data showed SRF expression was no significantly different between two groups, but MYH9 expression was obviously downregulated in CCG-1423-treated group (Figure S8D and S9B).
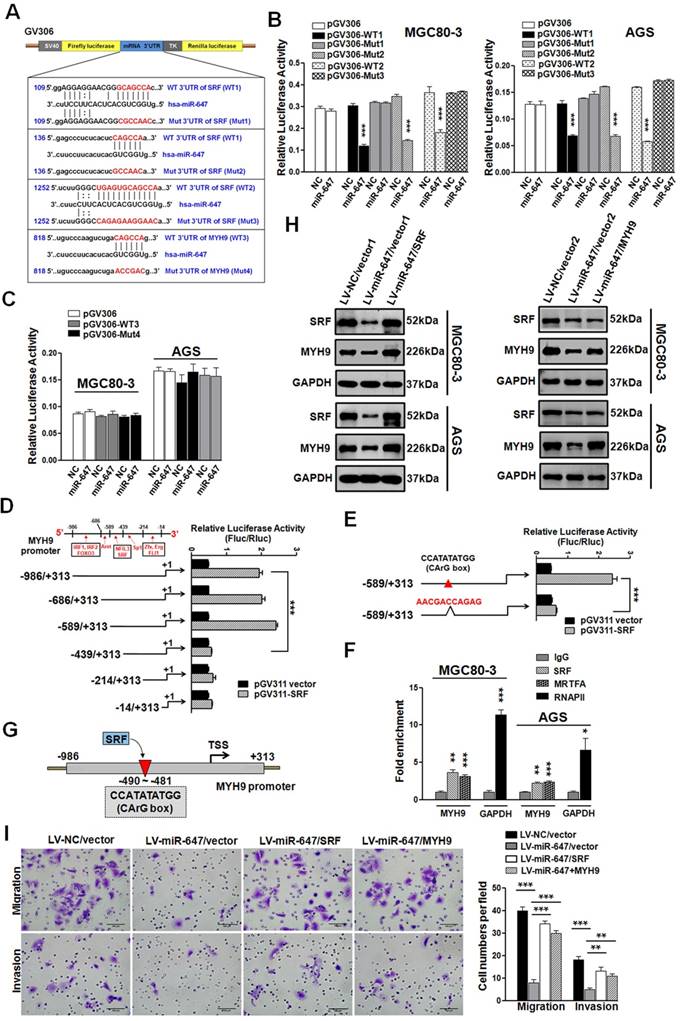
MiR-647 inhibits the migration and invasion of gastric cancer cell lines by directly targeting to SRF mRNA, which indirectly suppresses MYH9 expression. (A) Potential miR-647 binding sequences in the 3'UTR of SRF and MYH9 mRNAs predicted by TargetScan and miRanda databases. (B and C) Relative luciferase activity was analyzed after the described reporter plasmids (pGV306-3'UTR) or mock reporter plasmid (pGV306) were transfected into LV-miR-647 or LV-NC stably expressed MGC 80-3 and AGS (MGC 80-3-LV-miR-647 and AGS-LV-miR-647). The firefly luciferase activity was normalized to Renilla luciferase activity. ***p<0.001. (D to E) Dual-luciferase reporter assay was used to confirm SRF targeted to the promoter of MYH9 gene. ***p<0.001. (F) ChIP-qPCR analysis of SRF binding to the MYH9 promoter region in MGC 80-3 and AGS cells. RNA polymerase II (RNAPII) was used as a positive control. The fold enrichment over the IgG control is represented (mean ± SD; *p<0.05; **p<0.01; ***p<0.001). (G) Illustration of SRF protein targeting to the CArG box of MYH9 promoter. (H) Western blot analysis was used to detect the expression changes of SRF and MYH9 proteins followed by transfecting miR-647-overexpressed gastric cancer cell lines with SRF plasmid (SRF) or MYH9 plasmid (MYH9). GAPDH served as an internal control. (I) Cell 3D migration and invasion were tested using transwell chamber migration assay and invasion assay. AGS-LV-miR-647 transfected with SRF plasmid (SRF), MYH9 plasmid (MYH9) and vectors were used. Each bar represents the mean ± SD. ***p<0.001.
Expression of miR-647 is inversely correlated with the expression levels of SRF and MYH9 in gastric cancer samples. (A) Immunohistochemical staining of SRF protein in 109 paired of gastric cancer and their corresponding nontumorous tissues. Original magnification, 100×. (B) Statistical analysis of SRF expression according to the scoring (chi-square test; ***p<0.001.). (C) Correlation between SRF expression and miR-647 levels in the 109 gastric cancer tissue samples. ***p<0.001. (D) miR-647 expression patterns in human gastric cancer tissues and normal adjacent epithelial tissues as determined by in situ hybridization. Original magnification, 100× and 200×. DIG labeled LNA probes for U6 and scramble were used as positive and negative controls, respectively. (E and F) Expression associations of miR-647 with SRF and MYH9 expression in 109 GC patients. Two representative cases are shown (E). Original magnification, 200×. The percentage of specimens showing low or high miR-647 expression in relation to the expression levels of SRF and MYH9 (F); *p<0.05. (G) Kaplan-Meier survival analysis of miR-647 expression in patients with gastric cancer (log-rank test).
Univariate and Multivariate Analysis of Different Prognostic Parameters for GC Patients
| Cohort 1 | Cohort 2 | ||||
|---|---|---|---|---|---|
| Variables | HR (95% CI)a | p Valueb | HR (95% CI)a | p Valueb | |
| Univariate Analysis | |||||
| Gender (male vs female) | 1.120 (0.654-1.919) | 0.680 | 1.034 (0.603-1.773) | 0.903 | |
| Age (years) (<59 vs ≥59) | 1.073 (0.629-1.830) | 0.795 | 1.211 (0.706-2.077) | 0.486 | |
| Tumor size (cm) (<5.5 vs ≥5.5) | 1.247 (0.732-2.126) | 0.417 | 1.616 (0.968-2.699) | 0.067 | |
| Location (C/B vs A/W)c | 0.909 (0.528-1.567) | 0.733 | 0.637 (0.377-1.077) | 0.092 | |
| Histological grade (W/M vs P/U)d | 0.526 (0.310-0.891) | 0.017 | 2.465 (1.367-4.447) | 0.003 | |
| pT status (T1/T2 vs T3/T4) | 1.172 (0.620-2.215) | 0.626 | 3.726 (1.347-10.308) | 0.011 | |
| pN status (N0 vs N1-3) | 2.516 (1.190-5.320) | 0.016 | 2.199 (1.139-4.249) | 0.019 | |
| pM status (M0 vs M1) | 2.101 (1.086-4.065) | 0.027 | 2.829 (1.375-5.822) | 0.005 | |
| AJCC stage (I/II vs III/IV) | 2.057 (1.106-3.824) | 0.023 | 3.462 (1.936-6.191) | 0.000 | |
| miR-647 expression (low vs high) | 0.451 (0.233-0.871) | 0.018 | |||
| SRF expression (low vs high) | 2.090 (1.141-3.826) | 0.017 | 2.426 (1.327-4.435) | 0.004 | |
| MYH9 expression (low vs high) | 1.972 (1.077-3.608) | 0.028 | 2.497 (1.345-4.634) | 0.004 | |
| Multivariate Analysis I | |||||
| Histological grade (W/M vs P/U)d | 0.450 (0.263-0.772) | 0.004 | |||
| AJCC stage (I/II vs III/IV) | 1.883 (1.009-3.514) | 0.047 | |||
| miR-647 expression (low vs high) | 0.388 (0.198-0.762) | 0.006 | |||
| Multivariate Analysis II | |||||
| Histological grade (W/M vs P/U)d | 0.453 (0.245-0.728) | 0.004 | 1.915 (1.054-3.479) | 0.033 | |
| AJCC stage (I/II vs III/IV) | 1.788 (0.956-3.344) | 0.069 | 2.881 (1.597-5.198) | 0.000 | |
| SRF expression (low vs high) | 2.321 (1.245-4.327) | 0.008 | 1.922 (1.046-3.533) | 0.035 | |
| Multivariate Analysis III | |||||
| Histological grade (W/M vs P/U)d | 0.517 (0.304-0.877) | 0.014 | 1.940 (1.065-3.535) | 0.030 | |
| AJCC stage (I/II vs III/IV) | 1.831 (0.979-3.422) | 0.058 | 2.886 (1.596-5.220) | 0.000 | |
| MYH9 expression (low vs high) | 1.910 (1.037-3.518) | 0.038 | 2.120 (1.136-3.956) | 0.018 | |
aHazard ratios (HRs) and 95% confidence intervals (CIs) were calculated using univariate or multivariate Cox proportional hazards regression in SPSS.
bp values were calculated using univariate or multivariate Cox proportional hazards regression in SPSS. p values < 0.05 were considered to indicate statistical significance
cC, cardia; B, body; A, antrum; W, whole
dW, well; M, moderate; P, poor; U, undifferentiated
Agomir, an engineered miRNA mimic, is well confirmed to be effective in mice diseases models [35-37]. Since CCG-1423 and miR-647 inhibited GC migration and invasion by different targets of Rho/SRF/MYH9 axis (Figure S10), we further detected the synergistic inhibitory effect of CCG-1423 and agomir-647 on GC metastasis. The detailed procedure for constructing orthotopic-transplant nude-mouse models and drug administration were shown in Figure S7. As shown in Figure 6G-I, compared with DMSO-NC, CCG-1423-NC, DMSO-agomir-647 groups, the primary tumors volumes and numbers of metastatic nodules in combined treatment group (CCG-1423, 0.15 mg/kg/d, intraperitoneally; agomir-647 80mg/kg/3d, tail vein injection) were smaller and less, respectively (Figure 6G-I). Meanwhile, the qPCR data showed that miR-647 expression in DMSO-NC and CCG-1423-NC groups was no obviously different with its expression in cell lines used for orthotropic implantation, and miR-647 expression in primary tumors in DMSO-agomir-647 and CCG-1423-agomir-647 groups were significantly upregulated compared with its expression in DMSO-NC and CCG-1423-NC groups (Figure S8E), respectively. We also detected SRF and MYH9 protein expression in primary tumors and metastatic nodules. Firstly, SRF expression in CCG-1423-treated groups (CCG-1423-NC and CCG-1423-agomir-647 groups) was not significantly changed compared with their own controls (DMSO-NC and DMSO- agomir-647 groups), but SRF expression in agomir-647-treated groups (DMSO-agomir-647 and CCG-1423-agomir-647 groups) was significantly downregulated compared with their own controls (DMSO-NC and CCG-1423-NC groups) (Figure S8F and S9B). In addition, MYH9 expression in agomir-647-treated or CCG-1423-treated groups was downregulated compared with their respective controls. MYH9 expression was not significantly different between CCG-1423-NC and CCG-1423-agomir-647 groups. To be sure, all primary tumor, intestinal metastasis, hepatic metastasis and peritoneal metastasis were confirmed by HE staining before qPCR and IHC studies (Figure S8G). The expression levels of SRF and MYH9 were evaluated by IHC staining (Figure S8D, S8F and S9B).
SRF/MYH9 signaling increases gastric cancer cell lines invasion and its activity is associated with poor survival in gastric cancer patients. (A) Western blot and qPCR analyses were used to detect the expression changes of SRF and MYH9 in MGC 80-3 cells infected with SRF siRNA2 plasmid or vector control, which were selected with neomycin. (B) Western blot and qPCR analysis were used to detect SRF and MYH9 expression in the SRF knocked down MGC 80-3 cells (MGC 80-3-SRF-KD) infected with SRF plasmid or control. ***p<0.001. (C) Cell 3D invasion was tested using transwell chamber invasion assay (Matrigel-coated membrane). MGC 80-3 transfected with SRF siRNA2 plasmid, SRF plasmid and their corresponding vector controls were detected. CCG-1423 (7.5μmol/L) and its control Dimethyl sulfoxide (DMSO) were used to detect their influences on gastric cancer cell invasion. ***p<0.001; **p<0.01. (D) Western blot and qPCR analysis were used to detect the expression changes of SRF and MYH9 in the above MGC 80-3 cells. (E) Subcellular localization of SRF and MYH9 proteins in gastric cancer cell lines (MGC 80-3 and MKN45) and gastric cancer tissues were determined by confocal immunofluorescence microscopy. (F to I) The coordinate expression of SRF and MYH9 in GC patients. TMA analysis of SRF and MYH9 in primary human GC tissues (n=90) and adjacent normal tissues (F). Original magnification, 35× and 200×. Kaplan-Meier survival analysis of SRF and MYH9 expression (G) in patients with gastric cancer (log-rank test) were performed. Spearman rank correlation analysis was performed based on the IHC expression score of SRF and MYH9 from the TMA (H). Linear regression analysis of SRF and MYH9 was performed based on the RNA-seq data from TCGA (I).
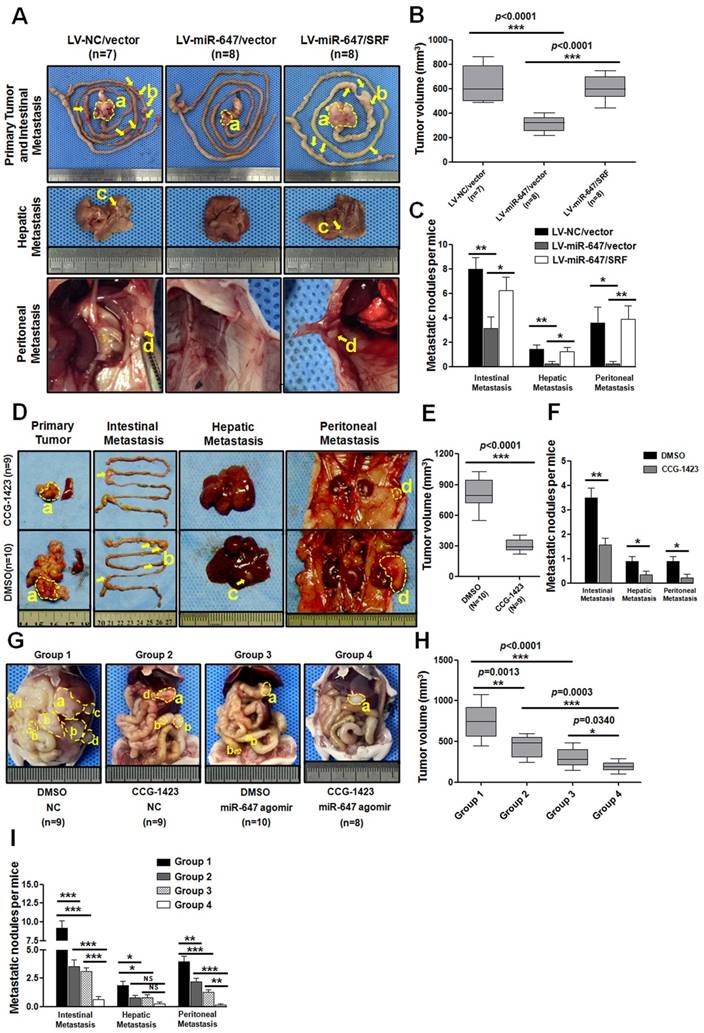
Agomir-647 and CCG-1423 synergistically inhibit tumor growth and metastasis of gastric cancer in vivo. (A-C) In vivo gastric cancer tumourigenesis analysis in the xenograft nude mouse models using gastric cancer cells (LV-NC/vector, LV-miR-647/vector and LV-miR-647/SRF). (A) White-light images of orthotopic tumors, intestinal, hepatic and peritoneal metastases of mice. Yellow arrows indicated metastatic nodules. Primary tumor volumes (B) and numbers of metastatic intestinal, hepatic and peritoneal nodules (C) per group were analyzed. (D-F) CCG-1423 (0.15 mg/kg/d, intraperitoneally) or vehicle alone (DMSO) was used to treat orthotopic-transplant nude-mouse for 4 weeks (2 procedures). Primary tumor volumes (E) and numbers of metastatic intestinal, hepatic and peritoneal nodules (F) per group were analyzed. (G-I) CCG-1423 (2 weeks per procedure, 2 procedures in all) and miR-647 agomir (80mg/kg/3d, tail vein injection, 5 times per procedure, 2 procedures) were combined to treat orthotopic-transplant nude-mice. Primary tumor volumes (H) and numbers of metastatic intestinal, hepatic and peritoneal nodules (I) per group were analyzed. *p<0.05, **p<0.01, ***p<0.001. a, primary tumor; b, intestinal metastatic nodules; c, hepatic metastatic nodules; d, peritoneal metastatic nodules.
Discussion
Though microRNAs' roles in cancer have been extensively studied, the significance and specific mechanism of miR-647 in cancer remains unknown. MiR-647 has been confirmed to be upregulated in Taxol-resistant ovarian cancer cells, and high expression of miR-647 was significantly associated with longer survival in chemosensitive ovarian cancer patients [16]. In prostate cancer, miR-647 has been reported to be one of the three genes in the predictive biomarker panel negatively associated with recurrence [15]. Besides, miR-647 is one of the 7 globally population-differentiated miRNAs associated with 37 diseases including GC, though the function of miR-647 is still unknown [17]. Meanwhile, relevant studies seemed to have different results. For example, the expression of miR-647 was found to be no difference between localized stage I non-small cell lung cancer patients with recurrence and those without recurrence [23]. Even due to its stable expression patterns across cases as well as the two case groups, miR-647 was suggested to be a potential reference miRNA for future stage I non-small cell lung cancer studies. Therefore, different cancers, even different stages, may have different miRNA expression profiles. Our study demonstrates that miR-647 is frequently downregulated in human GC, and functions as a tumor suppressor via SRF/MYH9 axis in GC.
In this study, we first performed qPCR in human GC tissues and corresponding normal gastric mucosa to identify miR-647 was associated with GC metastasis. Thereafter, using a series of in vitro and in vivo assays, we uncovered that miR-647 acted as an important invasion and metastasis suppressor in GC. Although miR-647 expression has been reported previously in other human cancers [15-17, 19, 23], ours is the first study that provides a novel and comprehensive insight into the functional role of miR-647 in GC metastasis and Rho-mediated signal pathways (Figure S10).
We confirmed that miR-647 was frequently downregulated in GC and significantly associated with local invasion, advanced clinical stage and lymph node-metastasis. After validating miR-647 acts as a metastasis-associated miRNA in vitro and in vivo, we performed a bioinformatics search for potential mRNAs targeted by miR-647 and carried out pathway enrichment analysis for miR-647-associated signal pathways and finally focused on Rho-mediated signal pathways due to its consentaneous function in cancer metastasis. Intriguingly, by qPCR and Luciferase reporter assays, SRF and MYH9, which are involved in Rho-mediated signal pathways and predicted to be potential miR-647 targets, were confirmed to be direct and indirect downstream targets of miR-647 in GC cells, respectively.
We found miR-647 directly targeted SRF mRNA and thus regulated Rho-mediated signaling pathways in GC cells. Rho signaling is strongly implicated in cancer metastasis [38], and this is generally thought to reflect direct effects on the cytoskeleton [39]. In addition to the direct cytoplasmic role of Rho effectors in cytoskeletal regulation, Rho GTPases also control activity of MAL/MRTF-A and MKL2/MRTF-B, which are transcriptional co-activators of SRF [40, 41]. SRF is an important regulator of genes through binding to the sequence CC(A/T)6GG termed the CArG box [42], which is involved in immediate early and tissue-specific gene expression, proliferation, differentiation, apoptotic and cancer metastasis [25, 28, 42]. Recently, SRF has been reported to be upregulated in GC and modulate the epithelial to mesenchymal transition by regulating miR-199a-5p expression [28]. However, little is known about the mechanism why SRF is overexpressed in GC tissues. In this study, we found miR-647, which was significantly downregulated in GC, could lead to the loss of inhibition of SRF via 3'UTR of SRF mRNA. The expression of miR-647 is also negatively associated SRF levels in GC tissue samples. In addition, our study also confirmed that MYH9 was a directly target of SRF in GC cells. MYH9, known as non-muscle myosin heavy chain IIA, was a known effector of Rho-ROCK signaling pathway and frequently found to be upregulated in GC [27, 43]. In our study, forced change of miR-647 expression not only led to SRF but also MYH9 dysregulation. Overexpression of MYH9 in LV-miR-647-infected GC cells could increase cells' migration and invasion. Thus, MYH9 was a critical downstream molecule of SRF in GC cells.
Currently, it is urgent to uncover a treatment strategy for GC metastasis, especially peritoneal metastasis. Our in vitro results showed that miR-647/SRF/MYH9 axis could effectively inhibit GC metastasis. MicroRNAs are thought to be promising next generation therapeutic targets in human diseases [44, 45]. Moreover, agomir is an engineered miRNA mimic, and has been well confirmed to be effective in mice diseases models [35-37]. CCG-1423, an inhibitor of Rho/MKL1/SRF signaling, has been used to improve glucose uptake and tolerance in insulin-resistant mice [34]. We further used the combined treatment strategy of agomiR-647 and CCG-1423 in vivo, which suppressed GC invasion and migration in different targets of miR-647/SRF/MYH9 axis. The proliferation and metastasis of orthotopic-transplant nude-mouse models of GC were dramatically inhibited. This synergistically inhibitory effect in vivo strengthened the potential clinically significance of miR-647/SRF/MYH9 axis in GC metastasis.
Our study has a few caveats. Since one miRNA often targets to many genes to regulate cell behavior, other molecules might also contribute to the invasion and metastasis suppressing effect of miR-647. ZAK, one of the predicted genes, was a factor of aggressiveness in cancer cells [46]. Our study found that the expression of ZAK mRNA was not significantly upregulated in AGS cells treated with miR-647 inhibitor, though its significant upregulation in MGC 80-3 cells treated with miR-647 mimics. Moreover, though miR-647 is negatively correlated with overall survival of GC patients, the potential clinical application of miR-647 remains a subject for further study. Whether it can be detected in patients' body fluids and serve as a diagnostic indicator are still unknown.
In conclusion, miR-647 is a potent tumor metastasis suppressor in the GC cells, and its migration and invasion inhibitory effects are, in part, mediated through its downstream target gene, SRF. MiR-647 may have a therapeutic potential to suppress GC metastasis by inhibiting Rho associated SRF/MYH9 signaling pathway.
Abbreviation
GC: gastric cancer; miRNAs: microRNAs; mRNA: messenger RNA; HR: hazard ratio; CI: confidence interval; AJCC: American Joint Committee on Cancer; TNM stage: tumor-node-metastasis stage; T: human gastric cancer tissue; N: normal gastric mucosa; LN: lymph node; SRF: serum response factor; MYH9: myosin heavy chain 9; NMHC IIA: non-muscle myosin heavy chain IIA; LV: lentivirus; NC: negative control; INC: inhibitor negative control; qRT-PCR(qPCR): quantitative real-time polymerase chain reaction; IHC: immunohistochemistry; WB: western blot; IF: immunofluorescence; DAPI: 4',6-diamidino-2-phenylindole; 3'UTR: three prime untranslated region; FLUC: Firefly luciferase; RLUC: Renilla luciferase; CArG box: 10-bp CC(A+T-rich)6GG sequence; MRTF-A: Myocardin-related transcription factor A; HE: hematoxylin-eosin staining; MGC 80-3/AGS-SRF-KD: the SRF knocked-down MGC 80-3/AGS cells; DMSO: Dimethyl sulfoxide; TMA: tissue microarray; GEO: Gene Expression Omnibus; TCGA: The Cancer Genome Atlas; RNAPII: RNA polymerase II
Acknowledgements
The authors thank Side Liu and Jide Wang (Southern Medical University, Guangzhou, China) for their technical assistance. The authors also thank James Y Yang (Xiamen University, xiamen, China) for his valuable guidance on this research.
Author contributions
This study was designed and supervised by L.G.X, L.H. and Q.X.L. Experiments were conducted by Y.G.T., Y.J., H.K.Z., Z.L.Y. and J.Y.M. Data analysis were conducted by Z.X.J. and Y.Q.B. Samples were obtained by S.Z.Y., Y.L, Z.H.P.The material and technical supports were obtained from L.W.D., Z.B.X., W.D. and Z.L. Funding was obtained by L.G.X. The manuscript was written by Y.G.T. and Q.X.L. The manuscript was revised by Q.X.L., H.Y.F., D.H.J and L.H. All authors discussed the results and commented on the manuscript.
Financial support
This work was supported by the grants from National Natural Science Foundation of China (81672446; 81600510), Guangdong Provincial Science and Technology Key Project (2014A020215014), the Research Fund of Public Welfare in the Health Industry, the National Health and Family Planning Commission of China (201402015) and the Key Clinical Specialty Discipline Construction Program, Guangzhou Industry-Academia-Research Collaborative Innovation Major Project (201704020015).
Supplementary Material
Supplementary figures and tables.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65:87-108
2. Tan P, Yeoh KG. Genetics and Molecular Pathogenesis of Gastric Adenocarcinoma. Gastroenterology. 2015;149:1153-62 e3
3. Wang K, Kan J, Yuen ST, Shi ST, Chu KM, Law S. et al. Exome sequencing identifies frequent mutation of ARID1A in molecular subtypes of gastric cancer. Nat Genet. 2011;43:1219-23
4. Zang ZJ, Cutcutache I, Poon SL, Zhang SL, McPherson JR, Tao J. et al. Exome sequencing of gastric adenocarcinoma identifies recurrent somatic mutations in cell adhesion and chromatin remodeling genes. Nat Genet. 2012;44:570-4
5. Yao F, Kausalya JP, Sia YY, Teo AS, Lee WH, Ong AG. et al. Recurrent Fusion Genes in Gastric Cancer: CLDN18-ARHGAP26 Induces Loss of Epithelial Integrity. Cell Rep. 2015;12:272-85
6. Nakamura J, Tanaka T, Kitajima Y, Noshiro H, Miyazaki K. Methylation-mediated gene silencing as biomarkers of gastric cancer: a review. World J Gastroenterol. 2014;20:11991-2006
7. Bass AJ TV, Shmulevich I, Reynolds SM, Miller M, Bernard B, Hinoue T, Laird PW, Curtis C, Shen H. et al. Comprehensive molecular characterization of gastric adenocarcinoma. Nature. 2014;513:202-9
8. Song JH, Meltzer SJ. MicroRNAs in pathogenesis, diagnosis, and treatment of gastroesophageal cancers. Gastroenterology. 2012;143:35-47 e2
9. Kim HP, Cho GA, Han SW, Shin JY, Jeong EG, Song SH. et al. Novel fusion transcripts in human gastric cancer revealed by transcriptome analysis. Oncogene. 2014;33:5434-41
10. Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281-97
11. Li XH, Zhang Y, Zhang YF, Ding J, Wu KC, Fan DM. Survival prediction of gastric cancer by a seven-microRNA signature. Gut. 2010;59:579-85
12. Zheng B, Liang L, Wang C, Huang S, Cao X, Zha R. et al. MicroRNA-148a suppresses tumor cell invasion and metastasis by downregulating ROCK1 in gastric cancer. Clin Cancer Res. 2011;17:7574-83
13. Tie J, Pan Y, Zhao L, Wu K, Liu J, Sun S. et al. MiR-218 inhibits invasion and metastasis of gastric cancer by targeting the Robo1 receptor. PLoS Genet. 2010;6:e1000879
14. Han TS, Hur K, Xu G, Choi B, Okugawa Y, Toiyama Y. et al. MicroRNA-29c mediates initiation of gastric carcinogenesis by directly targeting ITGB1. Gut. 2015;64:203-14
15. Long Q, Johnson BA, Osunkoya AO, Lai YH, Zhou W, Abramovitz M. et al. Protein-coding and microRNA biomarkers of recurrence of prostate cancer following radical prostatectomy. Am J Pathol. 2011;179:46-54
16. Kim YW, Kim EY, Jeon D, Liu JL, Kim HS, Choi JW. et al. Differential microRNA expression signatures and cell type-specific association with Taxol resistance in ovarian cancer cells. Drug Des Devel Ther. 2014;8:293-314
17. Rawlings-Goss RA, Campbell MC, Tishkoff SA. Global population-specific variation in miRNA associated with cancer risk and clinical biomarkers. Bmc Medical Genomics. 2014:7
18. Cao W, Wei W, Zhan Z, Xie D, Xie Y, Xiao Q. Role of miR-647 in human gastric cancer suppression. Oncol Rep. 2017;37:1401-11
19. Yang B, Jing C, Wang J, Guo X, Chen Y, Xu R. et al. Identification of microRNAs associated with lymphangiogenesis in human gastric cancer. Clin Transl Oncol. 2014;16:374-9
20. Bell JL, Haak AJ, Wade SM, Sun Y, Neubig RR, Larsen SD. Design and synthesis of tag-free photoprobes for the identification of the molecular target for CCG-1423, a novel inhibitor of the Rho/MKL1/SRF signaling pathway. Beilstein J Org Chem. 2013;9:966-73
21. Edge SB, Compton CC. The American Joint Committee on Cancer: the 7th Edition of the AJCC Cancer Staging Manual and the Future of TNM. Annals of Surgical Oncology. 2010;17:1471-4
22. Ji S, Ye G, Zhang J, Wang L, Wang T, Wang Z. et al. miR-574-5p negatively regulates Qki6/7 to impact beta-catenin/Wnt signalling and the development of colorectal cancer. Gut. 2013;62:716-26
23. Patnaik SK, Kannisto E, Knudsen S, Yendamuri S. Evaluation of microRNA expression profiles that may predict recurrence of localized stage I non-small cell lung cancer after surgical resection. Cancer Res. 2010;70:36-45
24. Nam S, Kim B, Shin S, Lee S. miRGator: an integrated system for functional annotation of microRNAs. Nucleic Acids Research. 2008;36:D159-D64
25. Medjkane S, Perez-Sanchez C, Gaggioli C, Sahai E, Treisman R. Myocardin-related transcription factors and SRF are required for cytoskeletal dynamics and experimental metastasis. Nat Cell Biol. 2009;11:257-68
26. Wang Y, Liu C, Luo M, Zhang Z, Gong J, Li J. et al. Chemotherapy-Induced miRNA-29c/Catenin-delta Signaling Suppresses Metastasis in Gastric Cancer. Cancer Res. 2015;75:1332-44
27. Liu D, Zhang L, Shen Z, Tan F, Hu Y, Yu J. et al. Clinicopathological significance of NMIIA Overexpression in Human Gastric Cancer. Int J Mol Sci. 2012;13:15291-304
28. Zhao X, He L, Li T, Lu Y, Miao Y, Liang S. et al. SRF expedites metastasis and modulates the epithelial to mesenchymal transition by regulating miR-199a-5p expression in human gastric cancer. Cell Death Differ. 2014;21:1900-13
29. Liu J, McCleland M, Stawiski EW, Gnad F, Mayba O, Haverty PM. et al. Integrated exome and transcriptome sequencing reveals ZAK isoform usage in gastric cancer. Nat Commun. 2014;5:3830
30. Hiramoto-Yamaki N, Takeuchi S, Ueda S, Harada K, Fujimoto S, Negishi M. et al. Ephexin4 and EphA2 mediate cell migration through a RhoG-dependent mechanism. Journal of Cell Biology. 2010;190:461-77
31. Bai SM, Nasser MW, Wang B, Hsu SH, Datta J, Kutay H. et al. MicroRNA-122 Inhibits Tumorigenic Properties of Hepatocellular Carcinoma Cells and Sensitizes These Cells to Sorafenib. Journal of Biological Chemistry. 2009;284:32015-27
32. Evelyn CR, Bell JL, Ryu JG, Wade SM, Kocab A, Harzdorf NL. et al. Design, synthesis and prostate cancer cell-based studies of analogs of the Rho/MKL1 transcriptional pathway inhibitor, CCG-1423. Bioorg Med Chem Lett. 2010;20:665-72
33. Furukawa T, Fu X, Kubota T, Watanabe M, Kitajima M, Hoffman RM. Nude mouse metastatic models of human stomach cancer constructed using orthotopic implantation of histologically intact tissue. Cancer Res. 1993;53:1204-8
34. Jin WZ, Goldfine AB, Boes T, Henry RR, Ciaraldi TP, Kim EY. et al. Increased SRF transcriptional activity in human and mouse skeletal muscle is a signature of insulin resistance. Journal of Clinical Investigation. 2011;121:918-29
35. Wang X, Guo B, Li Q, Peng J, Yang Z, Wang A. et al. miR-214 targets ATF4 to inhibit bone formation. Nat Med. 2013;19:93-100
36. Krutzfeldt J, Rajewsky N, Braich R, Rajeev KG, Tuschl T, Manoharan M. et al. Silencing of microRNAs in vivo with 'antagomirs'. Nature. 2005;438:685-9
37. Zhang LF, Lou JT, Lu MH, Gao C, Zhao S, Li B. et al. Suppression of miR-199a maturation by HuR is crucial for hypoxia-induced glycolytic switch in hepatocellular carcinoma. EMBO J. 2015;34:2671-85
38. Croft DR, Sahai E, Mavria G, Li SX, Tsai J, Lee WMF. et al. Conditional ROCK activation in vivo induces tumor cell dissemination and angiogenesis. Cancer Research. 2004;64:8994-9001
39. Yamaguchi H, Condeelis J. Regulation of the actin cytoskeleton in cancer cell migration and invasion. Biochimica Et Biophysica Acta-Molecular Cell Research. 2007;1773:642-52
40. Cen B, Selvaraj A, Burgess RC, Hitzler JK, Ma Z, Morris SW. et al. Megakaryoblastic leukemia 1, a potent transcriptional coactivator for serum response factor (SRF), is required for serum induction of SRF target genes. Mol Cell Biol. 2003;23:6597-608
41. Vartiainen MK, Guettler S, Larijani B, Treisman R. Nuclear actin regulates dynamic subcellular localization and activity of the SRF cofactor MAL. Science. 2007;316:1749-52
42. Psichari E, Balmain A, Plows D, Zoumpourlis V, Pintzas A. High activity of serum response factor in the mesenchymal transition of epithelial tumor cells is regulated by RhoA signaling. J Biol Chem. 2002;277:29490-5
43. Liang S, He L, Zhao X, Miao Y, Gu Y, Guo C. et al. MicroRNA let-7f inhibits tumor invasion and metastasis by targeting MYH9 in human gastric cancer. PLoS One. 2011;6:e18409
44. Srinivasan S, Selvan ST, Archunan G, Gulyas B, Padmanabhan P. MicroRNAs -the next generation therapeutic targets in human diseases. Theranostics. 2013;3:930-42
45. Sun S, Wang Y, Zhou R, Deng Z, Han Y, Han X. et al. Targeting and Regulating of an Oncogene via Nanovector Delivery of MicroRNA using Patient-Derived Xenografts. Theranostics. 2017;7:677-93
46. Rey C, Faustin B, Mahouche I, Ruggieri R, Brulard C, Ichas F. et al. The MAP3K ZAK, a novel modulator of ERK-dependent migration, is upregulated in colorectal cancer. Oncogene. 2016;35:3190-200
Author contact
![]() Corresponding authors: Xiaolong Qi, Guoxin Li & Hao Liu, Department of General Surgery, Nanfang Hospital, Southern Medical University, Guangdong Provincial Engineering Technology Research Center of Minimally Invasive Surgery; Tel.: +86 20 61641681/ fax: +86 20 61641683; Xiaolong Qi, E-mail: qixiaolong163.com; Guoxin Li, E-mail: gzliguoxincom; Hao Liu, E-mail: liuhaofbicom
Corresponding authors: Xiaolong Qi, Guoxin Li & Hao Liu, Department of General Surgery, Nanfang Hospital, Southern Medical University, Guangdong Provincial Engineering Technology Research Center of Minimally Invasive Surgery; Tel.: +86 20 61641681/ fax: +86 20 61641683; Xiaolong Qi, E-mail: qixiaolong163.com; Guoxin Li, E-mail: gzliguoxincom; Hao Liu, E-mail: liuhaofbicom
 Global reach, higher impact
Global reach, higher impact