Impact Factor
Theranostics 2012; 2(4):363-373. doi:10.7150/thno.3724 This issue Cite
Review
Theranostic Potential of Oncolytic Vaccinia Virus
Department of Surgery, University of Pittsburgh Cancer Institute, University of Pittsburgh, Pittsburgh, PA, USA
Received 2011-11-17; Accepted 2012-1-18; Published 2012-4-5
Abstract
Biological cancer therapies, such as oncolytic, or replication-selective viruses have advantages over traditional therapeutics as they can employ multiple different mechanisms to target and destroy cancers (including direct cell lysis, immune activation and vascular collapse). This has led to their rapid recent clinical development. However this also makes their pre-clinical and clinical study complex, as many parameters may affect their therapeutic potential and so defining reason for treatment failure or approaches that might enhance their therapeutic activity can be complicated. The ability to non-invasively image viral gene expression in vivo both in pre-clinical models and during clinical testing will considerably enhance the speed of oncolytic virus development as well as increasing the level and type of useful data produced from these studies. Further, subsequent to future clinical approval, imaging of reporter gene expression might be used to evaluate the likelihood of response to oncolytic viral therapy prior to changes in tumor burden. Here different reporter genes used in conjunction with oncolytic viral therapy are described, along with the imaging modalities used to measure their expression, while their applications both in pre-clinical and clinical testing are discussed. Possible future applications for reporter gene expression from oncolytic viruses in the phenotyping of tumors and the personalizing of treatment regimens are also discussed.
Keywords: Oncolytic Vaccinia Virus, Theranostics
1. Introduction
There has been a recent burst of interest in the development of biological therapies for the treatment of diseases such as cancer, with a variety of platforms and therapies displaying promising early clinical results or having moved into later stage clinical testing. The likelihood that a number of these will receive approval over the next decade means that the role of imaging of reporter gene expression in their pre-clinical and clinical applications will be a key factor in their future development. Here we will examine the role of imaging in the development and current and future applications of oncolytic viruses, focusing on the use of vaccinia viral vectors.
2. Oncolytic Viruses
A. Background
Oncolytic viruses are cancer therapeutics based on viruses whose replication is restricted to malignant cells (Fig. 1) [1-3]. In general this tumor selectivity can be achieved in one of two ways; (i) some viruses that normally do not cause disease in humans can nevertheless replicate in cancer cells (where the interferon (IFN) anti-viral response is frequently non-functional) or in tumors (where an immunosuppressive environment exists). These are typically small viruses with fast replication cycles, such as reovirus[4, 5], Newcastle Disease Virus[6] or VSV[7]; (ii) the second group of oncolytic vectors are based on viruses that are either used as vaccines against common disease-causing viruses (such as vaccinia virus[8] or the Edmonton strain of measles virus[9]) or on viruses that themselves cause known disease in humans (such as adenovirus[10], HSV[11] or poliovirus[12]). These tend to be larger viruses that are amenable to genetic engineering to produce or enhance their tumor selectivity. This increased selectivity is normally achieved through the deletion of viral virulence genes that are redundant for viral replication in tumor cells. As a result viral replication is attenuated in normal tissues, but proceeds normally in cancer cells. Because many of the alterations produced in a cancer cell during transformation are similar to the adaptations that a virus needs to induce in a cell for successful replication[13, 14], many such virulence genes exist (as witnessed by the oncogenic properties of some viral genes and the production of some cancers as a result of chronic viral infections). These viral gene products may fit into one of several different categories, including immune modulators (that are not required in the immunosuppressive tumor environment)[15]; anti-apoptotic proteins[16]; or inducers of cellular proliferation[17], meaning that different viral vectors (sometimes even produced from the same viral backbone) may target tumors based on unique or independent tumorogenic properties. Additional approaches to achieving tumor-selective replication of oncolytic viruses have also met with some success. These include the use of tissue or tumor specific promoters to drive expression of an essential viral gene[18-21]; and the alteration of viral-surface receptors to selectively target ligands that are highly expressed on tumor cells or in the tumor microenvironment[22-24].
Representation of the selective targeting and tumor-selective amplification of oncolytic viral vectors, such as oncolytic vaccinia.
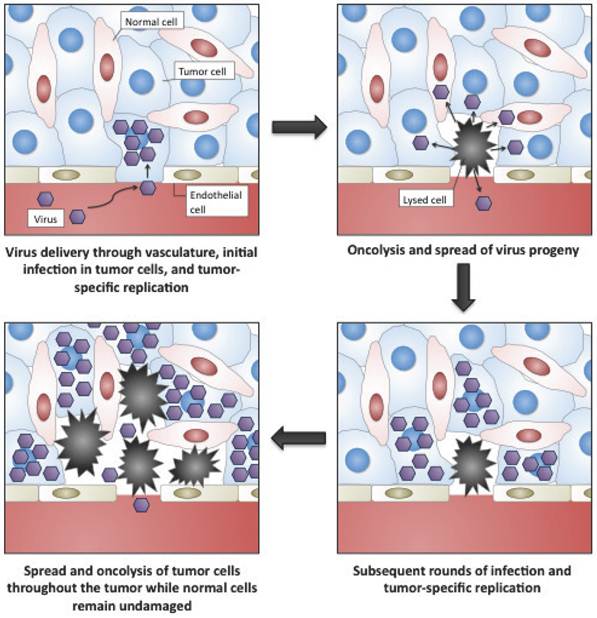
The first proposed use of replication competent viruses to treat cancer was over 100 years ago[25], while the first clinical trials were attempted over 60 years ago[26]. However, it was not until our understanding of cancer biology, virology and molecular biology reached a point that allowed for the logical construction of viruses with pre-determined tumor-targeting properties some 20 years ago that the field of oncolytic virology really come into being[27]. The rapid subsequent clinical progression of an oncolytic adenovirus (ONYX-015)[28, 29] towards Phase III testing reflected the early promise of the field[30, 31], however despite proven safety, Adenovirus was felt to have insufficient anti-tumor effects and so researchers turned to more rapidly replicating and lytic viruses to act as the backbone of oncolytic agents. Currently, the highly promising clinical data with several of these second generation vectors has re-ignited interest in the field[32-35], and it is likely that one or more of these viruses will be approved in the North American market in the next decade.
One advantage of oncolytic viruses is that they are known to destroy tumors by several distinct mechanisms, which typically do not overlap with the mechanisms induced by traditional therapies[8, 36, 37]. In addition to directly destroying infected tumor cells as a result of infection (which also leads to amplification of viral copies within the tumor), many oncolytic vectors can induce a potent immune response within the tumor. This immune response can overcome localized immune suppression, and may even create an in situ vaccination effect through cross-presentation of tumor associated antigens to the host immune response. Furthermore several viruses have been demonstrated to induce a robust vascular collapse within the tumor that is capable of destroying further tumor cells[38, 39]. Because oncolytic viruses express their genomes primarily within the tumor and amplify the copy numbers of their genes in the tumor microenvironment, their effects can be enhanced through the expression of therapeutic transgenes[40, 41]. A variety of such transgenes have been investigated expressed from these vectors, including those encoding cytokines, pro-drug converting enzymes and anti-angiogenic proteins. However, another approach that has begun to be more actively explored is the additional expression of reporter transgenes from these viruses, and this will be the focus of this review.
B. Oncolytic Vaccinia Virus
Here we will primarily focus on oncolytic vaccinia virus vectors for several reasons; (i) the field of oncolytic viruses is too large to satisfactorily cover all viruses in a single review; (ii) vaccinia is one of the most successful of the current generation of clinical vectors, with highly promising Phase I and Phase II clinical data being reported in melanoma[42] and HCC[33], and with systemic delivery demonstrated in the clinic[32]; (iii) vaccinia's large genome means that a variety of different oncolytic constructs have been reported[8]. It is also capable of expressing at least 25KB of foreign DNA[43], meaning that expression of multiple genes from a single vector is possible.
Vaccinia is an excellent candidate to produce oncolytic vectors for several further reasons[44, 45], including (i) it has a rapid and lytic replication cycle; (ii) it produces a potent immune response; (iii) it has been extensively used in humans during the smallpox eradication campaign[46], meaning that contraindications are well defined and approved anti-virals are available[47, 48]; and (iv) it has evolved to spread in the vascular system of the host[49], meaning that systemic delivery and spread between tumors has been demonstrated in pre-clinical[32, 50] and clinical testing. Vaccinia also has well defined molecular biology, including standard cloning techniques and a choice of natural and synthetic promoters[51, 52] with different strengths and expression patterns. This means that viral reporter gene expression can be linked to certain events, such as viral replication.
Several different oncolytic strains of vaccinia have been reported, with perhaps the most widely studied in pre-clinical models being vvDD[17, 50]. This is a version of the Western Reserve (WR) strain of vaccinia with deletions in the viral thymidine kinase gene[53] (meaning that replication is dependent on host cell TK expression, that is upregulated in most tumors[54]), and in the viral growth factor (VGF)(a secreted growth factor homolog that binds the EGF-R, inducing proliferation in surrounding cells[55], but that is redundant in >80% of cancers where mutations in the EGF-R signaling pathway mean it is unregulated). This virus has displayed significant selectivity, systemic delivery and robust anti-tumor potential in many pre-clinical models, and has recently completed Phase I clinical testing (Bartlett, unpublished data). However, the vaccinia strain that has been most extensively developed in the clinic is JX-594 (Jennerex, Inc), a version of the Wyeth strain of vaccinia with a single TK gene deletion and expressing GM-CSF to augment its anti-tumor effects[56]. This strain displayed highly promising Phase I data in a small intralesional trial in melanoma patients[33, 42], and in a Phase I study of HCC with intratumoral injection[33]. It has also demonstrated systemic delivery potential in the clinic and so further intravenous trials and Phase III testing are ongoing. A further vaccinia strain (Lister) with deletions in the viral thymidine kinase and the viral hemaglutinin gene has also been described in pre-clinical studies (GLV-1h68)[57, 58] and has entered clinical testing, although no clinical results have been reported to date.
However, in addition to these clinical strains a variety of additional oncolytic vaccinia vectors have been described in pre-clinical studies, these include a strain with deletion of the viral type I IFN binding protein (B18R) that selectively replicates in cells with loss of an IFN-response phenotype[15]; viruses with enhanced spread to treat metastatic tumors[59]; and a virus with deletions in the viral serpins that targets cells with a loss of anti-apoptotic potential[60]. Thus vaccinia has the capability to target different cancers with vectors targeted to different phenotypic properties.
The therapeutic activity of oncolytic vaccinia virus has been covered in several recent reviews [8, 37, 61, 62] and so we will focus on non-invasice imaging and diagnostic applications of this vector here.
C. Role of the Immune Response
One of the different mechanisms that oncolytic viruses, such as those based on vaccinia, utilize to destroy their tumor targets is the induction of a potent immune response within the tumor. Although in some situations the immune response may limit the direct oncolytic potential of these vectors, it has become apparent that the immune response raised by the virus can provide significant therapeutic benefit. The action of the virus in the tumor has been demonstrated to overcome localized immune suppression that is typically found within the tumor, and can even induce specific vaccination for relevant tumor antigens[37, 63, 64]. In this respect, oncolytic viruses such as those based on vaccinia may be thought of as personalized approaches to inducing relevant vaccination to help eradicate residual tumor cells within a host, and to provide memory immune surveillance for long term prevention of relapse. It is also likely that systemic measurements of the level and type of immune response induced by the viral treatment may ultimately be used either as an early prognostic indicator of therapeutic activity, or may help elucidate the immune properties of the tumor being treated, and so assist in the design of subsequent immunotherapeutic treatments.
3. Applications for the imaging of oncolytic viruses
Non-invasive imaging of reporter transgene expression is a powerful tool that can provide precise and unique spacio-temporal measurements of viral replication in vivo. In preclinical models, terminal assays in which cohorts of animals must be sacrificed at multiple time points in order to track viral biodistribution present numerous drawbacks, such as the inability to perform kinetic studies in one animal or the requirement to section the whole animal to not miss unexpected localizations. Similarly, in clinical trials, biopsies are also restricted to certain target organs and frequently involve surgery, which limits the number of samples available for the study. Arming oncolytic viruses with reporter genes that can be detected by imaging is an approach that can overcome these limitations [65]. Furthermore, these agents can be used for a wide range of other applications, such as early-stage tumor diagnosis or detection of residual disease after surgery. In this section, we focus on the different transgenes that have been used for imaging oncolytic viral replication, as well as the possible pre-clinical and clinical applications of these agents.
A. Bioluminescence imaging
Bioluminescence imaging exploits the emission of visible photons at specific wavelengths based on energy-dependent reactions catalyzed by luciferases [66]. These reactions require a substrate, and for insect luciferases also require ATP, magnesium and oxygen. The bioluminescence emitted can be detected and amplified using sensitive detection systems, revealing the sites of luciferase expression and activity. The luciferase family includes a variety of photoproteins, such as firefly luciferase, which metabolizes luciferin into oxyluciferin, or Gaussia and Renilla luciferases, which use coelenterazine as a substrate. By virtue of the wavelength of light produced leading to the most efficient transmission through tissues and the solubility of its substrate [67], the luc luciferase from firefly species (Photinus) is the most commonly used luciferase for in vivo imaging. The rapid imaging of many animals, ease of use of the imaging instrumentation and inexpensive nature of bioluminescence imaging means that it has become a powerful research tool for the pre-clinical development of novel therapies.
The applications of bioluminescence imaging in biomedicine are extensive. It has been used for tracking bacterial pathogens [68], to study gene expression patterns [69], to monitor tumor cell growth and regression [70], to determine the location and proliferation of stem cells [71] and to track gene expression [72]. In the context of oncolytic viruses, bioluminiscence has been used primarily to monitor viral replication in vivo and biodistribution in preclinical models (Fig 2). The luciferase gene has been cloned and used for tracking the replication of oncolytic parvoviruses [73], adenoviruses [74, 75], HSV-1 [76, 77], vaccinia virus [15, 58], measles virus [78] and VSV [79]. Furthermore, the infection of carrier cells with oncolytic viruses expressing luciferase can be used to evaluate the biodistribution of those cells. This strategy was used to evaluate the biodistribution of carrier CIK (Cytokine Induced Killer) cells loaded with vaccinia virus expressing luciferase [80] or T-cells loaded with measles virus [81].
Patterns of infection, replication and biodistribution and persistence within the tumor after intravenous delivery of oncolytic vaccinia (vvDD) expressing luciferase as determined by bioluminescence imaging on an IVIS200 (Xenogen, part of Caliper, now part of Perkin Elmer)
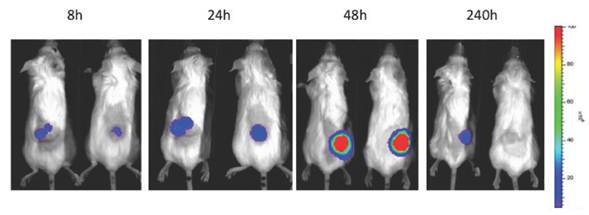
A novel and interesting strategy for a clinical application of luciferase-expressing oncolytic viruses is the use of such agents to predict the therapeutic outcome of the oncolytic therapy. In preclinical models, Davydova and collaborators demonstrated the predictive value of an early-time point imaging of adenoviral replication in tumors [82]. The in vivo luciferase bioluminescence measured 6 days after viral administration significantly correlated with the antitumor effect observed at day 36. On the other hand, bioluminescence can also be used for revealing the locations of primary tumors and metastases in animals. Yu and collaborators provided evidence that an intravenously delivered oncolytic vaccinia virus expressing the Renilla luciferase replicated in the tumor tissue and permitted the delineation of the location of tumors and metastases [83]. Despite the fact that luciferase expression and bioluminescence imaging cannot currently be used in humans, these studies provide proof-of-concept that oncolytic viruses expressing clinical-friendly imaging systems may improve the outcome of clinical protocols.
B. Fluorescence imaging
Fluorescent proteins have been extensively used as reporter genes in oncolytic viruses, especially for tracking the replication of the virus in vitro. They offer the possibility to image viral replication rapidly and inexpensively. The fluorescent protein most commonly cloned into oncolytic viruses is GFP, a 27-kDa protein form jellyfish Aequorea victoria that fluoresces green upon illumination with UV light [84]. This wild-type protein and its enhanced-fluorescent form have been cloned into the majority of oncolytic viruses, including Newcastle disease virus [85], VSV [86], HSV-1 [87], measles [88], adenovirus [89] and vaccinia virus [17].
The use of GFP for monitoring virus replication within organs in living animals is limited due to very low tissue penetrance of GFP excitation and emission wavelengths (1-2 mm). To overcome this limitation, novel strategies are being developed, such as the combination of fiber optic monitoring coupled with confocal microscopy to allow direct, rapid and sensitive visualization of fluorescent signals in the brain [90]. However, Li and collaborators recently successfully imaged a GFP-expressing Newcastle disease virus using a Maestro in vivo fluorescence imaging system (CRi, part of Caliper Life Sciences, part of Perkin Elmer), demonstrating the possibility of using this protein for in vivo imaging [91]. Despite this, the use of fluorescent proteins that fluoresce in the far-red and near-infrared spectrum maximize tissue penetrance and minimize the level of autofluorescence. Proteins such as the far-red fluorescent protein TurboFP635 [92] can be expressed from oncolytic viruses and imaged using advanced imaging methods, such as fluorescence molecular tomography (FMT, such as with the FMT2500, VisEn, part of Perkin Elmer). In FMT, the subject is exposed to continuous wave or pulsed light, and the emitted light is captured by detectors in an imaging chamber, arranged in a spatially defined order [72]. A tomographic image is reconstituted after the data is mathematically processed and quantitative, three-dimensional molecular information can be extracted.
Besides monitoring virus replication, oncolytic viruses expressing fluorescent proteins have been tested pre-clinically for a variety of putative clinical applications. As with luciferase-expressing viruses, oncolytic viruses expressing fluorescent proteins might also be applied to predict therapeutic outcome. A novel method under development is the infection of tissues from patients ex vivo prior to therapy in order to ascertain the likelihood of successful oncolytic virotherapy [93]. Demonstrating functionality of the oncolytic agent via fluorescence imaging within ex vivo tissues may be a useful bridge between pre-clinical studies and clinical trials. Alternatively, GFP-expressing viruses have also been proposed for precise surgical navigation due to its ability to infect and express GFP selectively in cancer cells. Kishimoto and collaborators used a GFP-expressing adenovirus for fluorescence-guided surgery in a model of intraperitoneal disseminated colon cancer and in a model of pleural disseminated lung cancer [94]. Five days after virus intraperitoneal or intrapleural administration, a fluorescence-guided laparotomy permitted the resection of disseminated cancer nodules, which would otherwise be undetectable. Furthermore, an HSV-1 expressing GFP was also used for this purpose. In a model of peritoneal carcinomatosis (PC), the administration of the oncolytic virus forty-eight hours prior to surgery allowed the detection of residual disease in 8 of 13 mice that experienced surgeons presumed complete cytoreduction [95]. Residual disease was identified in sites corresponding to patterns of recurrence in a published human series, highlighting the potential of combining surgery with virally-directed fluorescence imaging. Moreover, this technique has also been used for the detection of lymph node metastases. In a model of metastatic melanoma, Kelly and collaborators demonstrated that the administration of an engineered vaccinia virus expressing GFP into the primary tumor resulted in viral transmission to lymph nodes, infection of lymphatic metastases, and transgene expression that was reliably and easily detected [96]. The real-time fluorescence detection of nodal disease was 80% sensitive and 100% specific for identification of sentinel node disease. In a model of metastatic breast cancer, the injection of a GFP-expressing HSV-1 directly into primary breast tumors also resulted in viral transit to axillary lymph nodes, infection of lymphatic metastases, and GFP expression [97]. This technique permitted the real-time intraoperative identification of micrometastases otherwise undetectable due to the impracticality of performing extensive histologic analyses on a large number of resected nodes.
C. Nuclear medicine-based imaging
Nuclear medicine-based imaging is reliant on the ability of scanners to detect and localize gamma ray emission from the decay of a radiotracer. The scanners used are highly sensitive and provide good spatial resolution, as well as offering the greatest potential for translation into clinical applications, but require the use of more expensive cameras and radioactivity during their pre-clinical development [98]. The gamma camera, and PET (Positron Emission Tomography) and SPECT (Single-Photon Emission Computed Tomography) scanners are the principal types of scanners used in nuclear medicine. When applied to oncolytic viruses expressing a reporter transgene, viral replication can be monitored as a result of the alteration of the biodistribution of a tracer molecule at the sites of expression of the reporter gene, leading to a local concentration of the tracer at the site of viral replication.
Three types of reporter gene families have been cloned into oncolytic viruses. In the first family, the accumulation of the radio-labelled substrate is based on enzymatic activities that trap the tracer inside target cells when phosphorylated by the enzyme. When working with HSV-1, the endogenous thymidine kinase can be used for this purpose. FIAU and [18F]FHBG have been used previously as the substrate for HSV-TK to successfully image the viral replication in animal models using a PET scanner [99, 100]. Furthermore, a Sindbis virus expressing HSV-TK was also successfully imaged using 18F-FEAU as a substrate with a PET scanner [101]. However, in an attempt to monitor the replication of an oncolytic HSV-1 in glioma patients, no evidence of viral replication was found using a 123I-FIAU brain SPECT scanner [102]. This unsuccessful result was probably due to limited expression of endogenous HSV-TK and limited sensitivity of 123I and SPECT imaging, indicating the need for higher reporter gene expression, more potent reporter genes and/or greater sensitivity in radioisotope detection for the clinical setting.
The second family of reporter genes involves membrane receptors that bind and trap a radio-labeled substrate. The human somatostatin receptor SSTR2 was successfully used to monitor the replication of an oncolytic vaccinia virus in mice using 111In-pentetreotide as ligand and a gamma camera for the imaging [103]. Furthermore, the system dopamine DR2 receptor/123I-IBF or 11C-raclopride is also suitable for application to oncolytic viruses [104]. Finally, the third and most extensively examined family is that of membrane proteins that mediate ionic transport. The human norepinephrine transporter (hNES) has been used in the context of an oncolytic vaccinia virus to monitor its replication in mice [105], but the most broadly used ion transporter used for imaging is the human sodium iodide symporter (hNIS). This can be imaged using [123, 124, 131I]NaI, [99mTc]pertechnetate, or [186, 188Re]perrhenate, among others, and using scintigraphy, PET or SPECT scanners [65]. To monitor viral replication in preclinical models, it has been cloned into measles virus [106, 107], VSV [108], adenovirus [109, 110], and vaccinia virus [111]. In the clinical setting, the feasibility of using hNIS-mediated imaging was demonstrated in a phase I clinical trial treating prostate cancer patients with an oncolytic adenovirus coding for the hNIS gene [112]. In this study, viral replication was detected in 7 of 9 patients when 1x1012 vp were injected. However, Rajecki and collaborators failed to image the replication of a different oncolytic adenovirus expressing the hNIS gene [113]. In this study, a cervical metastatic carcinoma patient was treated with 3x1011 vp and a SPECT-CT scanner was used for the imaging. The difference in the success of the imaging may be explained by the dose administrated, the virus design, and the relative sensitivity of PET and SPECT scanners.
Alternative applications for the combination of nuclear medicine-based imaging with oncolytic viruses have also been described. As with oncolytic viruses expressing fluorescent proteins, the endogenous thymidine kinase coded by an oncolytic herpes virus has been used for the imaging and detection of lymph node micrometastases. In a pre-clinical model of metastatic melanoma, Brader and collaborators successfully identify 8 out of 8 tumor-positive nodes after an intratumoral injection of the oncolytic agent [114]. No overlap between radioactivity levels in tumor-positive and tumor-negative lymph nodes using [18F]FEAU PET was described. Furthermore, genes such as hNIS can be used for radiotherapy. In addition to imaging, the radiation physical crossfire effect permits the destruction of non-infected tumor by radiation emitted from neighboring cells transduced by recombinant oncolytic viruses [115]. This effect is of relevance because it allows for the destruction of intra-tumoral physical barriers such as stromal cells that can otherwise handicap the dispersion of the virus throughout the tumor. Importantly, results from xenograft tumors treated with an oncolytic measles virus expressing NIS showed complete regression of the tumor after 131I therapy [115, 116].
4. Future Directions
More extensive use of clinical imaging of reporter gene expression from oncolytic strains of vaccinia virus and other oncolytic backbones will be expected to validate the use of different reporter constructs to image viral gene expression, and to correlate this with viral replication patterns and therapeutic outcome, as has previously been shown in pre-clinical imaging models. This will allow the collection of significantly more data during clinical evaluation of oncolytic vectors, helping advance the clinical development of these therapeutics at a considerably increased pace. Furthermore, as one or more of these vectors may soon become approved agents for the treatments of some cancers in the North American and European markets, the ability to image at early times after their application in order to provide early markers of therapeutic activity will provide significant advantages, allowing for a more rapid switch to alternative therapies in the case of treatment failure.
However, beyond these basic clinical applications of imaging of reporter gene expression from oncolytic viruses to assess therapeutic response and possibly to define the margins of the tumor and locations of metastases for surgery, it is possible to also envisage other uses. Because different viruses may be targeted to different phenotypic properties within the tumor, it may be possible to create a virus that would specifically replicate in tumors that are also sensitive to particular targeted therapies. In this way viral replication (and gene expression) would act as a predictive marker for the likelihood of response to a secondary therapy. Alternatively, this may be extended with the possibility of producing multiple different oncolytic vectors (ideally based on the same viral backbone), targeted through different viral gene deletions to different phenotypic tumor properties and expressing different reporter genes. The simultaneous application of these viruses and the readout of their relative reporter gene expression may act as a more accurate method for defining the properties of the tumor and so would act as a means to direct personalized treatment regimens. For example, multiple vaccinia strains with different virulence gene deletions and expressing different fluorescent proteins could be applied into a single lesion of a melanoma patient, and an optical readout used to 'bar-code' the tumor and so define the treatment most likely to produce a response. Alternatively, this same approach may simply be used to determine which oncolytic viral strain would be most appropriate to subsequently treat the patient with at higher and/or systemic doses.
5. Discussion
The recent clinical advances with oncolytic viruses have led to a more focused interest on the expression of transgenes from within the therapeutic vectors themselves. Although this has traditionally involved the use of genes that might increase the therapeutic activity of the oncolytic virus (such as immune modulators, toxins, or pro-drug converting enzymes), advances in molecular imaging have led to the routine use of reporter genes for non-invasive imaging being additionally applied. This is of particular utility in larger viral agents (such as HSV-1 or vaccinia virus) that have the cloning capacity to express multiple reporter transgenes).
The benefits of reporter gene expression and non-invasive molecular imaging in pre-clinical models include more rapid and extensive descriptions of viral biodistribution and persistence of replication under different conditions. However, it is in the clinical application of reporter gene expression that the greatest benefits may ultimately be witnessed. These are likely to include early indicators of viral therapeutic activity and even as a means to locate tumor margins and the locations of metastases. It is likely that, at least initially, PET and SPECT imaging will be predominantly used in the clinic. This opens up the possibility of additionally applying radioisotopes that have therapeutic rather than imaging properties.
Finally, the ability to target viral oncolytic vectors to the tumor through different gene deletions, and so target different phenotypic properties of the tumor may also allow for the phenotyping of the tumor, leading to additional personalized medicine applications in tumor therapy.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Guo ZS, Thorne SH, Bartlett DL. Oncolytic virotherapy: Molecular targets in tumor-selective replication and carrier cell-mediated delivery of oncolytic viruses. Biochim Biophys Acta. 2008; doi:S0304-419X(08)00004-8 [pii] 10.1016/j.bbcan. 2008 02.001
2. Thorne SH, Hermiston T, Kirn D. Oncolytic virotherapy: approaches to tumor targeting and enhancing antitumor effects. Semin Oncol. 2005;32:537-48
3. Kirn D, Martuza RL, Zwiebel J. Replication-selective virotherapy for cancer: Biological principles, risk management and future directions. Nat Med. 2001;7:781-7
4. Harrington KJ, Vile RG, Melcher A, Chester J, Pandha HS. Clinical trials with oncolytic reovirus: moving beyond phase I into combinations with standard therapeutics. Cytokine Growth Factor Rev. 2010;21:91-8 doi:10.1016/j.cytogfr.2010.02.006
5. Van Den Wollenberg DJ, Van Den Hengel SK, Dautzenberg IJ, Kranenburg O, Hoeben RC. Modification of mammalian reoviruses for use as oncolytic agents. Expert opinion on biological therapy. 2009;9:1509-20 doi:10.1517/14712590903307370
6. Schirrmacher V, Fournier P. Newcastle disease virus: a promising vector for viral therapy, immune therapy, and gene therapy of cancer. Methods in molecular biology. 2009;542:565-605
7. Barber GN. VSV-tumor selective replication and protein translation. Oncogene. 2005;24:7710-9 doi:10.1038/sj.onc.1209042
8. Kirn DH, Thorne SH. Targeted and armed oncolytic poxviruses: a novel multi-mechanistic therapeutic class for cancer. Nat Rev Cancer. 2009;9:64-71 doi:nrc2545 [pii]10.1038/nrc2545
9. Msaouel P, Iankov ID, Dispenzieri A, Galanis E. Attenuated Oncolytic Measles Virus Strains as Cancer Therapeutics. Curr Pharm Biotechnol. 2011
10. Toth K, Wold WS. Increasing the efficacy of oncolytic adenovirus vectors. Viruses. 2010;2:1844-66 doi:10.3390/v2091844
11. Kaur B, Chiocca EA, Cripe TP. Oncolytic HSV-1 Virotherapy: Clinical Experience and Opportunities for Progress. Curr Pharm Biotechnol. 2011
12. Goetz C, Gromeier M. Preparing an oncolytic poliovirus recombinant for clinical application against glioblastoma multiforme. Cytokine Growth Factor Rev. 2010;21:197-203 doi:10.1016/j.cytogfr.2010.02.005
13. Hanahan D, Weinberg RA. The hallmarks of cancer. Cell. 2000;100:57-70
14. Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646-74 doi:10.1016/j.cell.2011.02.013
15. Kirn DH, Wang Y, Le Boeuf F, Bell J, Thorne SH. Targeting of interferon-beta to produce a specific, multi-mechanistic oncolytic vaccinia virus. PLoS Med. 2007;4:e353. doi:07-PLME-RA-0571 [pii]10.1371/journal.pmed.0040353
16. Guo ZS, Naik A, O'Malley ME, Popovic P, Demarco R, Hu Y. et al. The enhanced tumor selectivity of an oncolytic vaccinia lacking the host range and antiapoptosis genes SPI-1 and SPI-2. Cancer Research. 2005;65:9991-8 doi:10.1158/0008-5472.CAN-05-1630
17. McCart JA, Ward JM, Lee J, Hu Y, Alexander HR, Libutti SK. et al. Systemic cancer therapy with a tumor-selective vaccinia virus mutant lacking thymidine kinase and vaccinia growth factor genes. Cancer Res. 2001;61:8751-7
18. DeWeese TL, van der Poel H, Li S, Mikhak B, Drew R, Goemann M. et al. A phase I trial of CV706, a replication-competent, PSA selective oncolytic adenovirus, for the treatment of locally recurrent prostate cancer following radiation therapy. Cancer Res. 2001;61:7464-72
19. Li Y, Yu DC, Chen Y, Amin P, Zhang H, Nguyen N. et al. A hepatocellular carcinoma-specific adenovirus variant, CV890, eliminates distant human liver tumors in combination with doxorubicin. Cancer Res. 2001;61:6428-36
20. Yu DC, Chen Y, Dilley J, Li Y, Embry M, Zhang H. et al. Antitumor synergy of CV787, a prostate cancer-specific adenovirus, and paclitaxel and docetaxel. Cancer Res. 2001;61:517-25
21. Rojas JJ, Guedan S, Searle PF, Martinez-Quintanilla J, Gil-Hoyos R, Alcayaga-Miranda F. et al. Minimal RB-responsive E1A promoter modification to attain potency, selectivity, and transgene-arming capacity in oncolytic adenoviruses. Mol Ther. 2010;18:1960-71 doi:mt2010173 [pii]10.1038/mt.2010.173
22. Roelvink PW, Mi Lee G, Einfeld DA, Kovesdi I, Wickham TJ. Identification of a conserved receptor-binding site on the fiber proteins of CAR-recognizing adenoviridae. Science. 1999;286:1568-71
23. Douglas JT, Rogers BE, Rosenfeld ME, Michael SI, Feng M, Curiel DT. Targeted gene delivery by tropism-modified adenoviral vectors. Nat Biotechnol. 1996;14:1574-8
24. Peng KW, Donovan KA, Schneider U, Cattaneo R, Lust JA, Russell SJ. Oncolytic measles viruses displaying a single-chain antibody against CD38, a myeloma cell marker. Blood. 2003;101:2557-62
25. Dock G. Rabies virus vaccination in a patient with cervical carcinoma. American Journal of Medical Science. 1904;127:563
26. Southam CM, Moore AE. Clinical studies of viruses as antineoplastic agents with particular reference to Egypt 101 virus. Cancer. 1952;5:1025-34
27. Martuza RL, Malick A, Markert JM, Ruffner KL, Coen DM. Experimental therapy of human glioma by means of a genetically engineered virus mutant. Science. 1991;252:854-6
28. Bischoff JR, Kirn DH, Williams A, Heise C, Horn S, Muna M. et al. An adenovirus mutant that replicates selectively in p53-deficient human tumor cells. Science. 1996;274:373-6
29. Heise C, Sampson-Johannes A, Williams A, McCormick F, Von Hoff DD, Kirn DH. ONYX-015, an E1B gene-attenuated adenovirus, causes tumor-specific cytolysis and antitumoral efficacy that can be augmented by standard chemotherapeutic agents. Nat Med. 1997;3:639-45
30. Nemunaitis J, Khuri F, Ganly I, Arseneau J, Posner M, Vokes E. et al. Phase II trial of intratumoral administration of ONYX-015, a replication-selective adenovirus, in patients with refractory head and neck cancer. J Clin Oncol. 2001;19:289-98
31. Galanis E, Okuno SH, Nascimento AG, Lewis BD, Lee RA, Oliveira AM. et al. Phase I-II trial of ONYX-015 in combination with MAP chemotherapy in patients with advanced sarcomas. Gene Ther. 2005;12:437-45
32. Breitbach CJ, Burke J, Jonker D, Stephenson J, Haas AR, Chow LQ. et al. Intravenous delivery of a multi-mechanistic cancer-targeted oncolytic poxvirus in humans. Nature. 2011;477:99-102 doi:10.1038/nature10358
33. Park BH, Hwang T, Liu TC, Sze DY, Kim JS, Kwon HC. et al. Use of a targeted oncolytic poxvirus, JX-594, in patients with refractory primary or metastatic liver cancer: a phase I trial. Lancet Oncol. 2008;9:533-42 doi:S1470-2045(08)70107-4 [pii]10.1016/S1470-2045(08)70107-4
34. Schmidt C. Amgen spikes interest in live virus vaccines for hard-to-treat cancers. Nature biotechnology. 2011;29:295-6 doi:10.1038/nbt0411-295
35. Senzer NN, Kaufman HL, Amatruda T, Nemunaitis M, Reid T, Daniels G. et al. Phase II clinical trial of a granulocyte-macrophage colony-stimulating factor-encoding, second-generation oncolytic herpesvirus in patients with unresectable metastatic melanoma. J Clin Oncol. 2009;27:5763-71 doi:JCO.2009.24.3675 [pii]10.1200/JCO.2009.24.3675
36. Breitbach CJ, Thorne SH, Bell JC, Kirn DH. Targeted and Armed Oncolytic Poxviruses for Cancer: The Lead Example of JX-594. Curr Pharm Biotechnol. 2011
37. Thorne SH. Immunotherapeutic potential of oncolytic vaccinia virus. Immunol Res. 2011;50:286-93 doi:10.1007/s12026-011-8211-4
38. Breitbach CJ, Paterson JM, Lemay CG, Falls TJ, McGuire A, Parato KA. et al. Targeted inflammation during oncolytic virus therapy severely compromises tumor blood flow. Mol Ther. 2007;15:1686-93 doi:6300215 [pii]10.1038/sj.mt.6300215
39. Liu TC, Hwang T, Park BH, Bell J, Kirn DH. The targeted oncolytic poxvirus JX-594 demonstrates antitumoral, antivascular, and anti-HBV activities in patients with hepatocellular carcinoma. Mol Ther. 2008;16:1637-42 doi:mt2008143 [pii]10.1038/mt.2008.143
40. Hermiston T. Fighting fire with fire: attacking the complexity of human tumors with armed therapeutic viruses. Curr Opin Mol Ther. 2002;4:334-42
41. Hermiston TW, Kuhn I. Armed therapeutic viruses: strategies and challenges to arming oncolytic viruses with therapeutic genes. Cancer Gene Ther. 2002;9:1022-35
42. Mastrangelo MJ, Maguire HC Jr, Eisenlohr LC, Laughlin CE, Monken CE, McCue PA. et al. Intratumoral recombinant GM-CSF-encoding virus as gene therapy in patients with cutaneous melanoma. Cancer Gene Ther. 1999;6:409-22
43. Smith GL, Moss B. Infectious poxvirus vectors have capacity for at least 25 000 base pairs of foreign DNA. Gene. 1983;25:21-8
44. Moss B. Vaccinia virus: a tool for research and vaccine development. Science. 1991;252:1662-7
45. Moss B. Genetically engineered poxviruses for recombinant gene expression, vaccination, and safety. Proc Natl Acad Sci U S A. 1996;93:11341-8
46. Fenner F, Henderson DA, Arita I, Jezek Z, Ladnyi ID. Smallpox and its eradication. Geneva: World Health Organization. 1988
47. Cono J, Casey CG, Bell DM. Smallpox vaccination and adverse reactions. Guidance for clinicians. MMWR Recomm Rep. 2003;52:1-28
48. Wittek R. Vaccinia immune globulin: current policies, preparedness, and product safety and efficacy. Int J Infect Dis. 2006;10:193-201 doi:S1201-9712(06)00040-3 [pii]10.1016/j.ijid.2005.12.001
49. Buller RM, Palumbo GJ. Poxvirus pathogenesis. Microbiol Rev. 1991;55:80-122
50. Thorne SH, Hwang TH, O'Gorman WE, Bartlett DL, Sei S, Kanji F. et al. Rational strain selection and engineering creates a broad-spectrum, systemically effective oncolytic poxvirus, JX-963. J Clin Invest. 2007;117:3350-8 doi:10.1172/JCI32727
51. Davison AJ, Moss B. Structure of vaccinia virus late promoters. J Mol Biol. 1989;210:771-84
52. Davison AJ, Moss B. Structure of vaccinia virus early promoters. J Mol Biol. 1989;210:749-69
53. Puhlmann M, Brown CK, Gnant M, Huang J, Libutti SK, Alexander HR. et al. Vaccinia as a vector for tumor-directed gene therapy: biodistribution of a thymidine kinase-deleted mutant. Cancer Gene Ther. 2000;7:66-73
54. Hengstschlager M, Knofler M, Mullner EW, Ogris E, Wintersberger E, Wawra E. Different regulation of thymidine kinase during the cell cycle of normal versus DNA tumor virus-transformed cells. J Biol Chem. 1994;269:13836-42
55. Buller RM, Chakrabarti S, Moss B, Fredrickson T. Cell proliferative response to vaccinia virus is mediated by VGF. Virology. 1988;164:182-92
56. Kim JH, Oh JY, Park BH, Lee DE, Kim JS, Park HE. et al. Systemic Armed Oncolytic and Immunologic Therapy for Cancer with JX-594, a Targeted Poxvirus Expressing GM-CSF. Mol Ther. 2006;14:361-70
57. Yu YA, Timiryasova T, Zhang Q, Beltz R, Szalay AA. Optical imaging: bacteria, viruses, and mammalian cells encoding light-emitting proteins reveal the locations of primary tumors and metastases in animals. Anal Bioanal Chem. 2003;377:964-72
58. Zhang Q, Yu YA, Wang E, Chen N, Danner RL, Munson PJ. et al. Eradication of solid human breast tumors in nude mice with an intravenously injected light-emitting oncolytic vaccinia virus. Cancer Res. 2007;67:10038-46 doi:67/20/10038 [pii]10.1158/0008-5472.CAN-07-0146
59. Kirn DH, Wang Y, Liang W, Contag CH, Thorne SH. Enhancing poxvirus oncolytic effects through increased spread and immune evasion. Cancer Res. 2008;68:2071-5 doi:68/7/2071 [pii]10.1158/0008-5472.CAN-07-6515
60. Guo ZS, Naik A, O'Malley ME, Popovic P, Demarco R, Hu Y. et al. The enhanced tumor selectivity of an oncolytic vaccinia lacking the host range and antiapoptosis genes SPI-1 and SPI-2. Cancer Res. 2005;65:9991-8
61. Chen NG, Szalay AA. Oncolytic Vaccinia Virus: A theranostic agent for cancer. Future Virol. 2010;5:763-84
62. Thorne SH, Contag CH. Combining immune cell and viral therapy for the treatment of cancer. Cell Mol Life Sci. 2007;64:1449-51 doi:10.1007/s00018-007-6550-z
63. Contag CH, Sikorski R, Negrin RS, Schmidt T, Fan AC, Bachireddy P. et al. Definition of an enhanced immune cell therapy in mice that can target stem-like lymphoma cells. Cancer Research. 2010;70:9837-45 doi:10.1158/0008-5472.CAN-10-2650
64. Thorne SH, Liang W, Sampath P, Schmidt T, Sikorski R, Beilhack A. et al. Targeting localized immune suppression within the tumor through repeat cycles of immune cell-oncolytic virus combination therapy. Molecular therapy: the journal of the American Society of Gene Therapy. 2010;18:1698-705 doi:10.1038/mt.2010.140
65. Baril P, Martin-Duque P, Vassaux G. Visualization of gene expression in the live subject using the Na/I symporter as a reporter gene: applications in biotherapy. Br J Pharmacol. 2010;159:761-71 doi:BPH412 [pii]10.1111/j.1476-5381.2009.00412.x
66. Negrin RS, Contag CH. In vivo imaging using bioluminescence: a tool for probing graft-versus-host disease. Nat Rev Immunol. 2006;6:484-90 doi:nri1879 [pii]10.1038/nri1879
67. Zhao H, Doyle TC, Coquoz O, Kalish F, Rice BW, Contag CH. Emission spectra of bioluminescent reporters and interaction with mammalian tissue determine the sensitivity of detection in vivo. J Biomed Opt. 2005;10:41210
68. Contag CH, Contag PR, Mullins JI, Spilman SD, Stevenson DK, Benaron DA. Photonic detection of bacterial pathogens in living hosts. Mol Microbiol. 1995;18:593-603
69. Contag CH, Spilman SD, Contag PR, Oshiro M, Eames B, Dennery P. et al. Visualizing gene expression in living mammals using a bioluminescent reporter. Photochem Photobiol. 1997;66:523-31
70. Shah K, Tang Y, Breakefield X, Weissleder R. Real-time imaging of TRAIL-induced apoptosis of glioma tumors in vivo. Oncogene. 2003;22:6865-72
71. Cao YA, Wagers AJ, Beilhack A, Dusich J, Bachmann MH, Negrin RS. et al. Shifting foci of hematopoiesis during reconstitution from single stem cells. Proc Natl Acad Sci U S A. 2004;101:221-6
72. Weissleder R. Scaling down imaging: molecular mapping of cancer in mice. Nat Rev Cancer. 2002;2:11-8 doi:10.1038/nrc701
73. Maxwell IH, Chapman JT, Scherrer LC, Spitzer AL, Leptihn S, Maxwell F. et al. Expansion of tropism of a feline parvovirus to target a human tumor cell line by display of an alpha(v) integrin binding peptide on the capsid. Gene Ther. 2001;8:324-31 doi:10.1038/sj.gt.3301399
74. Nanda D, Vogels R, Havenga M, Avezaat CJ, Bout A, Smitt PS. Treatment of malignant gliomas with a replicating adenoviral vector expressing herpes simplex virus-thymidine kinase. Cancer Res. 2001;61:8743-50
75. Rivera AA, Wang M, Suzuki K, Uil TG, Krasnykh V, Curiel DT. et al. Mode of transgene expression after fusion to early or late viral genes of a conditionally replicating adenovirus via an optimized internal ribosome entry site in vitro and in vivo. Virology. 2004;320:121-34 doi:10.1016/j.virol.2003.11.028S0042682203008742 [pii]
76. Yamamoto S, Deckter LA, Kasai K, Chiocca EA, Saeki Y. Imaging immediate-early and strict-late promoter activity during oncolytic herpes simplex virus type 1 infection and replication in tumors. Gene Ther. 2006;13:1731-6 doi:3302831 [pii]10.1038/sj.gt.3302831
77. Veerapong J, Bickenbach KA, Shao MY, Smith KD, Posner MC, Roizman B. et al. Systemic delivery of (gamma1)34.5-deleted herpes simplex virus-1 selectively targets and treats distant human xenograft tumors that express high MEK activity. Cancer Res. 2007;67:8301-6 doi:67/17/8301 [pii]10.1158/0008-5472.CAN-07-1499
78. Msaouel P, Iankov ID, Allen C, Morris JC, von Messling V, Cattaneo R. et al. Engineered measles virus as a novel oncolytic therapy against prostate cancer. Prostate. 2009;69:82-91 doi:10.1002/pros.20857
79. Moussavi M, Fazli L, Tearle H, Guo Y, Cox M, Bell J. et al. Oncolysis of prostate cancers induced by vesicular stomatitis virus in PTEN knockout mice. Cancer Res. 2010;70:1367-76 doi:0008-5472.CAN-09-2377[pii]10.1158/0008-5472.CAN-09-2377
80. Thorne SH, Negrin RS, Contag CH. Synergistic antitumor effects of immune cell-viral biotherapy. Science. 2006;311:1780-4
81. Ong HT, Hasegawa K, Dietz AB, Russell SJ, Peng KW. Evaluation of T cells as carriers for systemic measles virotherapy in the presence of antiviral antibodies. Gene Ther. 2007;14:324-33 doi:3302880 [pii]10.1038/sj.gt.3302880
82. Davydova J, Gavrikova T, Brown EJ, Luo X, Curiel DT, Vickers SM. et al. In vivo bioimaging tracks conditionally replicative adenoviral replication and provides an early indication of viral antitumor efficacy. Cancer Sci. 2010;101:474-81 doi:CAS1407 [pii]10.1111/j.1349-7006.2009.01407.x
83. Yu YA, Shabahang S, Timiryasova TM, Zhang Q, Beltz R, Gentschev I. et al. Visualization of tumors and metastases in live animals with bacteria and vaccinia virus encoding light-emitting proteins. Nat Biotechnol. 2004;22:313-20
84. Chalfie M, Tu Y, Euskirchen G, Ward WW, Prasher DC. Green fluorescent protein as a marker for gene expression. Science. 1994;263:802-5
85. Song KY, Wong J, Gonzalez L, Sheng G, Zamarin D, Fong Y. Antitumor efficacy of viral therapy using genetically engineered Newcastle disease virus [NDV(F3aa)-GFP] for peritoneally disseminated gastric cancer. J Mol Med (Berl). 2010;88:589-96 doi:10.1007/s00109-010-0605-6
86. Ebert O, Shinozaki K, Huang TG, Savontaus MJ, Garcia-Sastre A, Woo SL. Oncolytic vesicular stomatitis virus for treatment of orthotopic hepatocellular carcinoma in immune-competent rats. Cancer Res. 2003;63:3605-11
87. Stanziale SF, Stiles BM, Bhargava A, Kerns SA, Kalakonda N, Fong Y. Oncolytic herpes simplex virus-1 mutant expressing green fluorescent protein can detect and treat peritoneal cancer. Hum Gene Ther. 2004;15:609-18 doi:10.1089/104303404323142051
88. Allen C, Vongpunsawad S, Nakamura T, James CD, Schroeder M, Cattaneo R. et al. Retargeted oncolytic measles strains entering via the EGFRvIII receptor maintain significant antitumor activity against gliomas with increased tumor specificity. Cancer Res. 2006;66:11840-50 doi:66/24/11840 [pii]10.1158/0008-5472.CAN-06-1200
89. Funston GM, Kallioinen SE, de Felipe P, Ryan MD, Iggo RD. Expression of heterologous genes in oncolytic adenoviruses using picornaviral 2A sequences that trigger ribosome skipping. J Gen Virol. 2008;89:389-96 doi:89/2/389 [pii]10.1099/vir.0.83444-0
90. Ilyin SE, Flynn MC, Plata-Salaman CR. Fiber-optic monitoring coupled with confocal microscopy for imaging gene expression in vitro and in vivo. J Neurosci Methods. 2001;108:91-6 doi:S0165-0270(01)00379-X [pii]
91. Li P, Chen CH, Li S, Givi B, Yu Z, Zamarin D. et al. Therapeutic effects of a fusogenic newcastle disease virus in treating head and neck cancer. Head Neck. 2011;33:1394-9 doi:10.1002/hed.21609
92. Shcherbo D, Merzlyak EM, Chepurnykh TV, Fradkov AF, Ermakova GV, Solovieva EA. et al. Bright far-red fluorescent protein for whole-body imaging. Nat Methods. 2007;4:741-6 doi:nmeth1083 [pii]10.1038/nmeth1083
93. Diallo JS, Roy D, Abdelbary H, De Silva N, Bell JC. Ex vivo infection of live tissue with oncolytic viruses. J Vis Exp. 2011 doi:2854 [pii]10.3791/2854
94. Kishimoto H, Zhao M, Hayashi K, Urata Y, Tanaka N, Fujiwara T. et al. In vivo internal tumor illumination by telomerase-dependent adenoviral GFP for precise surgical navigation. Proc Natl Acad Sci U S A. 2009;106:14514-7 doi:0906388106 [pii]10.1073/pnas.0906388106
95. Adusumilli PS, Eisenberg DP, Chun YS, Ryu KW, Ben-Porat L, Hendershott KJ. et al. Virally directed fluorescent imaging improves diagnostic sensitivity in the detection of minimal residual disease after potentially curative cytoreductive surgery. J Gastrointest Surg. 2005;9:1138-46 doi:S1091-255X(05)00556-1 [pii]10.1016/j.gassur.2005.06.029
96. Kelly KJ, Brader P, Woo Y, Li S, Chen N, Yu YA. et al. Real-time intraoperative detection of melanoma lymph node metastases using recombinant vaccinia virus GLV-1h68 in an immunocompetent animal model. Int J Cancer. 2009;124:911-8 doi:10.1002/ijc.24037
97. Eisenberg DP, Adusumilli PS, Hendershott KJ, Chung S, Yu Z, Chan MK. et al. Real-time intraoperative detection of breast cancer axillary lymph node metastases using a green fluorescent protein-expressing herpes virus. Ann Surg. 2006;243:824-30 doi:10.1097/01.sla.0000219738.56896.c000000658-200606000-00013 [pii]
98. Vassaux G, Groot-Wassink T. In Vivo Noninvasive Imaging for Gene Therapy. J Biomed Biotechnol. 2003;2003:92-101 doi:10.1155/S1110724303209050S1110724303209050 [pii]
99. Jacobs A, Tjuvajev JG, Dubrovin M, Akhurst T, Balatoni J, Beattie B. et al. Positron emission tomography-based imaging of transgene expression mediated by replication-conditional, oncolytic herpes simplex virus type 1 mutant vectors in vivo. Cancer Res. 2001;61:2983-95
100. Kuruppu D, Brownell AL, Zhu A, Yu M, Wang X, Kulu Y. et al. Positron emission tomography of herpes simplex virus 1 oncolysis. Cancer Res. 2007;67:3295-300 doi:67/7/3295 [pii]10.1158/0008-5472.CAN-06-4062
101. Tseng JC, Zanzonico PB, Levin B, Finn R, Larson SM, Meruelo D. Tumor-specific in vivo transfection with HSV-1 thymidine kinase gene using a Sindbis viral vector as a basis for prodrug ganciclovir activation and PET. J Nucl Med. 2006;47:1136-43 doi:47/7/1136 [pii]
102. Dempsey MF, Wyper D, Owens J, Pimlott S, Papanastassiou V, Patterson J. et al. Assessment of 123I-FIAU imaging of herpes simplex viral gene expression in the treatment of glioma. Nucl Med Commun. 2006;27:611-7 doi:00006231-200608000-00003 [pii]
103. McCart JA, Mehta N, Scollard D, Reilly RM, Carrasquillo JA, Tang N. et al. Oncolytic vaccinia virus expressing the human somatostatin receptor SSTR2: molecular imaging after systemic delivery using 111In-pentetreotide. Mol Ther. 2004;10:553-61
104. al-Tikriti MS, Baldwin RM, Zea-Ponce Y, Sybirska E, Zoghbi SS, Laruelle M. et al. Comparison of three high affinity SPECT radiotracers for the dopamine D2 receptor. Nucl Med Biol. 1994;21:179-88
105. Brader P, Kelly KJ, Chen N, Yu YA, Zhang Q, Zanzonico P. et al. Imaging a Genetically Engineered Oncolytic Vaccinia Virus (GLV-1h99) Using a Human Norepinephrine Transporter Reporter Gene. Clin Cancer Res. 2009;15:3791-801 doi:1078-0432.CCR-08-3236[pii]10.1158/1078-0432.CCR-08-323
106. Penheiter AR, Griesmann GE, Federspiel MJ, Dingli D, Russell SJ, Carlson SK. Pinhole micro-SPECT/CT for noninvasive monitoring and quantitation of oncolytic virus dispersion and percent infection in solid tumors. Gene Ther. 2011; doi:gt2011107 [pii]10.1038/gt. 2011 107
107. Msaouel P, Iankov ID, Allen C, Aderca I, Federspiel MJ, Tindall DJ. et al. Noninvasive imaging and radiovirotherapy of prostate cancer using an oncolytic measles virus expressing the sodium iodide symporter. Mol Ther. 2009;17:2041-8 doi:mt2009218 [pii]10.1038/mt.2009.218
108. Goel A, Carlson SK, Classic KL, Greiner S, Naik S, Power AT. et al. Radioiodide imaging and radiovirotherapy of multiple myeloma using VSV(Delta51)-NIS, an attenuated vesicular stomatitis virus encoding the sodium iodide symporter gene. Blood. 2007;110:2342-50 doi:blood-2007-01-065573 [pii]10.1182/blood-2007-01-065573
109. Hakkarainen T, Rajecki M, Sarparanta M, Tenhunen M, Airaksinen AJ, Desmond RA. et al. Targeted radiotherapy for prostate cancer with an oncolytic adenovirus coding for human sodium iodide symporter. Clin Cancer Res. 2009;15:5396-403 doi:1078-0432.CCR-08-2571[pii]10.1158/1078-0432.CCR-08-257
110. Merron A, Baril P, Martin-Duque P, de la Vieja A, Tran L, Briat A. et al. Assessment of the Na/I symporter as a reporter gene to visualize oncolytic adenovirus propagation in peritoneal tumours. Eur J Nucl Med Mol Imaging. 2010;37:1377-85 doi:10.1007/s00259-009-1379-3
111. Haddad D, Chen NG, Zhang Q, Chen CH, Yu YA, Gonzalez L. et al. Insertion of the human sodium iodide symporter to facilitate deep tissue imaging does not alter oncolytic or replication capability of a novel vaccinia virus. J Transl Med. 2011;9:36. doi:1479-5876-9-36 [pii]10.1186/1479-5876-9-36
112. Barton KN, Stricker H, Brown SL, Elshaikh M, Aref I, Lu M. et al. Phase I study of noninvasive imaging of adenovirus-mediated gene expression in the human prostate. Mol Ther. 2008;16:1761-9 doi:mt2008172 [pii]10.1038/mt.2008.172
113. Rajecki M, Kangasmaki A, Laasonen L, Escutenaire S, Hakkarainen T, Haukka J. et al. Sodium iodide symporter SPECT imaging of a patient treated with oncolytic adenovirus Ad5/3-Delta24-hNIS. Mol Ther. 2011;19:629-31 doi:mt201131 [pii]10.1038/mt.2011.31
114. Brader P, Kelly K, Gang S, Shah JP, Wong RJ, Hricak H. et al. Imaging of lymph node micrometastases using an oncolytic herpes virus and [F]FEAU PET. PLoS One. 2009;4:e4789. doi:10.1371/journal.pone.0004789
115. Dingli D, Peng KW, Harvey ME, Greipp PR, O'Connor MK, Cattaneo R. et al. Image-guided radiovirotherapy for multiple myeloma using a recombinant measles virus expressing the thyroidal sodium iodide symporter. Blood. 2004;103:1641-6 doi:10.1182/blood-2003-07-22332003-07-2233 [pii]
116. Mitrofanova E, Unfer R, Vahanian N, Link C. Rat sodium iodide symporter allows using lower dose of 131I for cancer therapy. Gene Ther. 2006;13:1052-6 doi:3302758 [pii]10.1038/sj.gt.3302758
Author contact
Corresponding author: thorneshedu
 Global reach, higher impact
Global reach, higher impact